
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,454,344 – 14,454,460 |
| Length | 116 |
| Max. P | 0.777697 |

| Location | 14,454,344 – 14,454,460 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.76 |
| Mean single sequence MFE | -27.08 |
| Consensus MFE | -19.99 |
| Energy contribution | -21.93 |
| Covariance contribution | 1.95 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.777697 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14454344 116 + 23771897 UUGUUCCUGUUGUUCGCG-ACGUUGUUGUAUUUGUAUUUUAUUUAAACAAUUUACAAGAAAGCAACAUUCCAGCUGCCUUAAAAUGGAUAAAAGUUGGCACAAGAACGA-GACG--GUGG .....(((.((((((...-..((((((...((((((..((........))..))))))..))))))...((((((.((.......)).....)))))).....))))))-.).)--)... ( -26.00) >DroVir_CAF1 82739 109 + 1 GCUUGGCUGCUGUUUGC---UGUUGUUG----UGUCUUUUAUUUAAACAAUUUACAAGAAAGCAACAUUCCAGCUGCCUUAAAAUGGACAAAAGUUGGCGCAAGAACUA-CGAA--U-AG .........(((((((.---.(((.(((----((((((.((...........)).))))..........((((((.((.......)).....))))))))))).)))..-))))--)-)) ( -22.60) >DroPse_CAF1 73924 120 + 1 UUGUUCUUGUUGUUCUUGGCCGUUGUUGCAUUUGUAUUUUAUUUAAACAAUUUACAAGAAAGCAACAUUCCAGCUGCCUUAAAAUGGACAAAAGUUGGCACAAGAGCCAAGACCAGCCAG .....((.((((.((((((((((((((...((((((..((........))..))))))..))))))...((((((.((.......)).....)))))).....).))))))).)))).)) ( -34.00) >DroEre_CAF1 66894 116 + 1 UUGUUCCUGUUGUUCGCG-ACGUUGUUGUAUUUGUAUUUUAUUUAAACAAUUUACAAGAAAGCAACAUUCCAGCUGCCUUAAAAUGGACAAAAGUUGGCACAGGAACGA-GACA--GAGG ((((((((((........-..((((((...((((((..((........))..))))))..))))))...((((((.((.......)).....)))))).))))))))))-....--.... ( -31.30) >DroAna_CAF1 65018 98 + 1 UUGUUCCUGUUC------------------UUUGUAUUUUAUUUAAACAAUUUACAAGAAAGCAACAUUCCAGCUGCCUUAAAAUGGACAAAAGUUGGCACAGCCACCA-AGCC---ACC .((((.((.(((------------------((.(((..((........))..)))))))))).))))..((((((.((.......)).....))))))...........-....---... ( -14.70) >DroPer_CAF1 74210 120 + 1 UUGUUCUUGUUGUUCUUGGCCGUUGUUUCAUUUGUAUUUUAUUUAAACAAUUUACAAGAAAGCAACAUUCCAGCUGCCUUAAAAUGGACAAAAGUUGGCACAAGAGCCAAGACCAGCCAG .....((.((((.(((((((((((((((..((((((..((........))..)))))).)))))))...((((((.((.......)).....)))))).....).))))))).)))).)) ( -33.90) >consensus UUGUUCCUGUUGUUCGCG__CGUUGUUG_AUUUGUAUUUUAUUUAAACAAUUUACAAGAAAGCAACAUUCCAGCUGCCUUAAAAUGGACAAAAGUUGGCACAAGAACCA_GACC__CAAG ..((((((((...........((((((...((((((..((........))..))))))..))))))...((((((.((.......)).....)))))).))))))))............. (-19.99 = -21.93 + 1.95)


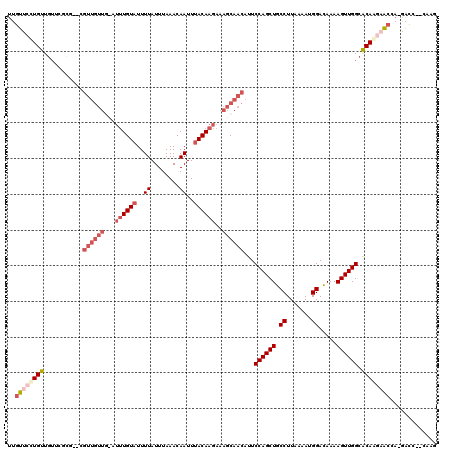
| Location | 14,454,344 – 14,454,460 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.76 |
| Mean single sequence MFE | -24.72 |
| Consensus MFE | -17.17 |
| Energy contribution | -17.95 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.536347 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14454344 116 - 23771897 CCAC--CGUC-UCGUUCUUGUGCCAACUUUUAUCCAUUUUAAGGCAGCUGGAAUGUUGCUUUCUUGUAAAUUGUUUAAAUAAAAUACAAAUACAACAACGU-CGCGAACAACAGGAACAA ....--(((.-.((((.(((((..................((((((((......)))))))).(((((..((((....))))..))))).))))).)))).-.))).............. ( -21.60) >DroVir_CAF1 82739 109 - 1 CU-A--UUCG-UAGUUCUUGCGCCAACUUUUGUCCAUUUUAAGGCAGCUGGAAUGUUGCUUUCUUGUAAAUUGUUUAAAUAAAAGACA----CAACAACA---GCAAACAGCAGCCAAGC ..-.--....-.....((((.((.......((((........))))((((.....(((((...((((....(((((.......)))))----..)))).)---)))).)))).)))))). ( -19.90) >DroPse_CAF1 73924 120 - 1 CUGGCUGGUCUUGGCUCUUGUGCCAACUUUUGUCCAUUUUAAGGCAGCUGGAAUGUUGCUUUCUUGUAAAUUGUUUAAAUAAAAUACAAAUGCAACAACGGCCAAGAACAACAAGAACAA .((..((.((((((((.((((((.................((((((((......)))))))).(((((..((((....))))..)))))..)).)))).)))))))).)).))....... ( -30.50) >DroEre_CAF1 66894 116 - 1 CCUC--UGUC-UCGUUCCUGUGCCAACUUUUGUCCAUUUUAAGGCAGCUGGAAUGUUGCUUUCUUGUAAAUUGUUUAAAUAAAAUACAAAUACAACAACGU-CGCGAACAACAGGAACAA ....--....-..((((((((.((.((..((((.......((((((((......)))))))).(((((..((((....))))..))))).....)))).))-.).)....)))))))).. ( -24.20) >DroAna_CAF1 65018 98 - 1 GGU---GGCU-UGGUGGCUGUGCCAACUUUUGUCCAUUUUAAGGCAGCUGGAAUGUUGCUUUCUUGUAAAUUGUUUAAAUAAAAUACAAA------------------GAACAGGAACAA (((---(((.-.((((((...))).)))...).)))))...(((((((......)))))))((((((...((((...........)))).------------------..)))))).... ( -22.20) >DroPer_CAF1 74210 120 - 1 CUGGCUGGUCUUGGCUCUUGUGCCAACUUUUGUCCAUUUUAAGGCAGCUGGAAUGUUGCUUUCUUGUAAAUUGUUUAAAUAAAAUACAAAUGAAACAACGGCCAAGAACAACAAGAACAA .((..((.((((((((.((((..((...............((((((((......)))))))).(((((..((((....))))..))))).))..)))).)))))))).)).))....... ( -29.90) >consensus CUGC__GGUC_UGGUUCUUGUGCCAACUUUUGUCCAUUUUAAGGCAGCUGGAAUGUUGCUUUCUUGUAAAUUGUUUAAAUAAAAUACAAAU_CAACAACG__CGCGAACAACAGGAACAA .............((((((((...................((((((((......)))))))).(((((..((((....))))..))))).....................)))))))).. (-17.17 = -17.95 + 0.78)


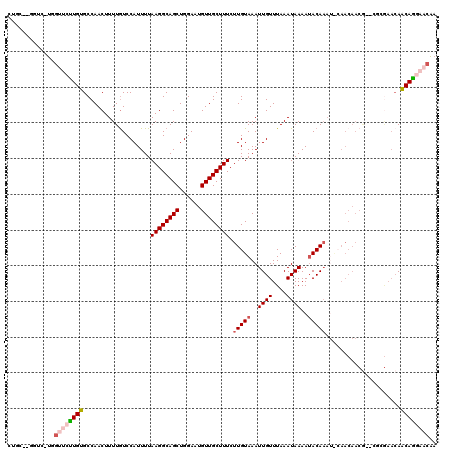
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:13 2006