
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,169,708 – 14,169,800 |
| Length | 92 |
| Max. P | 0.920879 |

| Location | 14,169,708 – 14,169,800 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 82.51 |
| Mean single sequence MFE | -40.10 |
| Consensus MFE | -32.30 |
| Energy contribution | -33.05 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.920879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14169708 92 + 23771897 GUCU--ACGUGGUUGGCCCUCAAUUAGCCUUAAAGGUGGCCGGCUGCCUAAUCGAGGCAGUUUCCAAGCCACUUU---GCGGGGGCCCA--ACAUUGUU----------------- ....--.....((((((((((.....((....((((((((.((((((((.....)))))))).....))))))))---))))))).)))--))......----------------- ( -36.20) >DroSec_CAF1 4363 92 + 1 GUCU--CCGUGGUUGGCCCUCAAUUAGCCUUAAAGGUGGCCGGCUGCCUAAUCGAGGCAGUUUCCAAGCCACUUU---GCGGGGGCCCA-AACAUUGU------------------ ((((--((((((((((.......))))))...((((((((.((((((((.....)))))))).....))))))))---))))))))...-........------------------ ( -37.10) >DroSim_CAF1 4321 92 + 1 GUCU--CCGUGGUUGGCCCUCAAUUAGCCUUAAAGGUGGCCGGCUGCCUAAUCGAGGCAGUUUCCAAGCCACUUU---GCGGGGGCCCA-AACAUUGU------------------ ((((--((((((((((.......))))))...((((((((.((((((((.....)))))))).....))))))))---))))))))...-........------------------ ( -37.10) >DroEre_CAF1 4812 95 + 1 GUCU--CCGCUGUUGGCCCUCAAUUAGCCUUAAAGGUGGCCGGCUGCCUAAUCGAGGCAGUUUCCAAGCCACUUU---GCGGGGGCCCA--ACAUUGUUGG--------------C ((((--((((....(((.........)))...((((((((.((((((((.....)))))))).....))))))))---))))))))(((--((...)))))--------------. ( -41.50) >DroYak_CAF1 4710 109 + 1 GUCU--CCGUGGUUGGCCCUCAAUUAGCCUUAAAGGUGGCCGGCUGCCUAAUCGAGGCAGUUUCCAAGCCACUUU---GCGGGGGCCCA--GCAUUGUUGGCAGUUUGCAGUUGGC ((((--((((((((((.......))))))...((((((((.((((((((.....)))))))).....))))))))---))))))))(((--((((((....)))).....))))). ( -42.50) >DroAna_CAF1 3787 102 + 1 GUGUAGCUGUUGGGGGCCCUCGAUUAGCCUUUUAGGUGGCCGGCUACCUAAUGGCGGCAUUUUCUGAGCCACCUUUAAGCGGCGGCCCCCAACUAACUUGG--------------C ........((((((((((.(((.((((((..((((((((....)))))))).)))(((.........))).....))).))).))))))))))........--------------. ( -46.20) >consensus GUCU__CCGUGGUUGGCCCUCAAUUAGCCUUAAAGGUGGCCGGCUGCCUAAUCGAGGCAGUUUCCAAGCCACUUU___GCGGGGGCCCA__ACAUUGUU_________________ ........((....(((((((.....((....((((((((.((((((((.....)))))))).....))))))))...)))))))))....))....................... (-32.30 = -33.05 + 0.75)
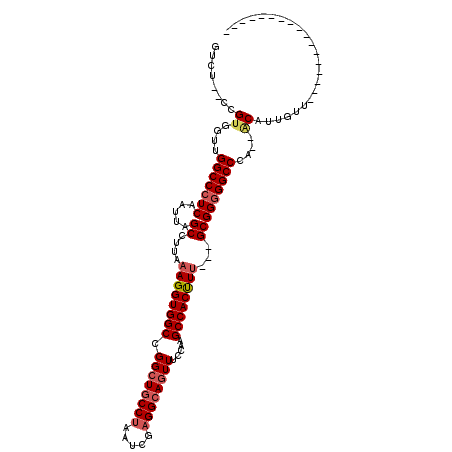



| Location | 14,169,708 – 14,169,800 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 82.51 |
| Mean single sequence MFE | -36.35 |
| Consensus MFE | -26.96 |
| Energy contribution | -27.60 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.807583 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 14169708 92 - 23771897 -----------------AACAAUGU--UGGGCCCCCGC---AAAGUGGCUUGGAAACUGCCUCGAUUAGGCAGCCGGCCACCUUUAAGGCUAAUUGAGGGCCAACCACGU--AGAC -----------------....((((--..(((((((..---(((((((((.((...((((((.....))))))))))))).))))..))........)))))....))))--.... ( -32.10) >DroSec_CAF1 4363 92 - 1 ------------------ACAAUGUU-UGGGCCCCCGC---AAAGUGGCUUGGAAACUGCCUCGAUUAGGCAGCCGGCCACCUUUAAGGCUAAUUGAGGGCCAACCACGG--AGAC ------------------.(..(((.-..(((((((..---(((((((((.((...((((((.....))))))))))))).))))..))........)))))....))).--.).. ( -31.30) >DroSim_CAF1 4321 92 - 1 ------------------ACAAUGUU-UGGGCCCCCGC---AAAGUGGCUUGGAAACUGCCUCGAUUAGGCAGCCGGCCACCUUUAAGGCUAAUUGAGGGCCAACCACGG--AGAC ------------------.(..(((.-..(((((((..---(((((((((.((...((((((.....))))))))))))).))))..))........)))))....))).--.).. ( -31.30) >DroEre_CAF1 4812 95 - 1 G--------------CCAACAAUGU--UGGGCCCCCGC---AAAGUGGCUUGGAAACUGCCUCGAUUAGGCAGCCGGCCACCUUUAAGGCUAAUUGAGGGCCAACAGCGG--AGAC .--------------(((((...))--)))....((((---...((((((.((...((((((.....))))))))))))))......((((.......))))....))))--.... ( -36.70) >DroYak_CAF1 4710 109 - 1 GCCAACUGCAAACUGCCAACAAUGC--UGGGCCCCCGC---AAAGUGGCUUGGAAACUGCCUCGAUUAGGCAGCCGGCCACCUUUAAGGCUAAUUGAGGGCCAACCACGG--AGAC (((.....(((...(((.....(((--.((....))))---)..((((((.((...((((((.....))))))))))))))......)))...)))..))).......(.--...) ( -35.60) >DroAna_CAF1 3787 102 - 1 G--------------CCAAGUUAGUUGGGGGCCGCCGCUUAAAGGUGGCUCAGAAAAUGCCGCCAUUAGGUAGCCGGCCACCUAAAAGGCUAAUCGAGGGCCCCCAACAGCUACAC .--------------...((((.((((((((((((..(((((.((((((.........)))))).)))))..)).((((........)))).......)))))))))))))).... ( -51.10) >consensus _________________AACAAUGU__UGGGCCCCCGC___AAAGUGGCUUGGAAACUGCCUCGAUUAGGCAGCCGGCCACCUUUAAGGCUAAUUGAGGGCCAACCACGG__AGAC .......................((....(((((.(((...((((((((.(((...((((((.....))))))))))))).))))...)).....).)))))....))........ (-26.96 = -27.60 + 0.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:43:18 2006