
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 14,097,512 – 14,097,611 |
| Length | 99 |
| Max. P | 0.998978 |

| Location | 14,097,512 – 14,097,611 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 69.22 |
| Mean single sequence MFE | -24.86 |
| Consensus MFE | -7.17 |
| Energy contribution | -7.69 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.29 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
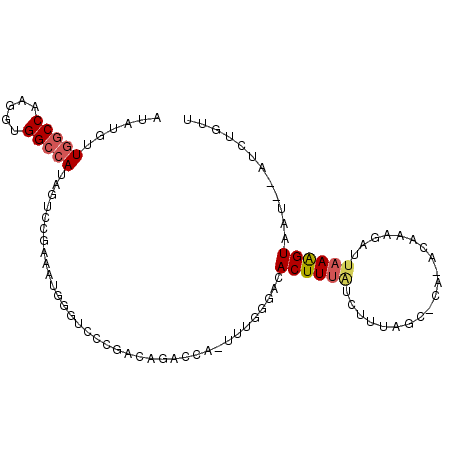
>3L_DroMel_CAF1 14097512 99 + 23771897 AUAUGUUAACCAAGGUGGCCAUAGUCCAAAAUUGGUCCAGACAGACCACUUUGGGACACUUUAUCUUUGGCUCAGACAAAGAUUAAAGUAGU--GUCUGUU .........((((((((((....(.(((....))).)......).)))))))))((((((..(((((((.......)))))))......)))--))).... ( -29.20) >DroSec_CAF1 95670 99 + 1 AUAUGUUGGCCAAGGUGGCCAUAGUCCGAAAUGCGUCUCGAUAGACCAUUUUGGGACACUUUAUCUUUAGCGCACACAAAGAUUAAAGUAAU--AUCUGUU ......(((((.....)))))..((((((((((.((((....)))))))))).))))(((((((((((.(......)))))).))))))...--....... ( -28.40) >DroSim_CAF1 97278 99 + 1 AUAUGUUGGCCAAGGUGGCCAUAGUCCGAAAUGCGUCCCGACAGACCACUUUGGGACACUUUAUCUUUAGCGCACACAAAGAUUAAAGUAAU--AUCUGUU ......(((((.....)))))..((((.(((((.(((......))))).))).))))(((((((((((.(......)))))).))))))...--....... ( -25.80) >DroEre_CAF1 94169 80 + 1 AUAUGUUGGCCAAGGUGGCCAUAAUACGAGC-AGAACGCUUUG----------GAACACUUUAUAUUUCG-----G-----UUGUAGGUUCUAACACAGUU ......(((((.....)))))......((((-.....))))((----------((((....((((.....-----.-----.)))).))))))........ ( -16.30) >DroYak_CAF1 98987 79 + 1 AUAUGUUGGCCAAGGUGGCCAUAAUCCGAAAUGGGUCUCGACA----------GACCACUUUGUUUUUAG-----A-----CUUAAAGUAAU--UUCGGUU ......(((((.....)))))....(((((((.(((((....)----------))))((((((.......-----.-----..)))))).))--))))).. ( -24.60) >consensus AUAUGUUGGCCAAGGUGGCCAUAGUCCGAAAUGGGUCCCGACAGACCA_UUUGGGACACUUUAUCUUUAGC_CA_ACAAAGAUUAAAGUAAU__AUCUGUU ......(((((.....)))))....................................((((((....................))))))............ ( -7.17 = -7.69 + 0.52)


| Location | 14,097,512 – 14,097,611 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 69.22 |
| Mean single sequence MFE | -25.46 |
| Consensus MFE | -10.28 |
| Energy contribution | -9.80 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.40 |
| Mean z-score | -2.81 |
| Structure conservation index | 0.40 |
| SVM decision value | 3.31 |
| SVM RNA-class probability | 0.998978 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
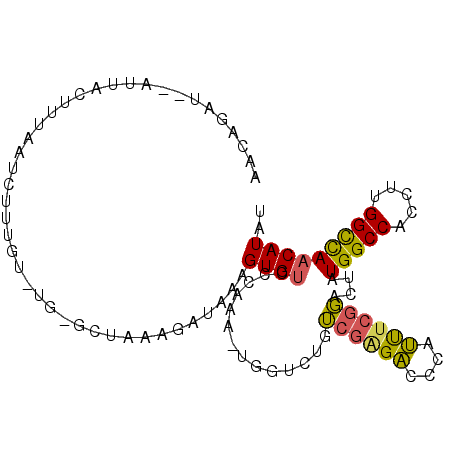
>3L_DroMel_CAF1 14097512 99 - 23771897 AACAGAC--ACUACUUUAAUCUUUGUCUGAGCCAAAGAUAAAGUGUCCCAAAGUGGUCUGUCUGGACCAAUUUUGGACUAUGGCCACCUUGGUUAACAUAU ....(((--(((......(((((((.......)))))))..))))))((((.((((((.(((..((.....))..)))...)))))).))))......... ( -31.30) >DroSec_CAF1 95670 99 - 1 AACAGAU--AUUACUUUAAUCUUUGUGUGCGCUAAAGAUAAAGUGUCCCAAAAUGGUCUAUCGAGACGCAUUUCGGACUAUGGCCACCUUGGCCAACAUAU .......--...((((((.(((((((....)).)))))))))))((((..(((((((((....)))).))))).))))..(((((.....)))))...... ( -29.00) >DroSim_CAF1 97278 99 - 1 AACAGAU--AUUACUUUAAUCUUUGUGUGCGCUAAAGAUAAAGUGUCCCAAAGUGGUCUGUCGGGACGCAUUUCGGACUAUGGCCACCUUGGCCAACAUAU .......--...((((((.(((((((....)).)))))))))))((((..(((((((((....)))).))))).))))..(((((.....)))))...... ( -28.70) >DroEre_CAF1 94169 80 - 1 AACUGUGUUAGAACCUACAA-----C-----CGAAAUAUAAAGUGUUC----------CAAAGCGUUCU-GCUCGUAUUAUGGCCACCUUGGCCAACAUAU ...(((((((((((((..((-----(-----(..........).))).----------...)).)))))-).........(((((.....)))))))))). ( -15.40) >DroYak_CAF1 98987 79 - 1 AACCGAA--AUUACUUUAAG-----U-----CUAAAAACAAAGUGGUC----------UGUCGAGACCCAUUUCGGAUUAUGGCCACCUUGGCCAACAUAU ..(((((--((.(((((..(-----(-----......)))))))((((----------(....))))).)))))))....(((((.....)))))...... ( -22.90) >consensus AACAGAU__AUUACUUUAAUCUUUGU_UG_GCUAAAGAUAAAGUGUCCCAAA_UGGUCUGUCGAGACCCAUUUCGGACUAUGGCCACCUUGGCCAACAUAU ..........................................((((..............((((((....))))))....(((((.....))))))))).. (-10.28 = -9.80 + -0.48)

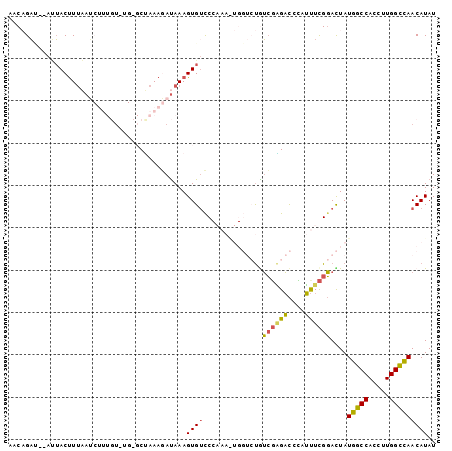
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:42:37 2006