
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,994,232 – 13,994,392 |
| Length | 160 |
| Max. P | 0.999863 |

| Location | 13,994,232 – 13,994,352 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.83 |
| Mean single sequence MFE | -31.64 |
| Consensus MFE | -27.22 |
| Energy contribution | -27.94 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.989939 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13994232 120 + 23771897 AUCUUGGCCAUUUGCGAACUUCGAAAAACGGUGUGCGAUGCGGUGCUUAUCGCGUAGCACAUAACAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUC ...((.((.....)).))....((((((.(((((((.(((((((....))))))).))))))...(((((((((((.........)))))))))))........)))))))......... ( -30.20) >DroSec_CAF1 7651 120 + 1 AUCUUGGCCGUUCGCGAACUUCGAAAAACGGUGUGCGACGCGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUC ((((((((((((..((.....))...)))))(((((.(((((((....))))))).)))))..))(((((((((((.........))))))))))).))))).................. ( -36.60) >DroSim_CAF1 5379 120 + 1 AUCUUGGUCGUUCGCGAACUUCGAAAAACGGUUUGCGAUGCGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUC ((((((((((((((((((((.........))))))))).)))(((((........)))))..)))(((((((((((.........))))))))))).))))).................. ( -33.80) >DroEre_CAF1 6430 119 + 1 AUCUUAGCCGUUCGCGAACUUCGGAAAACGGUGGGCGAUGCGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUACAACAGAAAAAUUGCAUUUAUAGAUCACUUUUU-UCUUUUAUC ......((.....)).......((((((.(((((((.(((((((....))))))).)).......(((((((((((.........))))))))))).....))))).)))-)))...... ( -28.90) >DroYak_CAF1 6995 120 + 1 AUCUUAGCCGUUCGCGAACUUCGGAAAACGGUGGGCGAUGAGGUGCUUAUCGCGUAGCACAUACCAGAUGCAAUUUACAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUC (((((.(((((((.((.....))))..)))))((((((((((...))))))))(.....)...))(((((((((((.........))))))))))).))))).................. ( -28.70) >consensus AUCUUGGCCGUUCGCGAACUUCGAAAAACGGUGUGCGAUGCGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUC (((((.((((((..((.....))...))))))((((.(((((((....))))))).)))).....(((((((((((.........))))))))))).))))).................. (-27.22 = -27.94 + 0.72)



| Location | 13,994,272 – 13,994,392 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.67 |
| Mean single sequence MFE | -31.00 |
| Consensus MFE | -27.98 |
| Energy contribution | -29.02 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.90 |
| SVM decision value | 4.29 |
| SVM RNA-class probability | 0.999863 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13994272 120 + 23771897 CGGUGCUUAUCGCGUAGCACAUAACAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUCACCGCUGCGCCUUCCGCCUAGAGUUGGUACAAAACACUGU (((((......(((((((.......(((((((((((.........)))))))))))..(((........)))...........))))))).....(((.......)))......))))). ( -30.10) >DroSec_CAF1 7691 120 + 1 CGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUCACCGCUGCGCCUUCCGCCUAGAGUUGGAACAAAACUCUGU .((((......(((((((.......(((((((((((.........)))))))))))..(((........)))...........)))))))....))))(((((((.......))))))). ( -33.50) >DroSim_CAF1 5419 120 + 1 CGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUCACCGCUGCGCCUUCCGCCUAGAGUUGGAACAAAACUCUGU .((((......(((((((.......(((((((((((.........)))))))))))..(((........)))...........)))))))....))))(((((((.......))))))). ( -33.50) >DroEre_CAF1 6470 119 + 1 CGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUACAACAGAAAAAUUGCAUUUAUAGAUCACUUUUU-UCUUUUAUCACCGCUCCGCCUUCCGCAUAGAGUUGGAACAAAACUCAGU ..((((.....(((.(((.......(((((((((((.........)))))))))))..(((.........-))).........))).))).....)))).(((((.......)))))... ( -25.50) >DroYak_CAF1 7035 120 + 1 AGGUGCUUAUCGCGUAGCACAUACCAGAUGCAAUUUACAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUCACCGCUGCGCCUUCCGCCUAGAGUGGAAACAAAACUCUGU (((((......(((((((.......(((((((((((.........)))))))))))..(((........)))...........)))))))....)))))((((((....)...))))).. ( -32.40) >consensus CGGUGCUUAUCGCGUAGCACAUACCAAAUGCAAUUUGGAACCGAAAAAUUGCAUUUAAAGAUCACUUUUUCUCUUUUAUCACCGCUGCGCCUUCCGCCUAGAGUUGGAACAAAACUCUGU .((((......(((((((.......(((((((((((.........)))))))))))(((((..........))))).......)))))))....))))(((((((.......))))))). (-27.98 = -29.02 + 1.04)


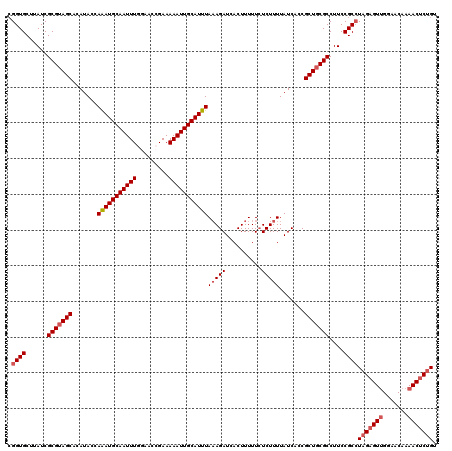
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:42:04 2006