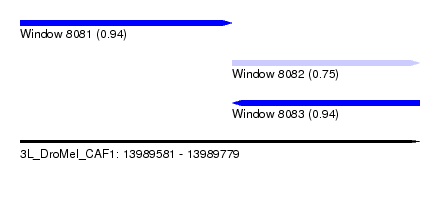

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,989,581 – 13,989,779 |
| Length | 198 |
| Max. P | 0.942238 |

| Location | 13,989,581 – 13,989,686 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 89.78 |
| Mean single sequence MFE | -42.98 |
| Consensus MFE | -39.03 |
| Energy contribution | -38.87 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.942238 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
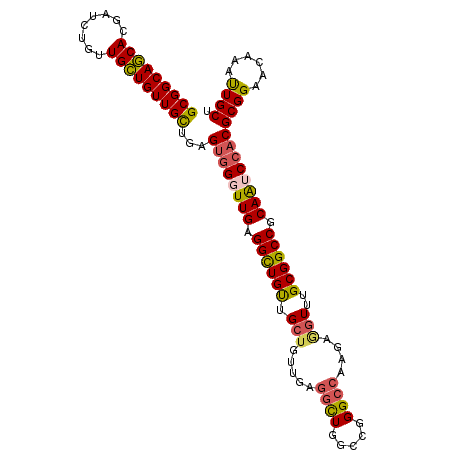
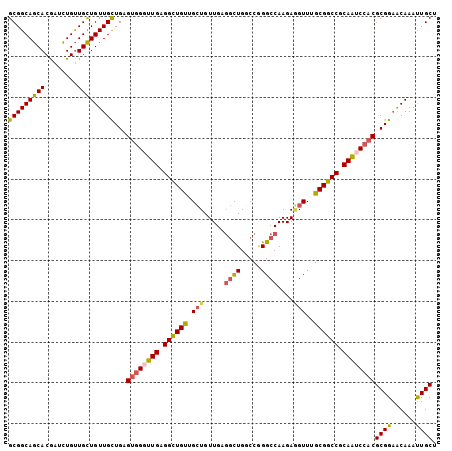
>3L_DroMel_CAF1 13989581 105 + 23771897 GCGGCAGCACGAUCUGUUGCUGUUGUUGAGUGGGUUGAGGCUGUUGCUGUUGAGGCUGGCCGGGCCAAGAGGUUUGCGGCCGCAAUCCACGCGGAACAAAUUGCU (((((((((.....)))))))))((((..((((((((.((((((.(((.....((((.....))))....)))..)))))).))))))))....))))....... ( -43.20) >DroPse_CAF1 3157 102 + 1 GCGGCAACACAAGUUGCUGUUGUUGCUGCGGAGCUUGAGGCUGCUGAUGUUG---CUGUCCUGGCCAAGAUGUUUGCGGCCGCAGUCCACGCGGUGCAAACUGCU (((((((((((......)).)))))))))(((..(((.((((((.((((((.---..((....))...)))))).)))))).))))))..(((((....))))). ( -42.60) >DroSec_CAF1 3094 105 + 1 GCGGCAGCACGAUCUGUUGCUGUUGUUGAGUGGGUUGAGGCUGUUGCUGUUGAGGCUGACCGGGCCAAGAGGUUUGCGGCCGCAAUCCACGCGGAACAAAUUGCU (((((((((.....)))))))))((((..((((((((.((((((.(((.....((((.....))))....)))..)))))).))))))))....))))....... ( -43.20) >DroSim_CAF1 3377 105 + 1 GCGGCAGCACGAUCUGUUGCUGUUGUUGAGUGGGUUGAGGCUGUUGCUGUUGAGGCUGGCCGGGCCAAGAGGUUUGCGGCCGCAAUCCACGCGGAACAAAUUGCU (((((((((.....)))))))))((((..((((((((.((((((.(((.....((((.....))))....)))..)))))).))))))))....))))....... ( -43.20) >DroEre_CAF1 3126 105 + 1 GCGGCAGCACGAUCUGUUGCUGUUGCUGGGUGGGUUGCGGUUGUUGCUGUUGAGGUUGGCCAGGCCAAGAGGUUUGCGGCCGCAAACCACGCGGAACAAAUUGCU (((((((((.....)))))))(((.(((.((((.((((((((((.(((.....((((.....))))....)))..)))))))))).)))).)))))).....)). ( -44.60) >DroYak_CAF1 3130 105 + 1 GCGGCAGCACGAUCUGUUGCUGUUGCUGAGUGGGUUGAGGUUGUUGCUGUUGAGGUUGGCCGGGCCAAGAAGUUUGCGGCCGCAAUCCACGCGGAACAAAUUGCU (((((((((.....)))))))(((.(((.((((((((.((((((.(((.......(((((...)))))..)))..)))))).)))))))).)))))).....)). ( -41.10) >consensus GCGGCAGCACGAUCUGUUGCUGUUGCUGAGUGGGUUGAGGCUGUUGCUGUUGAGGCUGGCCGGGCCAAGAGGUUUGCGGCCGCAAUCCACGCGGAACAAAUUGCU (((((((((........)))))))))...((((((((.((((((.(((.....((((.....))))....)))..)))))).))))))))((((......)))). (-39.03 = -38.87 + -0.16)

| Location | 13,989,686 – 13,989,779 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 90.32 |
| Mean single sequence MFE | -40.80 |
| Consensus MFE | -33.42 |
| Energy contribution | -33.74 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751768 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
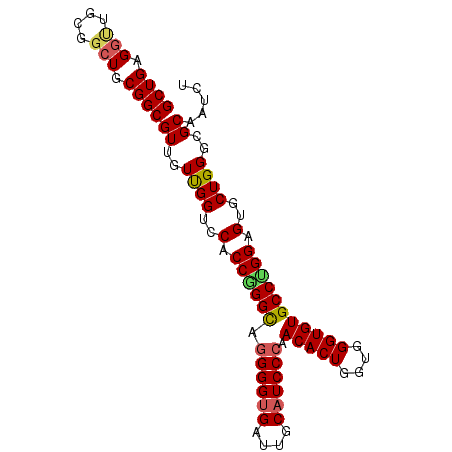
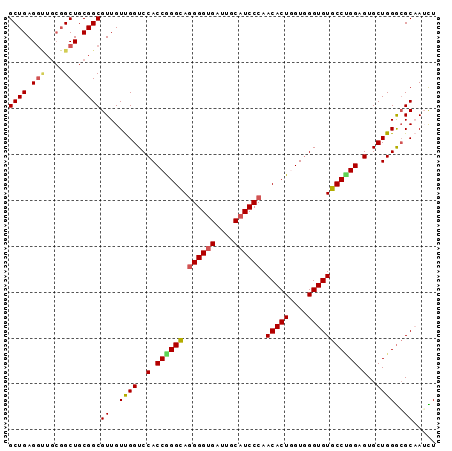
>3L_DroMel_CAF1 13989686 93 + 23771897 GCUGAGGCUGCGGCUGCGGCGUUGUUGGUCCACCGGGCAGGGGUGAUUGCAUCCCAACACUGGUGGGUGUGCCUGGAGUGCUGGACGCCAUUU ((((......))))...(((((..(..(..(.((((((.((((((....)))))).(((((....))))))))))).)..)..)))))).... ( -42.10) >DroSec_CAF1 3199 93 + 1 GCUGAGGUUGCGGCUGCGGCGUUGUUGGUCCACCAGGCAGGGGUGAUUGCAUCCCAACACUGGUGGGUGUGCCUGGAGUGCUGGGCGCAAGCU (((((.(((((....))))).)).....(((((((((..((((((....))))))..).))))))))((((((..(....)..))))))))). ( -43.90) >DroSim_CAF1 3482 93 + 1 GCUGAGGUUGCGGCUGCGGCGUUGUUGGUCCACCGGGCAGGGGUGAUUGCAUCCCAACACUGGUGGGUGUGCCUGGAGUGCUGGGCGCAAGCU (((((.(((((....))))).)).....(((((((((..((((((....))))))..).))))))))((((((..(....)..))))))))). ( -43.20) >DroEre_CAF1 3231 87 + 1 GCUGAG------GCUGCGGCGUUGUUGGUCCACCGGGUAGGGGUGAUUGCAUCCCAACACUAGUUGGUGUGCCCGGAGUGCUGGGCGCAAUCU ((((..------....))))...(((((((((((((((.((((((....)))))).(((((....))))))))))).....))))).)))).. ( -36.90) >DroYak_CAF1 3235 93 + 1 GCUGAGGUUGCGGCUGCGGCGUUGUCGGUCCACCGGGCAAGGGGGAUUGCAUCCCAACACUAGUGGGUGUGCCAGGAGUGCUGGGAGCCAUUU ((((......))))...(((..((.((((((.((......)).))))))))(((((.((((..(((.....)))..)))).)))))))).... ( -37.90) >consensus GCUGAGGUUGCGGCUGCGGCGUUGUUGGUCCACCGGGCAGGGGUGAUUGCAUCCCAACACUGGUGGGUGUGCCUGGAGUGCUGGGCGCAAUCU ((((.(((....))).))))((..((((..(.((((((.((((((....)))))).(((((....))))))))))).)..))))..))..... (-33.42 = -33.74 + 0.32)

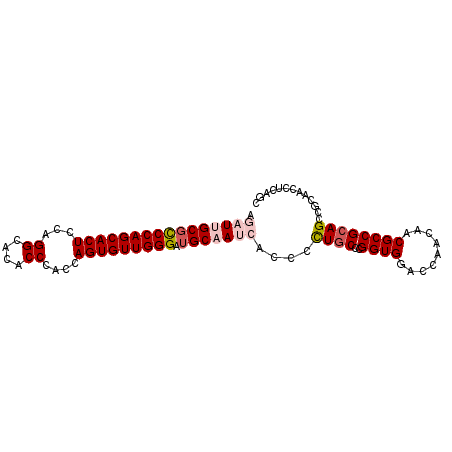
| Location | 13,989,686 – 13,989,779 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 90.32 |
| Mean single sequence MFE | -31.14 |
| Consensus MFE | -25.20 |
| Energy contribution | -26.48 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13989686 93 - 23771897 AAAUGGCGUCCAGCACUCCAGGCACACCCACCAGUGUUGGGAUGCAAUCACCCCUGCCCGGUGGACCAACAACGCCGCAGCCGCAGCCUCAGC ....(((((...(((..((((.(((........)))))))..)))..(((((.......))))).......)))))((....))......... ( -27.50) >DroSec_CAF1 3199 93 - 1 AGCUUGCGCCCAGCACUCCAGGCACACCCACCAGUGUUGGGAUGCAAUCACCCCUGCCUGGUGGACCAACAACGCCGCAGCCGCAACCUCAGC .((((((((((((((((...((.....))...))))))))).)))))......((((..((((.........))))))))..))......... ( -31.50) >DroSim_CAF1 3482 93 - 1 AGCUUGCGCCCAGCACUCCAGGCACACCCACCAGUGUUGGGAUGCAAUCACCCCUGCCCGGUGGACCAACAACGCCGCAGCCGCAACCUCAGC .((((((((((((((((...((.....))...))))))))).)))))......((((..((((.........))))))))..))......... ( -31.50) >DroEre_CAF1 3231 87 - 1 AGAUUGCGCCCAGCACUCCGGGCACACCAACUAGUGUUGGGAUGCAAUCACCCCUACCCGGUGGACCAACAACGCCGCAGC------CUCAGC .((((((((((((((((...((....))....))))))))).))))))).........(((((.........)))))....------...... ( -32.90) >DroYak_CAF1 3235 93 - 1 AAAUGGCUCCCAGCACUCCUGGCACACCCACUAGUGUUGGGAUGCAAUCCCCCUUGCCCGGUGGACCGACAACGCCGCAGCCGCAACCUCAGC ....(((((((((((((..(((.....)))..)))))))))).((((......)))).(((((.........)))))..)))........... ( -32.30) >consensus AGAUUGCGCCCAGCACUCCAGGCACACCCACCAGUGUUGGGAUGCAAUCACCCCUGCCCGGUGGACCAACAACGCCGCAGCCGCAACCUCAGC .((((((((((((((((...((....))....))))))))).)))))))....((((..((((.........))))))))............. (-25.20 = -26.48 + 1.28)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:42:01 2006