
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,928,913 – 13,929,029 |
| Length | 116 |
| Max. P | 0.795789 |

| Location | 13,928,913 – 13,929,029 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 81.49 |
| Mean single sequence MFE | -41.42 |
| Consensus MFE | -32.74 |
| Energy contribution | -34.33 |
| Covariance contribution | 1.59 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.572066 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
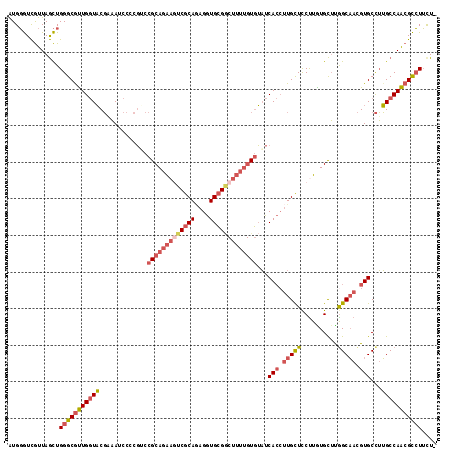
>3L_DroMel_CAF1 13928913 116 + 23771897 AUGGGUCGUUAGCUGGGCAUUGGUACGAAAUCCCCGACCGCAGAAAUCGCAGAGGUGCGGCUUUUGUGUAUCACCUUGCUCCUUGUGCUUGGCAACGUGCCUUGCCAACGCCUUUU- ..(((.((((.((.((((((.(((.((.......)))))(((.((...(((.(((((..((......))..))))))))...)).)))..(....))))))).)).)))))))...- ( -38.60) >DroSec_CAF1 37164 116 + 1 AUGGGUCGUUAGCUGGGCGUUGGUACGAAAUCCUCGUCCGCAGAAGUCGCAGAGGUGCGGCUUUUGUGUAUCACCUUGCUCCUUGUGCUUGGCAACGUGCCUUGCCAACGCCUUCU- ..(((.((((.((.((((((((..((((.....)))).(((((((((((((....))))))))))))).....((..((.......))..))))))..)))).)).)))))))...- ( -43.30) >DroSim_CAF1 37519 116 + 1 AUGGGUCGUUAGCUGGGCGUUGGUACGAAAUCCGCGUCCGCAGAAGUCGCAGAGGUGCGGCUUUUGUGUAUCACCUUGCUCCUUGUGCUUGGCAACGUGCCUUGCCAACGCCUUCU- ..............(((((((((((.(.....(((((.(((((((((((((....))))))))))))).....((..((.......))..))..))))).).)))))))))))...- ( -44.10) >DroEre_CAF1 36471 116 + 1 GUGGGUCGUUAGCUGGGCGUUGGUACGAAAUCCCCGUCCGCAGAAGUCGCAGAGGUGCGUCUUUUGUGUAUCACCUUGCUUCCUAUGCUUGGCAACGUGCCUUGCCAACGCCUUUU- ..............(((((((((((.(...........((((((((.((((....)))).))))))))...(((.(((((..........))))).))).).)))))))))))...- ( -40.20) >DroYak_CAF1 38667 116 + 1 AUGGGUCGUUAGCUGGGCGUUGGUACGGAAUCCCCGACCGCAGAAGUCGCAUAGGUGCGGCUUUUGUGUAUCACCUUGCUCCUUAUGCUUAGCAACGUGCCUUGCCAACGCCUUUU- ..............(((((((((((.((..........(((((((((((((....)))))))))))))...(((.(((((..........))))).))))).)))))))))))...- ( -46.30) >DroPer_CAF1 38209 111 + 1 GUGUAUUAUUAAUUGGGCGCUGCUUCUGUAGCCCUUGCCGCAGGUCUCGCAAAGAUGGGA------UCGCUCAAACUGUUUCUUGUGUGCGGCCAUGUGCUGGGCCAGCGUGUACUU ..............(((.(((((....))))))))((((((.(((((.(((...((((..------((((....((........))..)))))))).))).))))).))).)))... ( -36.00) >consensus AUGGGUCGUUAGCUGGGCGUUGGUACGAAAUCCCCGUCCGCAGAAGUCGCAGAGGUGCGGCUUUUGUGUAUCACCUUGCUCCUUGUGCUUGGCAACGUGCCUUGCCAACGCCUUCU_ ..............(((((((((((.............(((((((((((((....)))))))))))))...(((.(((((..........))))).)))...))))))))))).... (-32.74 = -34.33 + 1.59)


| Location | 13,928,913 – 13,929,029 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 81.49 |
| Mean single sequence MFE | -38.62 |
| Consensus MFE | -27.42 |
| Energy contribution | -28.37 |
| Covariance contribution | 0.95 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.795789 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
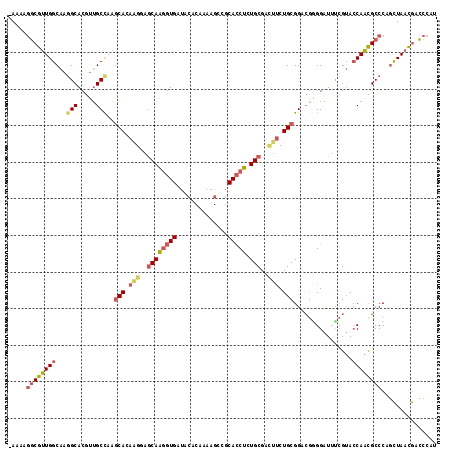
>3L_DroMel_CAF1 13928913 116 - 23771897 -AAAAGGCGUUGGCAAGGCACGUUGCCAAGCACAAGGAGCAAGGUGAUACACAAAAGCCGCACCUCUGCGAUUUCUGCGGUCGGGGAUUUCGUACCAAUGCCCAGCUAACGACCCAU -....(((((((((..(((.....)))........((.((((((((...(......)...))))).(((((..(((.(....).)))..)))))....))))).))))))).))... ( -38.70) >DroSec_CAF1 37164 116 - 1 -AGAAGGCGUUGGCAAGGCACGUUGCCAAGCACAAGGAGCAAGGUGAUACACAAAAGCCGCACCUCUGCGACUUCUGCGGACGAGGAUUUCGUACCAACGCCCAGCUAACGACCCAU -....(((((((((..(((..........(((.(((..((((((((...(......)...))))).)))..))).)))(((((((...))))).))...)))..))))))).))... ( -42.00) >DroSim_CAF1 37519 116 - 1 -AGAAGGCGUUGGCAAGGCACGUUGCCAAGCACAAGGAGCAAGGUGAUACACAAAAGCCGCACCUCUGCGACUUCUGCGGACGCGGAUUUCGUACCAACGCCCAGCUAACGACCCAU -....(((((((((..(((.....)))........((.((.(((((...(......)...))))).(((((..(((((....)))))..))))).....)))).))))))).))... ( -41.00) >DroEre_CAF1 36471 116 - 1 -AAAAGGCGUUGGCAAGGCACGUUGCCAAGCAUAGGAAGCAAGGUGAUACACAAAAGACGCACCUCUGCGACUUCUGCGGACGGGGAUUUCGUACCAACGCCCAGCUAACGACCCAC -....(((((((((..(((..........(((.(((..((((((((...(......)...))))).)))..))).)))((((((.....)))).))...)))..))))))).))... ( -39.30) >DroYak_CAF1 38667 116 - 1 -AAAAGGCGUUGGCAAGGCACGUUGCUAAGCAUAAGGAGCAAGGUGAUACACAAAAGCCGCACCUAUGCGACUUCUGCGGUCGGGGAUUCCGUACCAACGCCCAGCUAACGACCCAU -....(((((((((..(((..........(((.(((..((((((((...(......)...))))).)))..))).)))((((((.....))).)))...)))..))))))).))... ( -44.10) >DroPer_CAF1 38209 111 - 1 AAGUACACGCUGGCCCAGCACAUGGCCGCACACAAGAAACAGUUUGAGCGA------UCCCAUCUUUGCGAGACCUGCGGCAAGGGCUACAGAAGCAGCGCCCAAUUAAUAAUACAC .......(((((((((..(....)((((((..((((......)))).((((------........))))......))))))..))))........)))))................. ( -26.60) >consensus _AAAAGGCGUUGGCAAGGCACGUUGCCAAGCACAAGGAGCAAGGUGAUACACAAAAGCCGCACCUCUGCGACUUCUGCGGACGGGGAUUUCGUACCAACGCCCAGCUAACGACCCAU .....((((((((...(((.....)))..(((.(((..((((((((..............))))).)))..))).)))................))))))))............... (-27.42 = -28.37 + 0.95)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:41:23 2006