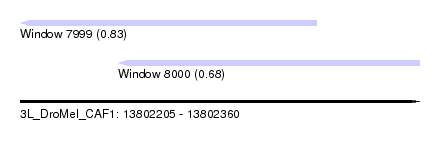

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,802,205 – 13,802,360 |
| Length | 155 |
| Max. P | 0.833066 |

| Location | 13,802,205 – 13,802,320 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 87.39 |
| Mean single sequence MFE | -21.46 |
| Consensus MFE | -17.04 |
| Energy contribution | -17.23 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.833066 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
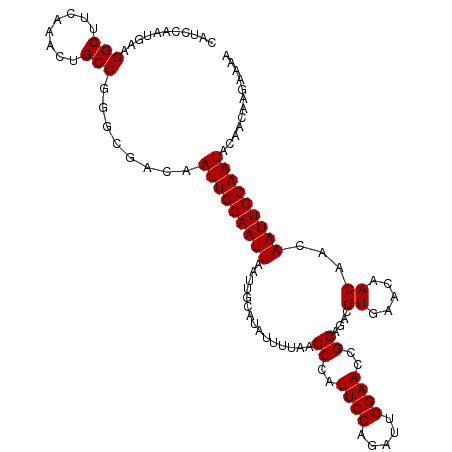
>3L_DroMel_CAF1 13802205 115 - 23771897 CAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCGCAGACGUGAACAACAAAAAUUGCAAUACAACAAGAAAA ..((((((...(((........))).(((......(((((....)))))........))).......))))))...((((...((.....))....))))............... ( -22.94) >DroSec_CAF1 39046 115 - 1 CAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCGCAGACGUGAACAACAACAAUUGCAAUACAACAAGAAAA ..((((((...(((........))).(((......(((((....)))))........))).......))))))...((((..((........))..))))............... ( -23.94) >DroEre_CAF1 39796 115 - 1 CAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCGCAGACGUGAACAACAACAAUUGCAAUACAACAAGAAAA ..((((((...(((........))).(((......(((((....)))))........))).......))))))...((((..((........))..))))............... ( -23.94) >DroYak_CAF1 37860 87 - 1 ----------------------------CGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCGCAGACGUGAACAACAACAAUUGCAAUACAACAAGAAAA ----------------------------.....(((((((((...............((..((((.....))))..))....((.....))...)))))))))............ ( -15.00) >consensus CAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCGCAGACGUGAACAACAACAAUUGCAAUACAACAAGAAAA ...........(((........)))........(((((((((...............((..((((.....))))..))....((.....))...)))))))))............ (-17.04 = -17.23 + 0.19)


| Location | 13,802,243 – 13,802,360 |
|---|---|
| Length | 117 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 99.43 |
| Mean single sequence MFE | -31.97 |
| Consensus MFE | -30.93 |
| Energy contribution | -31.27 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.681526 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
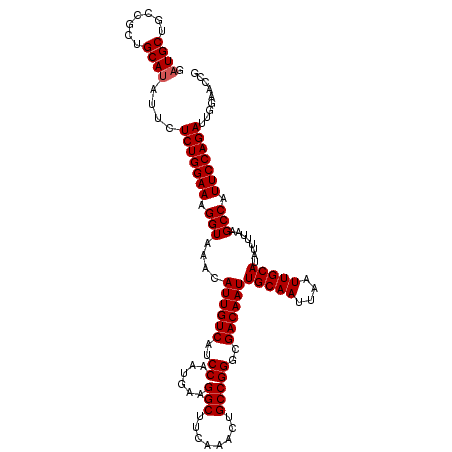
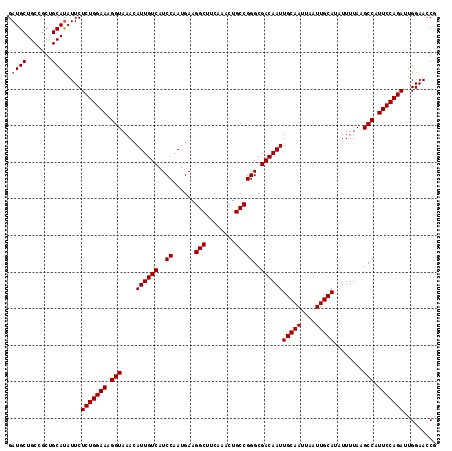
>3L_DroMel_CAF1 13802243 117 - 23771897 GAUGCUGCCGCUGCACAUUCUCUGGAAAGGUAAACAUUGUCAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCG ((((.(((....))))))).(((((((.(((....((((((..((......(((........)))))..))))))(((((....)))))........))).)))))))......... ( -31.90) >DroSec_CAF1 39084 117 - 1 GAUGCUGCCGCUGCAUAUUCUCUGGAAAGGUAAACAUUGUCAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCG .((((.......))))....(((((((.(((....((((((..((......(((........)))))..))))))(((((....)))))........))).)))))))......... ( -32.00) >DroEre_CAF1 39834 117 - 1 GAUGCUGCCGCUGCAUAUUCUCUGGAAAGGUAAACAUUGUCAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCG .((((.......))))....(((((((.(((....((((((..((......(((........)))))..))))))(((((....)))))........))).)))))))......... ( -32.00) >consensus GAUGCUGCCGCUGCAUAUUCUCUGGAAAGGUAAACAUUGUCAUCCAAUGAAGGCUUCAAACUGCCGGGCGACAAUUGCAAUUAAUUGCAUAUUUUAAGCCAUUCCAGAUUGGAACCG .((((.......))))....(((((((.(((....((((((..((......(((........)))))..))))))(((((....)))))........))).)))))))......... (-30.93 = -31.27 + 0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:40 2006