
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,791,488 – 13,791,621 |
| Length | 133 |
| Max. P | 0.925654 |

| Location | 13,791,488 – 13,791,586 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 90.92 |
| Mean single sequence MFE | -19.68 |
| Consensus MFE | -18.62 |
| Energy contribution | -18.62 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.842411 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13791488 98 + 23771897 AAGAAUUAGUAUUCAUGCCCCACAGUGUCAUUUCCGCGAAUGCGUGCGGGCGCAGAAAACUGGCAAAAAA--AA----------GAACAGAAAUGGAAAAAAGAAAAACU .....(((((.(((.((((((.((.((.(((((....))))))))).))).)))))).))))).......--..----------...((....))............... ( -20.00) >DroSec_CAF1 28532 100 + 1 AAGAAUUAGUAUUCAUGCCCCACAGUGUCAUUUCCGAGAAUGCGUGCGGGCGCAGAAGACUGGCAAAAAAAAAA----------AAACACAAAUCGAAAAAAGAAAAACU .....(((((.(((.((((((.((.((.(((((....))))))))).))).)))))).)))))...........----------.......................... ( -18.70) >DroSim_CAF1 24696 98 + 1 AAGAAUUAGUAUUCAUGCCCCACAGUGUCAUUUCCGCGAAUGCGUGCGGGCGCAGAAGACUGGCAAAAAA--AA----------GAACAGAAAUGGAAAAAAGAAAAACU .....(((((.(((.((((((.((.((.(((((....))))))))).))).)))))).))))).......--..----------...((....))............... ( -20.00) >DroEre_CAF1 27976 110 + 1 AAGAAUUAGUAUUCAUGCCCCACAGUGUCAUUUCCGCGAAUGCGUGCGGGCGCAGAAGACUGGCAAAAAAGAAAAAAAAGUAAAGCACAGAAAUGGAAAAAAGAAAAACU .....(((((.(((.((((((.((.((.(((((....))))))))).))).)))))).)))))........................((....))............... ( -20.00) >consensus AAGAAUUAGUAUUCAUGCCCCACAGUGUCAUUUCCGCGAAUGCGUGCGGGCGCAGAAGACUGGCAAAAAA__AA__________GAACAGAAAUGGAAAAAAGAAAAACU .....(((((.(((.((((((.((.((.(((((....))))))))).))).)))))).)))))............................................... (-18.62 = -18.62 + 0.00)


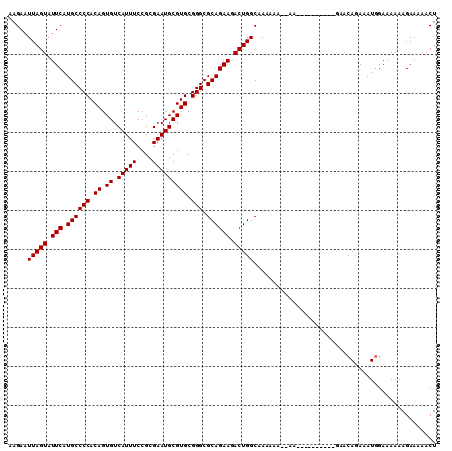
| Location | 13,791,528 – 13,791,621 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 90.47 |
| Mean single sequence MFE | -20.07 |
| Consensus MFE | -17.48 |
| Energy contribution | -17.48 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13791528 93 + 23771897 UGCGUGCGGGCGCAGAAAACUGGCAAAAAA--AA----------GAACAGAAAUGGAAAAAAGAAAAACUUUUUGCAAGCAUUUCUUGAUGGGAAUGGCGUGAGA .((((((..((.(((....)))))......--..----------...((....))..((((((.....))))))))...(((((((....))))))))))).... ( -18.70) >DroSec_CAF1 28572 95 + 1 UGCGUGCGGGCGCAGAAGACUGGCAAAAAAAAAA----------AAACACAAAUCGAAAAAAGAAAAACUUUUUGCAAGCAUUUCUUGAUGGGAAUGGCGUGAGA .((((((..((.(((....)))))..........----------.............((((((.....))))))))...(((((((....))))))))))).... ( -17.70) >DroSim_CAF1 24736 93 + 1 UGCGUGCGGGCGCAGAAGACUGGCAAAAAA--AA----------GAACAGAAAUGGAAAAAAGAAAAACUUUUUGCAAGCAUUUCUUGAUGGGAAUGGCGUGAGA .((((((..((.(((....)))))......--..----------...((....))..((((((.....))))))))...(((((((....))))))))))).... ( -18.70) >DroEre_CAF1 28016 105 + 1 UGCGUGCGGGCGCAGAAGACUGGCAAAAAAGAAAAAAAAGUAAAGCACAGAAAUGGAAAAAAGAAAAACUUUUUGCCAGCAUUUCUUGAUGGGAAUGGCGUGAGA (((((....))))).....((((((((((..........(.....).((....))..............))))))))))(((((((....))))))).(....). ( -25.20) >consensus UGCGUGCGGGCGCAGAAGACUGGCAAAAAA__AA__________GAACAGAAAUGGAAAAAAGAAAAACUUUUUGCAAGCAUUUCUUGAUGGGAAUGGCGUGAGA .((((((..((.(((....))))).................................((((((.....))))))))...(((((((....))))))))))).... (-17.48 = -17.48 + -0.00)


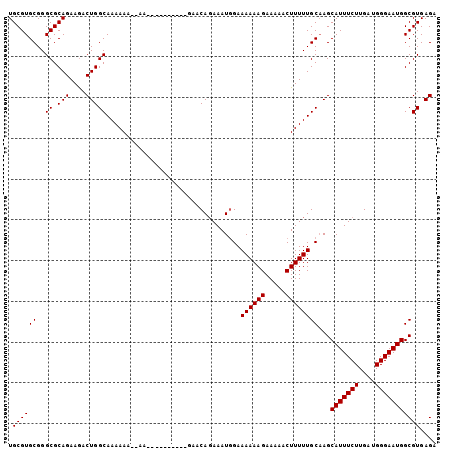
| Location | 13,791,528 – 13,791,621 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 90.47 |
| Mean single sequence MFE | -14.82 |
| Consensus MFE | -11.62 |
| Energy contribution | -11.88 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.610439 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13791528 93 - 23771897 UCUCACGCCAUUCCCAUCAAGAAAUGCUUGCAAAAAGUUUUUCUUUUUUCCAUUUCUGUUC----------UU--UUUUUUGCCAGUUUUCUGCGCCCGCACGCA ......((...........(((((.(((.((((((((........................----------..--)))))))).))))))))((....))..)). ( -13.77) >DroSec_CAF1 28572 95 - 1 UCUCACGCCAUUCCCAUCAAGAAAUGCUUGCAAAAAGUUUUUCUUUUUUCGAUUUGUGUUU----------UUUUUUUUUUGCCAGUCUUCUGCGCCCGCACGCA ....((((...((.....((((((.((((.....)))).)))))).....))...))))..----------..........(((((....))).))......... ( -13.80) >DroSim_CAF1 24736 93 - 1 UCUCACGCCAUUCCCAUCAAGAAAUGCUUGCAAAAAGUUUUUCUUUUUUCCAUUUCUGUUC----------UU--UUUUUUGCCAGUCUUCUGCGCCCGCACGCA ......((...........(((((((...(.((((((.....)))))).))))))))....----------..--......(((((....))).))..))..... ( -13.00) >DroEre_CAF1 28016 105 - 1 UCUCACGCCAUUCCCAUCAAGAAAUGCUGGCAAAAAGUUUUUCUUUUUUCCAUUUCUGUGCUUUACUUUUUUUUCUUUUUUGCCAGUCUUCUGCGCCCGCACGCA ......((...........((((..((((((((((((.............(((....)))...............)))))))))))).))))((....))..)). ( -18.69) >consensus UCUCACGCCAUUCCCAUCAAGAAAUGCUUGCAAAAAGUUUUUCUUUUUUCCAUUUCUGUUC__________UU__UUUUUUGCCAGUCUUCUGCGCCCGCACGCA ........................(((.(((((((((.....)))))).................................(((((....))).))..))).))) (-11.62 = -11.88 + 0.25)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:07 2006