
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,775,508 – 13,775,725 |
| Length | 217 |
| Max. P | 0.866534 |

| Location | 13,775,508 – 13,775,619 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 94.43 |
| Mean single sequence MFE | -29.26 |
| Consensus MFE | -29.48 |
| Energy contribution | -28.44 |
| Covariance contribution | -1.04 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.05 |
| Structure conservation index | 1.01 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.694948 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13775508 111 + 23771897 AAGGUUGGAAAUAAUCAAUAAUGAAAUGAAGGAGUUUGCCGAAACUUUGGCAAUAAGUGGUUUUGGGCUUGGCUUCCCUCG-GGCAGACUCCGUAGGUUUUGCUUAAUUUUG ..(((((....)))))......(((((...((((((((((((((((...((.....))))))))(((........)))...-)))))))))).((((.....))))))))). ( -28.10) >DroSec_CAF1 22129 108 + 1 AAGGUUGGAAA---UCAAUAAUGAAAUGAAGGAGUUUGCCGAAACUUUGGCAAUAAGUGGUUUUGGGCUUGGCUUCCCUCG-GGCAGACUCCGUAGGUUUUGCUUAAUUUUG ...((((....---.))))...(((((...((((((((((((((((...((.....))))))))(((........)))...-)))))))))).((((.....))))))))). ( -26.70) >DroSim_CAF1 17797 111 + 1 AAGGUUGGAAAUAAUCAAUAAUGAAAUGAAGGAGUUUGCCGAAACUUUGGCAAUAAGUGGUUUUGGGCUUGACUUCCCUCG-GGCAGACUCCGUAGGUUUUGCUUAAUUUUG ..(((((....)))))......(((((...((((((((((((((((...((.....))))))))(((........)))...-)))))))))).((((.....))))))))). ( -28.10) >DroEre_CAF1 21567 111 + 1 AAGGUUGAAAAUAAUCAAUAAUGGAAUGAAGGAGUUUGCCGAAACUUUGGCAAUAAGUGGUUUCGGGCUUGGUUUCCCUCG-GGCAGACUCUGAAGGUUUCGCUUAAUUUGG ((((((((......)))))...(((((...((((((((((((((((...((.....))))))))(((........)))...-))))))))))....))))).)))....... ( -30.50) >DroYak_CAF1 20302 112 + 1 AAGGUUGAAAAUAAUCAAUAAUGAAAUGAAGGAGUUUGCCGAAACUUUGGCAAUAAGUGGUUUCGGGCUUGGUUUCCCUCGGGGCAGACUCCGAAGGUUUCGCUUAAUUUUG ((((((((......)))))...(((((...((((((((((((((((...((.....))))))))(((........)))....))))))))))....))))).)))....... ( -32.90) >consensus AAGGUUGGAAAUAAUCAAUAAUGAAAUGAAGGAGUUUGCCGAAACUUUGGCAAUAAGUGGUUUUGGGCUUGGCUUCCCUCG_GGCAGACUCCGUAGGUUUUGCUUAAUUUUG ((((((((......)))))...(((((...((((((((((((((((...((.....))))))))(((........)))....))))))))))....))))).)))....... (-29.48 = -28.44 + -1.04)



| Location | 13,775,508 – 13,775,619 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 94.43 |
| Mean single sequence MFE | -21.20 |
| Consensus MFE | -17.10 |
| Energy contribution | -17.30 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.739089 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13775508 111 - 23771897 CAAAAUUAAGCAAAACCUACGGAGUCUGCC-CGAGGGAAGCCAAGCCCAAAACCACUUAUUGCCAAAGUUUCGGCAAACUCCUUCAUUUCAUUAUUGAUUAUUUCCAACCUU ...((((((...........(((((.((((-.(((((........)))......((((.......)))).)))))).)))))............))))))............ ( -20.10) >DroSec_CAF1 22129 108 - 1 CAAAAUUAAGCAAAACCUACGGAGUCUGCC-CGAGGGAAGCCAAGCCCAAAACCACUUAUUGCCAAAGUUUCGGCAAACUCCUUCAUUUCAUUAUUGA---UUUCCAACCUU ..(((((((...........(((((.((((-.(((((........)))......((((.......)))).)))))).)))))............))))---)))........ ( -20.40) >DroSim_CAF1 17797 111 - 1 CAAAAUUAAGCAAAACCUACGGAGUCUGCC-CGAGGGAAGUCAAGCCCAAAACCACUUAUUGCCAAAGUUUCGGCAAACUCCUUCAUUUCAUUAUUGAUUAUUUCCAACCUU ...((((((...........(((((.((((-.(((((........)))......((((.......)))).)))))).)))))............))))))............ ( -20.10) >DroEre_CAF1 21567 111 - 1 CCAAAUUAAGCGAAACCUUCAGAGUCUGCC-CGAGGGAAACCAAGCCCGAAACCACUUAUUGCCAAAGUUUCGGCAAACUCCUUCAUUCCAUUAUUGAUUAUUUUCAACCUU .....................((((.((((-...((....))......(((((..............))))))))).)))).............((((......)))).... ( -20.24) >DroYak_CAF1 20302 112 - 1 CAAAAUUAAGCGAAACCUUCGGAGUCUGCCCCGAGGGAAACCAAGCCCGAAACCACUUAUUGCCAAAGUUUCGGCAAACUCCUUCAUUUCAUUAUUGAUUAUUUUCAACCUU ...........((((.....(((((.((((....((....))......(((((..............))))))))).)))))....))))....((((......)))).... ( -25.14) >consensus CAAAAUUAAGCAAAACCUACGGAGUCUGCC_CGAGGGAAGCCAAGCCCAAAACCACUUAUUGCCAAAGUUUCGGCAAACUCCUUCAUUUCAUUAUUGAUUAUUUCCAACCUU ....................(((((.((((....(((........)))......((((.......))))...)))).))))).............................. (-17.10 = -17.30 + 0.20)

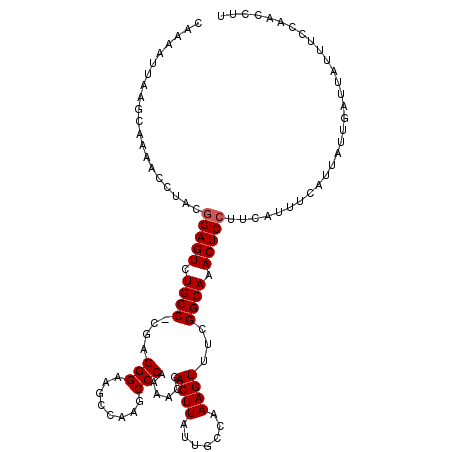

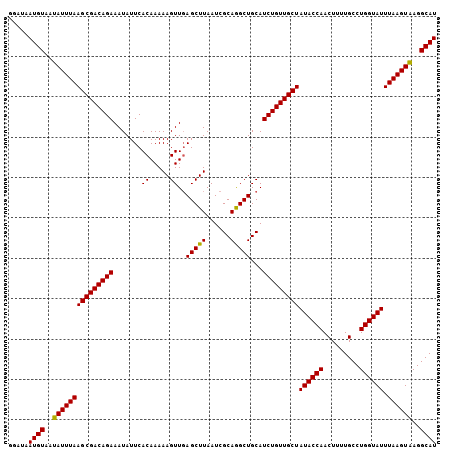
| Location | 13,775,619 – 13,775,725 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 97.64 |
| Mean single sequence MFE | -25.77 |
| Consensus MFE | -24.65 |
| Energy contribution | -24.27 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.866534 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13775619 106 - 23771897 GACUAAUGUAAUAUUUAAGCGACAGAAAUAUUCACAAAAAGUUCAGCUUAAUCGCAAGCUGCAUCUGUUGCUAUACCAACUUUUGCCUGGUAUUUAAGUAAGGCAU .....((((..(((((((((((((((.......((.....)).((((((......))))))..)))))))))((((((.........)))))).))))))..)))) ( -26.60) >DroSim_CAF1 17908 106 - 1 GGAUAAUGUAAUAUUUAAGCGACAGAAAUAUUCACAAAAAGUUGAGCUUAAUCGCAGGCUGCAUCUGUUGCUAUACCAACUUUUGCCUGGUAUUUAAGUAAGGCAU .....((((..(((((((((((((((........(((....)))(((((......)))))...)))))))))((((((.........)))))).))))))..)))) ( -25.60) >DroEre_CAF1 21678 106 - 1 GGAUAAUGUAAUAUUUAAGCGACAGAAAUAUUCACAAAAAGUUGAGCUUAAUCGCAGGCUGCAUCUGUUGCUAUACCAACUUUUGCCUGGUAUUUAAGUGAGGCAU .....((((..(((((((((((((((........(((....)))(((((......)))))...)))))))))((((((.........)))))).))))))..)))) ( -25.30) >DroYak_CAF1 20414 106 - 1 GGAUAAUGUAAUAUUUAAGCGACAGAAAUAUUCACAAAAAGUUGAGCUUAAUCGCAGGCUGCAUCUGUUGCUAUACCAACUUUUGCCUGGUAUUUAAGUAAGGCAU .....((((..(((((((((((((((........(((....)))(((((......)))))...)))))))))((((((.........)))))).))))))..)))) ( -25.60) >consensus GGAUAAUGUAAUAUUUAAGCGACAGAAAUAUUCACAAAAAGUUGAGCUUAAUCGCAGGCUGCAUCUGUUGCUAUACCAACUUUUGCCUGGUAUUUAAGUAAGGCAU .....((((..(((((((((((((((.......((.....))..(((((......)))))...)))))))))((((((.........)))))).))))))..)))) (-24.65 = -24.27 + -0.37)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:54 2006