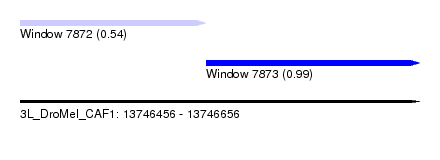
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,746,456 – 13,746,656 |
| Length | 200 |
| Max. P | 0.988673 |

| Location | 13,746,456 – 13,746,549 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 98.71 |
| Mean single sequence MFE | -20.48 |
| Consensus MFE | -18.64 |
| Energy contribution | -19.04 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.543568 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13746456 93 + 23771897 GGAAACGAACAAAUAAAAGGCUCAGCCCGCAAUCAUUGCGGUAAACAAAAGACAACAGCAGGGUUCACGAGGACACAUGUUUAUCAAUGUCCU (....)((((........(((...)))((((.....))))......................))))...((((((..((.....)).)))))) ( -18.70) >DroSec_CAF1 153214 93 + 1 GGAAACGAACAAAUAAAAGGCUCAGCCCGCAAUCAUUGCGGUAAACAAAAGACAACUGCAGGGUUCACGAGGACACAUGUUUAUCAAUGUCCU (....)((((........(((...)))........(((((((............))))))).))))...((((((..((.....)).)))))) ( -20.70) >DroSim_CAF1 164354 93 + 1 GGAAACGAACAAAUAAAAGGCUCAGCCCGCAAUCAUUGCGGUAAACAAAAGACAACUGCAGGGUUCACGAGGACACAUGUUUAUCAAUGUCCU (....)((((........(((...)))........(((((((............))))))).))))...((((((..((.....)).)))))) ( -20.70) >DroEre_CAF1 160058 92 + 1 GGAAACGAACAAAUAAAAGGCUCAGC-CGCAAUCAUUGCGGUAAACAAAAGACAACUGCAGGGUUCACGAGGACACAUGUUUAUCAAUGUCCU (....)((((........(((...))-).......(((((((............))))))).))))...((((((..((.....)).)))))) ( -21.60) >DroYak_CAF1 164303 93 + 1 GGAGACGAACAAAUAAAAGGCUCAGCCCGCAAUCAUUGCGGUAAACAAAAGACAACUGCAGGGUUCACGAGGACACAUGUUUAUCAAUGUCCU (....)((((........(((...)))........(((((((............))))))).))))...((((((..((.....)).)))))) ( -20.70) >consensus GGAAACGAACAAAUAAAAGGCUCAGCCCGCAAUCAUUGCGGUAAACAAAAGACAACUGCAGGGUUCACGAGGACACAUGUUUAUCAAUGUCCU (....)((((........(((...)))........(((((((............))))))).))))...((((((..((.....)).)))))) (-18.64 = -19.04 + 0.40)



| Location | 13,746,549 – 13,746,656 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 93.13 |
| Mean single sequence MFE | -25.33 |
| Consensus MFE | -20.90 |
| Energy contribution | -22.12 |
| Covariance contribution | 1.22 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.13 |
| SVM RNA-class probability | 0.988673 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
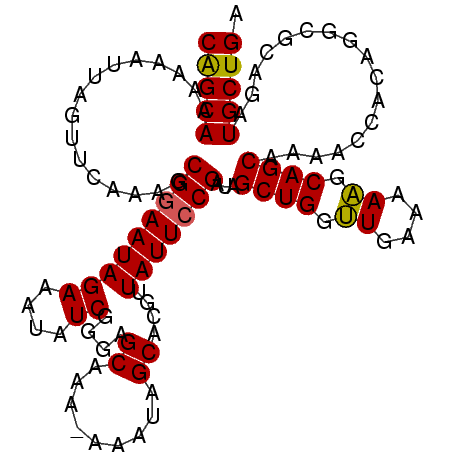
>3L_DroMel_CAF1 13746549 107 + 23771897 CAGCAAAAAAUUAGUUCAAAGCGGAAUAGAAAUAUCGGAAGCAAA-AAAUAGCACGGUAUUCCGAUAGCUGGUUGAAAAAGCAGCAAAACCACAGGCGCAGAUGCUGA (((((........((.....))((........((((((((((...-.....))......))))))))((((.((....)).))))....))...........))))). ( -26.00) >DroSec_CAF1 153307 107 + 1 CAGCAAAAAAUUAGUUCAAAGCGGAAUAGAAAUAUCGGGGGCAAA-AAAUUGCACGUUAUUCCGAUAGCUGAUUGAAAAAGCAGCAAAACCACAGGCGCAGAUGCUGA (((((...((((((((.....((((((((......((...((((.-...)))).))))))))))..))))))))......((.((..........))))...))))). ( -25.80) >DroSim_CAF1 164447 107 + 1 CAGCAAAAAAUUAGUUCAAAGCGGAAUAGAAAUAUCGGGGGCAAA-AAAUUGCACGUUAUUCCGAUAGCUGGUUGAAAAAGCAGCAAAACCACAGGCGCAGAUGCUGA (((((................((((((((......((...((((.-...)))).))))))))))...((((.((....)).)))).................))))). ( -26.90) >DroEre_CAF1 160150 108 + 1 CGGCAAAAAAUUAGUUCAAAGCGGAAUAGAAACAUCGGGAGCAAAAAAAUAGCACGUUAUUGCGAUAGCUGGUUGAAAAAGCAGCAAAGCCACAGACGCAGAUGCUGA (((((........((((......)))).......((.(..((.........(((......)))....((((.((....)).))))...))..).))......))))). ( -21.90) >DroYak_CAF1 164396 108 + 1 CAGCAAAAAAUUAGUUCAAAGCGGAAUAGAAGCAUCGAGAGCAAAAAAAUAGCACGUUAUUCCGAUAGCUGGCUGAAAAGGCAGCCAAACCACAGACGCAGAUGCUGA (((((................((((((((..((..................))...))))))))...((((.((....)).)))).................))))). ( -26.07) >consensus CAGCAAAAAAUUAGUUCAAAGCGGAAUAGAAAUAUCGGGAGCAAA_AAAUAGCACGUUAUUCCGAUAGCUGGUUGAAAAAGCAGCAAAACCACAGGCGCAGAUGCUGA (((((................(((((((((....))....((.........))....)))))))...((((.((....)).)))).................))))). (-20.90 = -22.12 + 1.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:38:42 2006