
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,710,366 – 13,710,458 |
| Length | 92 |
| Max. P | 0.748481 |

| Location | 13,710,366 – 13,710,458 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 86.84 |
| Mean single sequence MFE | -22.60 |
| Consensus MFE | -17.67 |
| Energy contribution | -17.80 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.559016 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
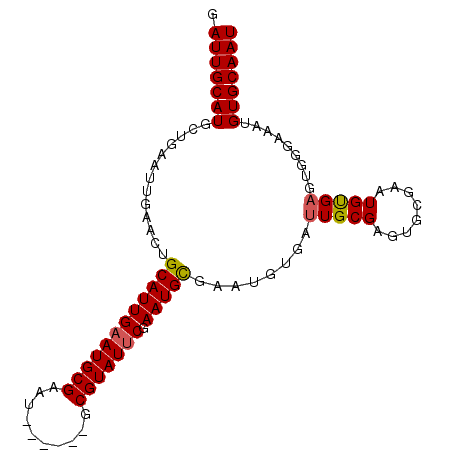
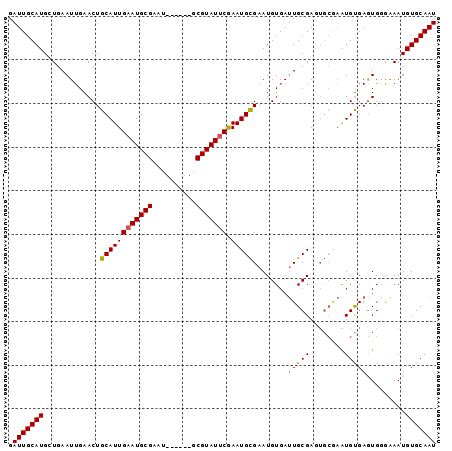
>3L_DroMel_CAF1 13710366 92 + 23771897 GAUUGCAUGGUGAAUUGAACUGCAUUGAAUGCGAAU------GCGUAUUCGAAUGCGAAUGUGAUUGCGAGUGCGAAUGUGAGUGGAAAAUGUGCAAU .((((((((.........(((((((((((((((...------.)))))))...((((..(((....)))..))))))))).)))......)))))))) ( -23.06) >DroSec_CAF1 117371 92 + 1 GAUUGCAUGCUGAAUUGAACUGCAUUGAAUGCGAAU------GCGUAUUCGAAUGCGAAUGUGAUUGCGAGUGCAAAUGUGAGUGGGAAAUGUGCAAU .(((((((.((..(((.(..(((((((((((((...------.)))))))...(((((......)))))))))))....).))).))....))))))) ( -23.20) >DroSim_CAF1 128281 92 + 1 GAUUGCAUGGUGAAUUGAACUGCAUUGAAUGCGAAU------GCGUAUUCGAAUGCGAAUGUGAUUGCGAGUGCGAAUGUGAGUGGGAAAUGUGCAAU .((((((((.........(((((((((((((((...------.)))))))...((((..(((....)))..))))))))).)))......)))))))) ( -23.06) >DroYak_CAF1 127534 92 + 1 GAUUGCAUGCUGAAUUGAACUGCAUUGAAUGCGAUUGCGAAUGCGUAUCCAAAUGUGAAUGUGAAUGCGA------GUGCGAAUGUGCAAUGUGCAAU .(((((((((...........))......((((.(((((..(((((.((((........)).))))))).------.)))))...))))..))))))) ( -21.10) >consensus GAUUGCAUGCUGAAUUGAACUGCAUUGAAUGCGAAU______GCGUAUUCGAAUGCGAAUGUGAUUGCGAGUGCGAAUGUGAGUGGGAAAUGUGCAAU .(((((((.............((((((((((((..........))))))).)))))........(((((........))))).........))))))) (-17.67 = -17.80 + 0.13)

| Location | 13,710,366 – 13,710,458 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 86.84 |
| Mean single sequence MFE | -15.38 |
| Consensus MFE | -9.79 |
| Energy contribution | -9.22 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.748481 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
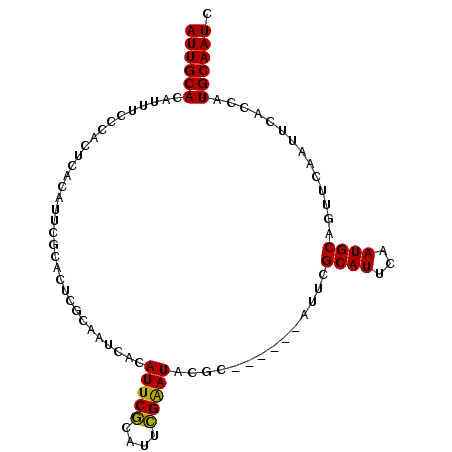
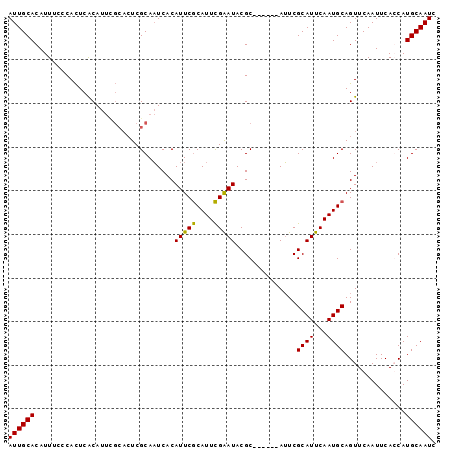
>3L_DroMel_CAF1 13710366 92 - 23771897 AUUGCACAUUUUCCACUCACAUUCGCACUCGCAAUCACAUUCGCAUUCGAAUACGC------AUUCGCAUUCAAUGCAGUUCAAUUCACCAUGCAAUC ((((((..................((....))..........(((((.((((.((.------...)).)))))))))..............)))))). ( -14.60) >DroSec_CAF1 117371 92 - 1 AUUGCACAUUUCCCACUCACAUUUGCACUCGCAAUCACAUUCGCAUUCGAAUACGC------AUUCGCAUUCAAUGCAGUUCAAUUCAGCAUGCAAUC ((((((................((((....))))........(((((.((((.((.------...)).)))))))))..............)))))). ( -15.90) >DroSim_CAF1 128281 92 - 1 AUUGCACAUUUCCCACUCACAUUCGCACUCGCAAUCACAUUCGCAUUCGAAUACGC------AUUCGCAUUCAAUGCAGUUCAAUUCACCAUGCAAUC ((((((..................((....))..........(((((.((((.((.------...)).)))))))))..............)))))). ( -14.60) >DroYak_CAF1 127534 92 - 1 AUUGCACAUUGCACAUUCGCAC------UCGCAUUCACAUUCACAUUUGGAUACGCAUUCGCAAUCGCAUUCAAUGCAGUUCAAUUCAGCAUGCAAUC ((((((...(((......((..------..))..............((((((..(((((.((....))....))))).))))))....))))))))). ( -16.40) >consensus AUUGCACAUUUCCCACUCACAUUCGCACUCGCAAUCACAUUCGCAUUCGAAUACGC______AUUCGCAUUCAAUGCAGUUCAAUUCACCAUGCAAUC ((((((................................(((((....)))))..............((((...))))..............)))))). ( -9.79 = -9.22 + -0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:37:58 2006