
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,601,688 – 13,601,906 |
| Length | 218 |
| Max. P | 0.996950 |

| Location | 13,601,688 – 13,601,791 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.78 |
| Mean single sequence MFE | -26.03 |
| Consensus MFE | -13.60 |
| Energy contribution | -13.49 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.628077 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13601688 103 + 23771897 AUACACAACAAUAACUGUCAGAGGAAAACUGUUGGGUGCUGCUGGUC-------CACC---CCAA-------AGAAGCGGCAUUUAUUGUUGUUAUUAUUGCAUUAUCCGUUGCAUACUC ....(((((((((((((.(((.......))).)))((((((((((..-------..))---....-------...))))))))))))))))))......((((........))))..... ( -24.91) >DroSec_CAF1 8605 100 + 1 AUACACAACAAUAACUGUCAGUGGAAAACUGUUGGGUG---CUGGUC-------CACC---CCAA-------AGAAGCGGCAUUUAUUGUUGUUAUUAUUGCAUUAUCCGUUGCAUACUU ....((((((((((.(((((((.....)))...(((((---......-------))))---)...-------......)))).))))))))))......((((........))))..... ( -26.40) >DroSim_CAF1 17864 102 + 1 AUACACAACAAUAACUGUCAGAGGAAAACUGCUGGGUGC-GCUGGUC-------CACC---CCAA-------AGAAGCGGCAUUUAUUGUUGUUAUUAUUGCAUUAUCCGUUGCAUACUU ....((((((((((.((((..((.....))((((((((.-.......-------))))---)...-------...))))))).))))))))))......((((........))))..... ( -26.60) >DroEre_CAF1 11418 100 + 1 AUACACAACAAUAACUGUCAGGGAGAAACGGGUGGGUGC---UACUC-------CACC---CGAA-------AGAAGCGGCAUUUAUUGUUGUUAUUAUUGCAUUAUCCGUUGCAUACUU ....((((((((((.((((.........((((((((...---...))-------))))---))..-------......)))).))))))))))......((((........))))..... ( -30.83) >DroYak_CAF1 14760 117 + 1 AUACACAACAAUAACUGUCGGGGAGAAACUGGUGGCUGC---CCCUCUCCUUUCCACCCCCCAAAAAUAAAUAAAAGCGGCAUUUAUUGUUGUUAUUAUUGCAUUAUCCGUUGCAUACUU ....((((((((((.(((((((((((....(((....))---)..))))))..........................))))).))))))))))......((((........))))..... ( -26.86) >DroAna_CAF1 44449 84 + 1 AUACACAAUAAUAACUGUCAGGA--------------G---UUGGCC-------CGCC---CA---------GAUAGCAGCGUUUAUUGUUGUUAUUAUUGCAUUAUCCGCUGCACUCUU ...................((((--------------(---((((..-------...)---))---------....((((((..((.(((.((....)).))).))..)))))))))))) ( -20.60) >consensus AUACACAACAAUAACUGUCAGGGGAAAACUGGUGGGUGC__CUGGUC_______CACC___CCAA_______AGAAGCGGCAUUUAUUGUUGUUAUUAUUGCAUUAUCCGUUGCAUACUU ....((((((((((.((((...........................................................)))).))))))))))......((((........))))..... (-13.60 = -13.49 + -0.11)


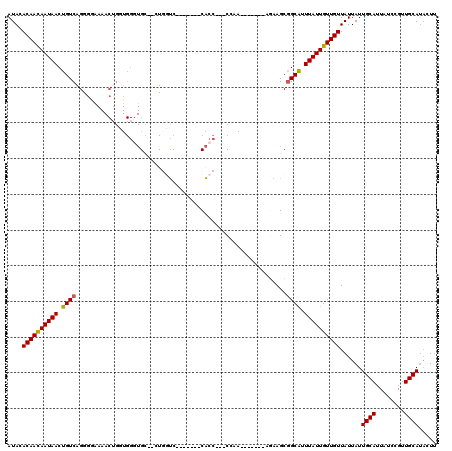
| Location | 13,601,751 – 13,601,871 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.33 |
| Mean single sequence MFE | -25.85 |
| Consensus MFE | -23.93 |
| Energy contribution | -24.43 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.77 |
| SVM RNA-class probability | 0.996950 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13601751 120 - 23771897 AGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGAGAGUAUGCAACGGAUAAUGCAAUAAUAACAACAAUAAAUG .((...(((((...(((.((((((((..(((((..(......)..)))))..))))))))....(((((....))))).......)))..)))))...)).................... ( -25.50) >DroSec_CAF1 8665 120 - 1 AGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGAAAGUAUGCAACGGAUAAUGCAAUAAUAACAACAAUAAAUG .((...(((((...(((.((((((((..(((((..(......)..)))))..))))))))....(((((....))))).......)))..)))))...)).................... ( -25.50) >DroSim_CAF1 17926 120 - 1 AGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGAAAGUAUGCAACGGAUAAUGCAAUAAUAACAACAAUAAAUG .((...(((((...(((.((((((((..(((((..(......)..)))))..))))))))....(((((....))))).......)))..)))))...)).................... ( -25.50) >DroEre_CAF1 11478 120 - 1 AGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGAAAGUAUGCAACGGAUAAUGCAAUAAUAACAACAAUAAAUG .((...(((((...(((.((((((((..(((((..(......)..)))))..))))))))....(((((....))))).......)))..)))))...)).................... ( -25.50) >DroYak_CAF1 14837 120 - 1 AGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUCGAAAGUAUGCAACGGAUAAUGCAAUAAUAACAACAAUAAAUG .((...(((((...(((.((((((((..(((((..(......)..)))))..)))))))).....((((....))))........)))..)))))...)).................... ( -24.00) >DroAna_CAF1 44493 120 - 1 AGUUACGUCUGCCGGCAAUAAUUGCAGCUGUAUUUUUAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCCCGAAAGAGUGCAGCGGAUAAUGCAAUAAUAACAACAAUAAACG .((...((((((......((((((((..(((((..((....))..)))))..))))))))............((((.(....).)))).))))))...)).................... ( -29.10) >consensus AGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGAAAGUAUGCAACGGAUAAUGCAAUAAUAACAACAAUAAAUG .((...(((((...(((.((((((((..(((((..(......)..)))))..))))))))....(((((....))))).......)))..)))))...)).................... (-23.93 = -24.43 + 0.50)



| Location | 13,601,791 – 13,601,906 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.17 |
| Mean single sequence MFE | -29.00 |
| Consensus MFE | -25.09 |
| Energy contribution | -25.45 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.871260 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13601791 115 - 23771897 CC----CCC-AGUGGAACAGACGAUAAAUUUCUGCCCAAGAGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGA ..----...-.((.(..((((((.(((...(((.....))).))))))))).).))..((((((((..(((((..(......)..)))))..))))))))....(((((....))))).. ( -28.40) >DroSec_CAF1 8705 115 - 1 CC----CCC-AGUGGAACAGACGAUAAAUUUCCGCCCAAGAGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGA ..----...-.((.(..((((((...(((((........))))).)))))).).))..((((((((..(((((..(......)..)))))..))))))))....(((((....))))).. ( -25.90) >DroSim_CAF1 17966 115 - 1 CC----CCC-AGUGGAACAGACGAUAAAUUUCUGCCCAAGAGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGA ..----...-.((.(..((((((.(((...(((.....))).))))))))).).))..((((((((..(((((..(......)..)))))..))))))))....(((((....))))).. ( -28.40) >DroEre_CAF1 11518 114 - 1 CC-----CC-ACUGGAACAGACGAUAAAUUUCUGCCCAAGAGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGA ..-----..-.((((..((((((.(((...(((.....))).)))))))))))))...((((((((..(((((..(......)..)))))..))))))))....(((((....))))).. ( -30.20) >DroYak_CAF1 14877 114 - 1 CC-----CC-ACUGGAACAGACGAUAAAUUUCUGCCCAAGAGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUCGA ..-----..-.((((..((((((.(((...(((.....))).)))))))))))))...((((((((..(((((..(......)..)))))..)))))))).....((((....))))... ( -28.70) >DroAna_CAF1 44533 119 - 1 CAGCGCUCGAGCUCGAACAGACGAUAAAUUUCAGC-CAAGAGUUACGUCUGCCGGCAAUAAUUGCAGCUGUAUUUUUAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCCCGA ..(((((((.((.((..((((((.(((...((...-...)).))))))))).))))..((((((((..(((((..((....))..)))))..)))))))).......)))..)))).... ( -32.40) >consensus CC____CCC_ACUGGAACAGACGAUAAAUUUCUGCCCAAGAGUUACGUCUGCCGGCAAUAAUUGCAACUGUAUUUUAAAAGAAUUAUGCACGUGCAAUUAAAUAAAGCGACAGCGCUUGA .............((..((((((.(((...(((.....))).)))))))))))(((..((((((((..(((((..(......)..)))))..))))))))......((....)))))... (-25.09 = -25.45 + 0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:35:32 2006