
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,598,016 – 13,598,168 |
| Length | 152 |
| Max. P | 0.999781 |

| Location | 13,598,016 – 13,598,128 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.06 |
| Mean single sequence MFE | -28.15 |
| Consensus MFE | -26.59 |
| Energy contribution | -26.95 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.94 |
| SVM decision value | 4.07 |
| SVM RNA-class probability | 0.999781 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
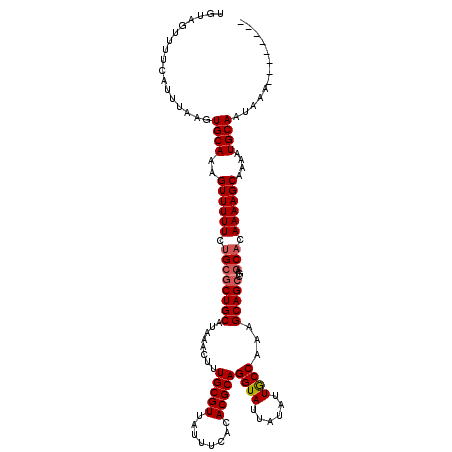
>3L_DroMel_CAF1 13598016 112 - 23771897 UGUAGUUUUCAUUUAAGUGCAAAGUUUUUCUGCGCUGCAUAAACUUUGCGUUAUUUCACACGCAGGUAUUAUAUUGCCAAAGCAGCCGAGCACAAAAGCAAAAUGCAAUAAA-------- .................((((..((((((.((((((((........(((((........)))))((((......))))...)))))...))).))))))....)))).....-------- ( -28.60) >DroSec_CAF1 4855 112 - 1 UGUAGUUUUCAUUUAAGUGCAAAGUUUUUCUGCGCUGCAUAAACUUUGCGUUAUUUCACACGCAGGUAUUAUAUUGCCAAAGCAGCCGAGCACAAAAGCAAAAUGCAAUAAA-------- .................((((..((((((.((((((((........(((((........)))))((((......))))...)))))...))).))))))....)))).....-------- ( -28.60) >DroSim_CAF1 14233 112 - 1 UGUAGUUUUCAUUUAAGUGCAAAGUUUUUCUGCGCUGCAUAAACUUUGCGUUAUUUCACACGCAGGUAUUAUAUUGCCAAAGCAGCCGAGCACAAAAGCAAAAUGCAAUAAA-------- .................((((..((((((.((((((((........(((((........)))))((((......))))...)))))...))).))))))....)))).....-------- ( -28.60) >DroEre_CAF1 6063 112 - 1 UGUAGUUUUCAUUUAAGUGCAAAGUUUUUCUGCGCUGCAUAAACUUUGCGUUAUUUCACACGCAGGUAUUAUAUUACCAAAGCAGCCGAGCACAAAAGCAAAAUGCAAUAAA-------- .................((((..((((((.((((((((........(((((........)))))((((......))))...)))))...))).))))))....)))).....-------- ( -28.60) >DroYak_CAF1 11012 112 - 1 UGUAGUUUUCAUUUAAGUGCAAAGUUUUUCUGCGCUGCAUAAACUUUGCGUUAUUUCACACGCAGGUAUUAUAUUGCCAAAGCAGCCGAGCACAAAAGCAAAAUGCAAUAAA-------- .................((((..((((((.((((((((........(((((........)))))((((......))))...)))))...))).))))))....)))).....-------- ( -28.60) >DroAna_CAF1 40613 114 - 1 UGUAGUUUUCAUUUAAGUGCAAAGUUUUUCCGCGCUGCAUAAACUUUGCGUUAUUUCACACGCAGGUAUUAUAUUGCCAAAGCAG-----CACAAAAGCA-AAUGCAAUAAGGCCAGAAG .......(((.((((..((((..((((((..(.(((((........(((((........)))))((((......))))...))))-----).))))))).-..)))).))))....))). ( -25.90) >consensus UGUAGUUUUCAUUUAAGUGCAAAGUUUUUCUGCGCUGCAUAAACUUUGCGUUAUUUCACACGCAGGUAUUAUAUUGCCAAAGCAGCCGAGCACAAAAGCAAAAUGCAAUAAA________ .................((((..((((((.((((((((........(((((........)))))((((......))))...)))))...))).))))))....))))............. (-26.59 = -26.95 + 0.36)


| Location | 13,598,048 – 13,598,168 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.33 |
| Mean single sequence MFE | -23.85 |
| Consensus MFE | -22.12 |
| Energy contribution | -21.90 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969800 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13598048 120 + 23771897 UUGGCAAUAUAAUACCUGCGUGUGAAAUAACGCAAAGUUUAUGCAGCGCAGAAAAACUUUGCACUUAAAUGAAAACUACAAAACAUUUAACUUUCGUCCAAAAAGGAGGCGACAGAAUUU ...............((((((..((((....(((((((((.(((...)))...)))))))))..(((((((............))))))).)))).(((.....))).))).)))..... ( -23.60) >DroSec_CAF1 4887 120 + 1 UUGGCAAUAUAAUACCUGCGUGUGAAAUAACGCAAAGUUUAUGCAGCGCAGAAAAACUUUGCACUUAAAUGAAAACUACAAAACAUUUAACUUUCGUCCAAAAUGGAGGCGACAGAAUUU ...............((((((..((((....(((((((((.(((...)))...)))))))))..(((((((............))))))).)))).(((.....))).))).)))..... ( -23.90) >DroSim_CAF1 14265 120 + 1 UUGGCAAUAUAAUACCUGCGUGUGAAAUAACGCAAAGUUUAUGCAGCGCAGAAAAACUUUGCACUUAAAUGAAAACUACAAAACAUUUAACUUUCGUCCAAAAUGGAGGCGACAGAAUUU ...............((((((..((((....(((((((((.(((...)))...)))))))))..(((((((............))))))).)))).(((.....))).))).)))..... ( -23.90) >DroEre_CAF1 6095 120 + 1 UUGGUAAUAUAAUACCUGCGUGUGAAAUAACGCAAAGUUUAUGCAGCGCAGAAAAACUUUGCACUUAAAUGAAAACUACAAAACAUUUAACUUUCGUCCAAAAAGGAGGCGACAGAAUUU ...............((((((..((((....(((((((((.(((...)))...)))))))))..(((((((............))))))).)))).(((.....))).))).)))..... ( -23.60) >DroYak_CAF1 11044 120 + 1 UUGGCAAUAUAAUACCUGCGUGUGAAAUAACGCAAAGUUUAUGCAGCGCAGAAAAACUUUGCACUUAAAUGAAAACUACAAAACAUUUAACUUUCGUCCAAAAAGGAGGCGACAGAAUUU ...............((((((..((((....(((((((((.(((...)))...)))))))))..(((((((............))))))).)))).(((.....))).))).)))..... ( -23.60) >DroAna_CAF1 40647 118 + 1 UUGGCAAUAUAAUACCUGCGUGUGAAAUAACGCAAAGUUUAUGCAGCGCGGAAAAACUUUGCACUUAAAUGAAAACUACAAAACAUUUAACUUUUGG-CAAAAAGGAGACG-CGGAAUUU ...............((((((.(........(((((((((.(((...)))...)))))))))..(((((((............)))))))(((((..-..))))).).)))-)))..... ( -24.50) >consensus UUGGCAAUAUAAUACCUGCGUGUGAAAUAACGCAAAGUUUAUGCAGCGCAGAAAAACUUUGCACUUAAAUGAAAACUACAAAACAUUUAACUUUCGUCCAAAAAGGAGGCGACAGAAUUU ...............((((((..((((....(((((((((.(((...)))...)))))))))..(((((((............))))))).)))).(((.....))).))).)))..... (-22.12 = -21.90 + -0.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:35:22 2006