
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,597,140 – 13,597,260 |
| Length | 120 |
| Max. P | 0.935147 |

| Location | 13,597,140 – 13,597,260 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.78 |
| Mean single sequence MFE | -40.80 |
| Consensus MFE | -32.96 |
| Energy contribution | -33.68 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.81 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
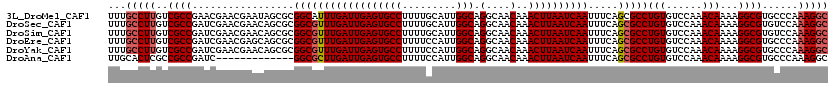
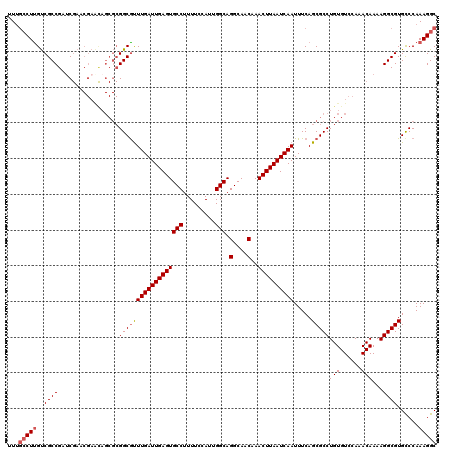
>3L_DroMel_CAF1 13597140 120 + 23771897 GCCUUUGGGCACGCCUUUUGUUUGGACACAGGCGCUGAAAUUGAUUAAGUUUGUUGCCUGCCAAUGCAAAAGGCACUCAAUCAAAUGCCGCGCUAUUCGUUCGUUCGGCGACAAGGCAAA (((((((((...(((((((((((((...((((((...(((((.....)))))..)))))))))).))))))))).))))......(((((.((.........)).)))))..)))))... ( -41.90) >DroSec_CAF1 3975 120 + 1 GCCUUUGGACACGCCUUUUGUUUGGACACAGGCGCUGAAAUUGAUUAAGUUUGUUGCCUGCCAAUGCAAAAGGCACUCAAUCAAACGCCGCGCUGUUCGUUCGAUCGGCGACAAGGCAAA (((((.......(((((((((((((...((((((...(((((.....)))))..)))))))))).)))))))))...............((((((.((....)).))))).))))))... ( -41.60) >DroSim_CAF1 13349 120 + 1 GCCUUUGGACACGCCUUUUGUUUGGACACAGGCGCUGAAAUUGAUUAAGUUUGUUGCCUGCCAAUGCAAAAGGCACUCAAUCAAACGCCGCGCUGUUCGUUCGAUCGGCGACAAGGCAAA (((((.......(((((((((((((...((((((...(((((.....)))))..)))))))))).)))))))))...............((((((.((....)).))))).))))))... ( -41.60) >DroEre_CAF1 5185 120 + 1 GCCUUUGGGCACGCCUUUUGUUUGGACACAGGCGCUGAAAUUGAUUAAGUUUGUUGCCUGCCAAUGGAAAAGGCACUCAAUCAAACGCCGCGCUGCUCGUUCGAUCGGCGACAAGGCAAA (((....)))..(((((((.(((((...((((((...(((((.....)))))..)))))))))).).)))))))............(((.(((((.((....)).)))))....)))... ( -41.20) >DroYak_CAF1 10141 120 + 1 GCCUUUGGGCACGCCUUUUGUUUGGACACAGGCGCUGAAAUUGAUUAAGUUUGUUGCCUGCCAAUGGAAAAGGCACUCAAUCAAACGCCGCGCUGUUCGUUCGAUCGGCGACAAGGCAAA (((....)))..(((((((.(((((...((((((...(((((.....)))))..)))))))))).).)))))))............(((.(((((.((....)).)))))....)))... ( -41.30) >DroAna_CAF1 39733 107 + 1 GCCUUUGGGCACGCCUUUUGUUUGGACACAGGCGCUGAAAUUGAUUAAGUUUGUUGCCUGCCAAUGGAAAAGGCACUCAAUCAAGCGCC-------------GAUCGGCGGCGAGUGCAA ........(((((((((((.(((((...((((((...(((((.....)))))..)))))))))).).)))))))...........((((-------------(.....))))).)))).. ( -37.20) >consensus GCCUUUGGGCACGCCUUUUGUUUGGACACAGGCGCUGAAAUUGAUUAAGUUUGUUGCCUGCCAAUGCAAAAGGCACUCAAUCAAACGCCGCGCUGUUCGUUCGAUCGGCGACAAGGCAAA (((((((((...(((((((..((((...((((((...(((((.....)))))..))))))))))...))))))).))))......(((((..(.........)..)))))..)))))... (-32.96 = -33.68 + 0.72)



| Location | 13,597,140 – 13,597,260 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.78 |
| Mean single sequence MFE | -38.85 |
| Consensus MFE | -34.14 |
| Energy contribution | -34.67 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.935147 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13597140 120 - 23771897 UUUGCCUUGUCGCCGAACGAACGAAUAGCGCGGCAUUUGAUUGAGUGCCUUUUGCAUUGGCAGGCAACAAACUUAAUCAAUUUCAGCGCCUGUGUCCAAACAAAAGGCGUGCCCAAAGGC ...(((((..((((.............((((.....(((((((((((((.........))).(....)..)))))))))).....)))).(((......)))...))))......))))) ( -37.50) >DroSec_CAF1 3975 120 - 1 UUUGCCUUGUCGCCGAUCGAACGAACAGCGCGGCGUUUGAUUGAGUGCCUUUUGCAUUGGCAGGCAACAAACUUAAUCAAUUUCAGCGCCUGUGUCCAAACAAAAGGCGUGUCCAAAGGC ...(((((..(((((((((((((..........))))))))))...((((((((..(((((((((......................)))))...)))).)))))))))))....))))) ( -39.35) >DroSim_CAF1 13349 120 - 1 UUUGCCUUGUCGCCGAUCGAACGAACAGCGCGGCGUUUGAUUGAGUGCCUUUUGCAUUGGCAGGCAACAAACUUAAUCAAUUUCAGCGCCUGUGUCCAAACAAAAGGCGUGUCCAAAGGC ...(((((..(((((((((((((..........))))))))))...((((((((..(((((((((......................)))))...)))).)))))))))))....))))) ( -39.35) >DroEre_CAF1 5185 120 - 1 UUUGCCUUGUCGCCGAUCGAACGAGCAGCGCGGCGUUUGAUUGAGUGCCUUUUCCAUUGGCAGGCAACAAACUUAAUCAAUUUCAGCGCCUGUGUCCAAACAAAAGGCGUGCCCAAAGGC ...(((((((.((((......)).)).))(.(((..(((((((((((((.........))).(....)..)))))))))).....((((((.(((....)))..)))))))))).))))) ( -40.30) >DroYak_CAF1 10141 120 - 1 UUUGCCUUGUCGCCGAUCGAACGAACAGCGCGGCGUUUGAUUGAGUGCCUUUUCCAUUGGCAGGCAACAAACUUAAUCAAUUUCAGCGCCUGUGUCCAAACAAAAGGCGUGCCCAAAGGC ...((((((.(((.(.((....)).).))))(((..(((((((((((((.........))).(....)..)))))))))).....((((((.(((....)))..)))))))))..))))) ( -40.20) >DroAna_CAF1 39733 107 - 1 UUGCACUCGCCGCCGAUC-------------GGCGCUUGAUUGAGUGCCUUUUCCAUUGGCAGGCAACAAACUUAAUCAAUUUCAGCGCCUGUGUCCAAACAAAAGGCGUGCCCAAAGGC ........((((((....-------------((((((((((((((((((.........))).(....)..))))))))).....))))))(((......)))...)))).))........ ( -36.40) >consensus UUUGCCUUGUCGCCGAUCGAACGAACAGCGCGGCGUUUGAUUGAGUGCCUUUUCCAUUGGCAGGCAACAAACUUAAUCAAUUUCAGCGCCUGUGUCCAAACAAAAGGCGUGCCCAAAGGC ...(((((..((((.................((((((((((((((((((.........))).(....)..)))))))))).....)))))(((......)))...))))......))))) (-34.14 = -34.67 + 0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:35:19 2006