
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,570,295 – 13,570,385 |
| Length | 90 |
| Max. P | 0.838661 |

| Location | 13,570,295 – 13,570,385 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 76.41 |
| Mean single sequence MFE | -17.42 |
| Consensus MFE | -9.63 |
| Energy contribution | -9.67 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.838661 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
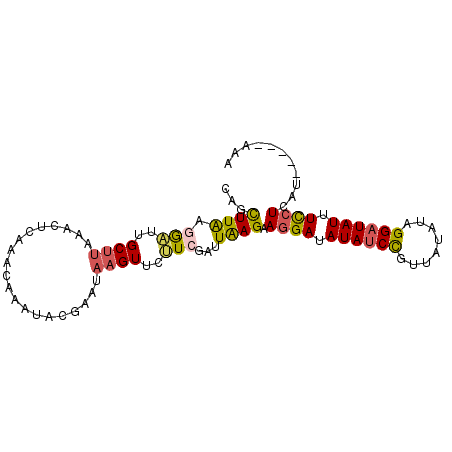
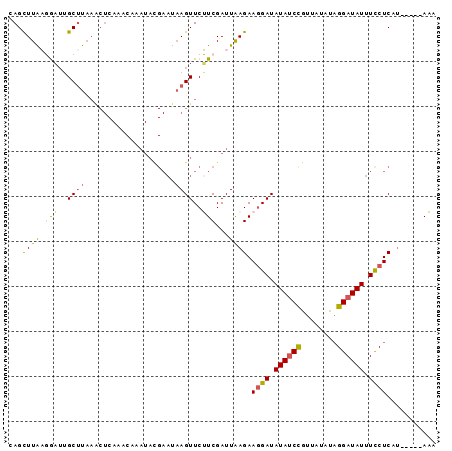
>3L_DroMel_CAF1 13570295 90 + 23771897 CAGCUUAAGGAUUGCUUUAGCUCAAACAAAUACGAAUAAGUUCUUCGAUUAAGAAGGAUAUAACCGUUAUAUAGGAUAUUUCCUCAU-----AAA .((((.(((.....))).)))).........(((..((.(((((((......))))))).))..)))..(((((((....)))).))-----).. ( -16.90) >DroSec_CAF1 51840 90 + 1 CAGCUUAAGAAUGGCUUAAACUCAAACAAAUACGAAUAAGUUCUUCGAUUAAGAAGGAUAUAUCCGUUAUAUAGGAUAUUUCCUCAU-----AAA ...((((.(((..(((((...((..........)).)))))..)))...)))).((((.((((((........)))))).))))...-----... ( -18.00) >DroSim_CAF1 38010 90 + 1 CAGCUUAAGAAUGGCUUAAACUCAAACAAAUACGAAUAAGUUCUUCGAUUAAGAAGGAUAUAUCCGUUAUAUAGGAUAUUUCCUCAU-----AAA ...((((.(((..(((((...((..........)).)))))..)))...)))).((((.((((((........)))))).))))...-----... ( -18.00) >DroEre_CAF1 53013 70 + 1 CAGCUUGUUGAAUGCCUA--------------------AGUUCAUCGAUUAAGAAAGAUAUAUCCGUUAUUUAGGAUAUAUCCUCGA-----AUA ....(((.(((((.....--------------------.))))).)))....((..(((((((((........))))))))).))..-----... ( -15.90) >DroYak_CAF1 54419 95 + 1 CAGUUUGUUGUUUGCUUAAACUCAAACAAAUAUGAAGAAGUUCAUCGAUUGAGAAGGAUAUAUCUAUUAUAUUGGAUAUUUUCUCGUGUACUAUA ..(((((..((((....)))).)))))..((((((.(((((((.((......)).))))(((((((......))))))).)))))))))...... ( -18.30) >consensus CAGCUUAAGGAUUGCUUAAACUCAAACAAAUACGAAUAAGUUCUUCGAUUAAGAAGGAUAUAUCCGUUAUAUAGGAUAUUUCCUCAU_____AAA ...((((.(((..((((....................))))..)))...)))).((((.((((((........)))))).))))........... ( -9.63 = -9.67 + 0.04)

| Location | 13,570,295 – 13,570,385 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 76.41 |
| Mean single sequence MFE | -16.78 |
| Consensus MFE | -9.14 |
| Energy contribution | -11.54 |
| Covariance contribution | 2.40 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.54 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.511075 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
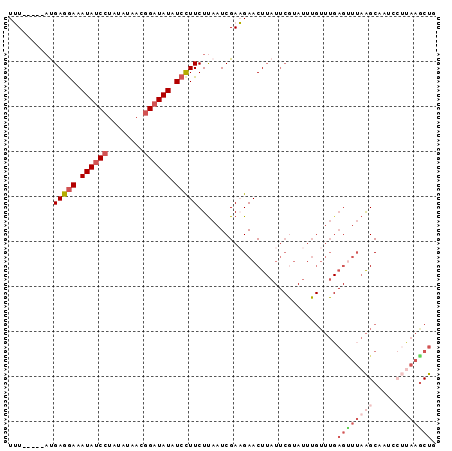
>3L_DroMel_CAF1 13570295 90 - 23771897 UUU-----AUGAGGAAAUAUCCUAUAUAACGGUUAUAUCCUUCUUAAUCGAAGAACUUAUUCGUAUUUGUUUGAGCUAAAGCAAUCCUUAAGCUG ...-----.((((((.............(((..((....((((......))))....))..)))..((((((......))))))))))))..... ( -15.40) >DroSec_CAF1 51840 90 - 1 UUU-----AUGAGGAAAUAUCCUAUAUAACGGAUAUAUCCUUCUUAAUCGAAGAACUUAUUCGUAUUUGUUUGAGUUUAAGCCAUUCUUAAGCUG ..(-----(((((((.((((((........)))))).)))(((((.....))))).....)))))........((((((((.....)))))))). ( -20.40) >DroSim_CAF1 38010 90 - 1 UUU-----AUGAGGAAAUAUCCUAUAUAACGGAUAUAUCCUUCUUAAUCGAAGAACUUAUUCGUAUUUGUUUGAGUUUAAGCCAUUCUUAAGCUG ..(-----(((((((.((((((........)))))).)))(((((.....))))).....)))))........((((((((.....)))))))). ( -20.40) >DroEre_CAF1 53013 70 - 1 UAU-----UCGAGGAUAUAUCCUAAAUAACGGAUAUAUCUUUCUUAAUCGAUGAACU--------------------UAGGCAUUCAACAAGCUG ...-----((((((((((((((........)))))))))).......))))......--------------------..(((.........))). ( -14.91) >DroYak_CAF1 54419 95 - 1 UAUAGUACACGAGAAAAUAUCCAAUAUAAUAGAUAUAUCCUUCUCAAUCGAUGAACUUCUUCAUAUUUGUUUGAGUUUAAGCAAACAACAAACUG ..((((....(((((.(((((..........)))))....))))).....(((((....)))))..(((((((........)))))))...)))) ( -12.80) >consensus UUU_____AUGAGGAAAUAUCCUAUAUAACGGAUAUAUCCUUCUUAAUCGAAGAACUUAUUCGUAUUUGUUUGAGUUUAAGCAAUCCUUAAGCUG ..........(((((.((((((........)))))).)))))...............................((((((((.....)))))))). ( -9.14 = -11.54 + 2.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:34:52 2006