
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,535,513 – 13,535,604 |
| Length | 91 |
| Max. P | 0.995680 |

| Location | 13,535,513 – 13,535,604 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 90.85 |
| Mean single sequence MFE | -26.44 |
| Consensus MFE | -23.03 |
| Energy contribution | -22.87 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.846246 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13535513 91 + 23771897 ---UCCGCUGUCUCACUGGCAGUGUCUCGCUUUUUUGUUUGCCAGCAAGUCCUUGCAUCCU--UGGGCUCUGCAUGGGCAAAAGUUCAGCGCAAAA ---..(((((((.....)))))))...((((......((((((.((((((((.........--.))))).)))...)))))).....))))..... ( -30.90) >DroSec_CAF1 22734 89 + 1 ---UCCGCUGUCUCACUGGCAGUGUCUCACUUUUUUGUUUGCCAGCGAGUCCUUGCAUCCU--UUGGC--UGCAUGGGCAAAAGUUCAGCGCAAAA ---..(((((..((.(((((((................))))))).))((((.((((.((.--..)).--)))).)))).......)))))..... ( -29.89) >DroSim_CAF1 8857 89 + 1 ---UCCGCUGUCUCACUGGCAGUGUCUCACUUUUUUGUUUGCCAGCGGGUCCUUGCAUCCU--UUGGC--UGCAUGGACAAAAGUUCAGCGCAAAA ---..(((((..((.(((((((................))))))).))((((.((((.((.--..)).--)))).)))).......)))))..... ( -29.39) >DroEre_CAF1 23003 89 + 1 ---UCCGUUGUCUCACUUGCAGUGUCUCGCUUUUUUGUUUGCCGGCGAGUCCUUGCAUCCU--UUGGC--UGCAUGGGUAAAAGUUCAGCGCAAAA ---..(((((....((((((((.(.((((((............)))))).).))))((((.--.((..--..)).))))..)))).)))))..... ( -21.90) >DroYak_CAF1 24538 94 + 1 CGCUCCGUUGUCUCACUUGCAGUGUCUCCCUUUUUUGUUUGCCGCCGAGUCCUUGCAUCCUUUUUGGC--UGCAUGGGUAAAAGUUCAGUGCAAAA ................(((((.((.....(((((.....(((.((((((.............))))))--.))).....)))))..)).))))).. ( -20.12) >consensus ___UCCGCUGUCUCACUGGCAGUGUCUCACUUUUUUGUUUGCCAGCGAGUCCUUGCAUCCU__UUGGC__UGCAUGGGCAAAAGUUCAGCGCAAAA .....(((((.(((.(((((((................))))))).)))(((.((((.((.....))...)))).)))........)))))..... (-23.03 = -22.87 + -0.16)

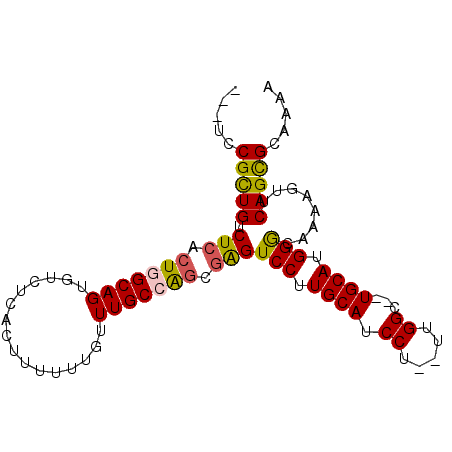
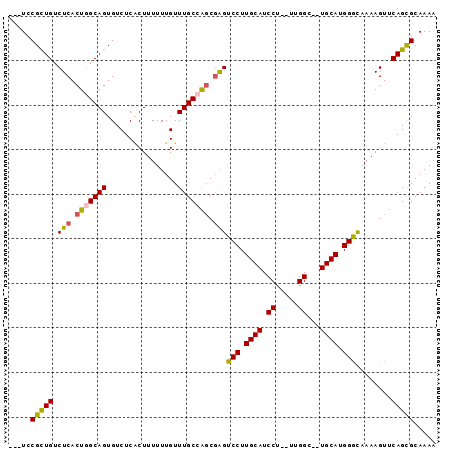
| Location | 13,535,513 – 13,535,604 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 90.85 |
| Mean single sequence MFE | -27.25 |
| Consensus MFE | -24.33 |
| Energy contribution | -25.97 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.60 |
| SVM RNA-class probability | 0.995680 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13535513 91 - 23771897 UUUUGCGCUGAACUUUUGCCCAUGCAGAGCCCA--AGGAUGCAAGGACUUGCUGGCAAACAAAAAAGCGAGACACUGCCAGUGAGACAGCGGA--- .....(((((.........((.((((...((..--.)).)))).)).((..((((((..................))))))..)).)))))..--- ( -30.07) >DroSec_CAF1 22734 89 - 1 UUUUGCGCUGAACUUUUGCCCAUGCA--GCCAA--AGGAUGCAAGGACUCGCUGGCAAACAAAAAAGUGAGACACUGCCAGUGAGACAGCGGA--- .....(((((.........((.((((--.((..--.)).)))).)).((((((((((..................)))))))))).)))))..--- ( -32.57) >DroSim_CAF1 8857 89 - 1 UUUUGCGCUGAACUUUUGUCCAUGCA--GCCAA--AGGAUGCAAGGACCCGCUGGCAAACAAAAAAGUGAGACACUGCCAGUGAGACAGCGGA--- .....(((((.......((((.((((--.((..--.)).)))).)))).((((((((..................))))))))...)))))..--- ( -31.87) >DroEre_CAF1 23003 89 - 1 UUUUGCGCUGAACUUUUACCCAUGCA--GCCAA--AGGAUGCAAGGACUCGCCGGCAAACAAAAAAGCGAGACACUGCAAGUGAGACAACGGA--- .((((((.((.........((.((((--.((..--.)).)))).)).(((((..............))))).)).))))))............--- ( -19.84) >DroYak_CAF1 24538 94 - 1 UUUUGCACUGAACUUUUACCCAUGCA--GCCAAAAAGGAUGCAAGGACUCGGCGGCAAACAAAAAAGGGAGACACUGCAAGUGAGACAACGGAGCG ....((.(((...((((((((.((((--.((.....)).)))).)).....((((....(......)(....).))))..))))))...))).)). ( -21.90) >consensus UUUUGCGCUGAACUUUUGCCCAUGCA__GCCAA__AGGAUGCAAGGACUCGCUGGCAAACAAAAAAGCGAGACACUGCCAGUGAGACAGCGGA___ .....(((((.........((.((((...((.....)).)))).)).((((((((((..................)))))))))).)))))..... (-24.33 = -25.97 + 1.64)

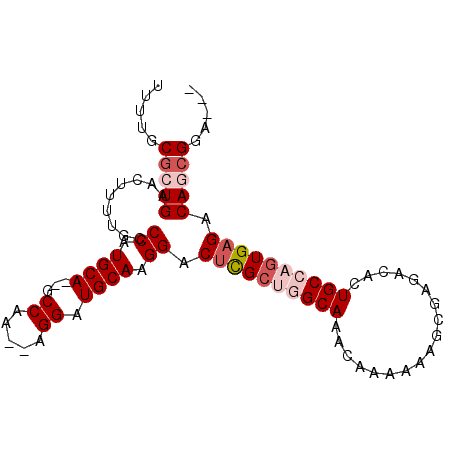

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:34:28 2006