
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,514,489 – 13,514,605 |
| Length | 116 |
| Max. P | 0.999554 |

| Location | 13,514,489 – 13,514,605 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.51 |
| Mean single sequence MFE | -35.07 |
| Consensus MFE | -33.77 |
| Energy contribution | -33.97 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.96 |
| SVM decision value | 3.72 |
| SVM RNA-class probability | 0.999554 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
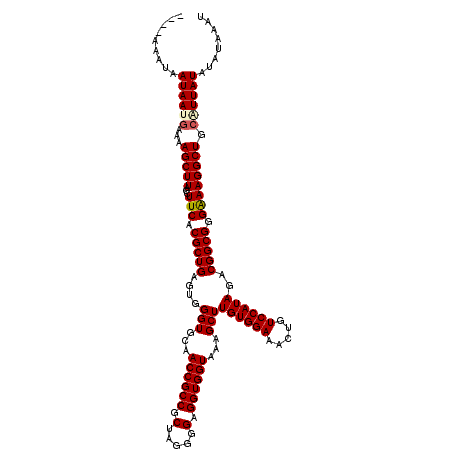
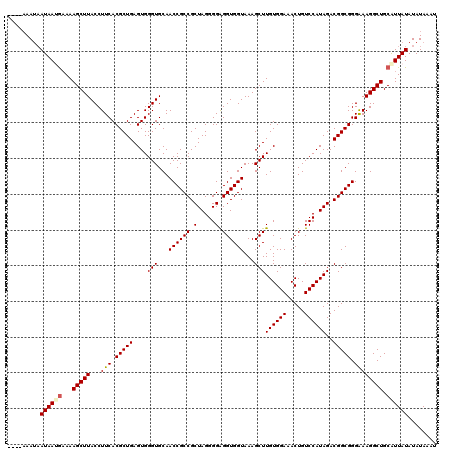
>3L_DroMel_CAF1 13514489 116 + 23771897 ----AAAUUAUAAAGAAAAGCUUACCUUCACGCUGAGUGGGUGCAACCGCCGCUAGGGGAGGUGGUAAAGCUUGUGGAAACUGUCCAUAGACGGCGGGAAAGGCUGCAUUAUAUAUAAAU ----....(((((.(...(((((...(((.(((((....(((...((((((.(.....).))))))...)))((((((.....))))))..))))).)))))))).).)))))....... ( -32.50) >DroPse_CAF1 1944 116 + 1 ----AAAUAAUAAUGAAAAGCUUACCUUCACGCUGAGUGGGUGCAACCGCCGCUAGGGGAGGUGGUAAAGCUUGUGGAAACUGUCCAUAGACGGCGGGGAAGGCUGCAUUAUAUAUAGAU ----.....((((((...(((((.(((...(((((....(((...((((((.(.....).))))))...)))((((((.....))))))..)))))))).))))).))))))........ ( -37.60) >DroWil_CAF1 2903 120 + 1 AAAUCAAAAAUAAUAUAAAGCUUACCUUCACGCUGAGUGGGUGCAACCGCCGCUAGGGGGGGUGGUAAAGCUUGUGGAAACUGUCCAUAGACGGCGGGAAAGGCUGCAUUAUAUAUAAAU ............((((((.((...(((((.(((((....(((...((((((.(.....).))))))...)))((((((.....))))))..))))).).))))..)).))))))...... ( -36.00) >DroMoj_CAF1 1843 116 + 1 ----AAAUAAUAAUGAAAAGCUUACCUUCACGCUGAGUGGGUGCAACCGCCGCUAGGGGAGGUGGUAAAGCUUGUGGAAACUGUCCAUAGACGGCGGGAAAGGCUGCGUUAUAUAUAAAU ----.....((((((...(((((...(((.(((((....(((...((((((.(.....).))))))...)))((((((.....))))))..))))).)))))))).))))))........ ( -35.30) >DroAna_CAF1 1764 116 + 1 ----AAAUAAUAAAGAAAAGCUUACCUUCACGCUGAGUGGGUGCAACCGCCGCUAGGGGAGGUGGUAAAGCUUGUGGAAACUGUCCAUAGACGGCGGGAAAGGCUGCAUUAUAUAUAAAU ----.....((((.(...(((((...(((.(((((....(((...((((((.(.....).))))))...)))((((((.....))))))..))))).)))))))).).))))........ ( -31.40) >DroPer_CAF1 2036 116 + 1 ----AAAUAAUAAUGAAAAGCUUACCUUCACGCUGAGUGGGUGCAACCGCCGCUAGGGGAGGUGGUAAAGCUUGUGGAAACUGUCCAUAGACGGCGGGGAAGGCUGCAUUAUAUAUAGAU ----.....((((((...(((((.(((...(((((....(((...((((((.(.....).))))))...)))((((((.....))))))..)))))))).))))).))))))........ ( -37.60) >consensus ____AAAUAAUAAUGAAAAGCUUACCUUCACGCUGAGUGGGUGCAACCGCCGCUAGGGGAGGUGGUAAAGCUUGUGGAAACUGUCCAUAGACGGCGGGAAAGGCUGCAUUAUAUAUAAAU .........((((((...(((((...(((.(((((....(((...((((((.(.....).))))))...)))((((((.....))))))..))))).)))))))).))))))........ (-33.77 = -33.97 + 0.19)

| Location | 13,514,489 – 13,514,605 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.51 |
| Mean single sequence MFE | -24.64 |
| Consensus MFE | -23.98 |
| Energy contribution | -23.76 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -0.98 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.886209 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
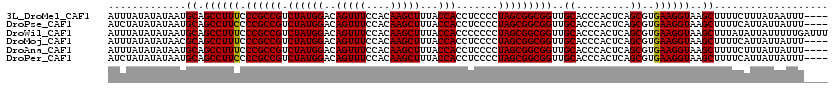
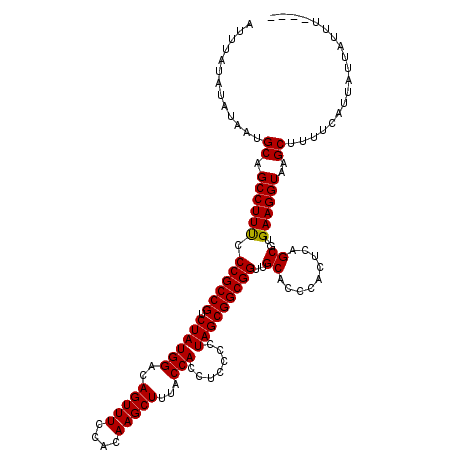
>3L_DroMel_CAF1 13514489 116 - 23771897 AUUUAUAUAUAAUGCAGCCUUUCCCGCCGUCUAUGGACAGUUUCCACAAGCUUUACCACCUCCCCUAGCGGCGGUUGCACCCACUCAGCGUGAAGGUAAGCUUUUCUUUAUAAUUU---- ..(((((......((.(((((..((((((.((((((..(((((....)))))...))).......))))))))).......(((.....))))))))..)).......)))))...---- ( -24.33) >DroPse_CAF1 1944 116 - 1 AUCUAUAUAUAAUGCAGCCUUCCCCGCCGUCUAUGGACAGUUUCCACAAGCUUUACCACCUCCCCUAGCGGCGGUUGCACCCACUCAGCGUGAAGGUAAGCUUUUCAUUAUUAUUU---- .............((.((((((.((((((.((((((..(((((....)))))...))).......)))))))))..((.........))..))))))..))...............---- ( -25.11) >DroWil_CAF1 2903 120 - 1 AUUUAUAUAUAAUGCAGCCUUUCCCGCCGUCUAUGGACAGUUUCCACAAGCUUUACCACCCCCCCUAGCGGCGGUUGCACCCACUCAGCGUGAAGGUAAGCUUUAUAUUAUUUUUGAUUU ......((((((.((.(((((..((((((.((((((..(((((....)))))...))).......))))))))).......(((.....))))))))..)).))))))............ ( -26.41) >DroMoj_CAF1 1843 116 - 1 AUUUAUAUAUAACGCAGCCUUUCCCGCCGUCUAUGGACAGUUUCCACAAGCUUUACCACCUCCCCUAGCGGCGGUUGCACCCACUCAGCGUGAAGGUAAGCUUUUCAUUAUUAUUU---- .............((.(((((..((((((.((((((..(((((....)))))...))).......))))))))).......(((.....))))))))..))...............---- ( -23.51) >DroAna_CAF1 1764 116 - 1 AUUUAUAUAUAAUGCAGCCUUUCCCGCCGUCUAUGGACAGUUUCCACAAGCUUUACCACCUCCCCUAGCGGCGGUUGCACCCACUCAGCGUGAAGGUAAGCUUUUCUUUAUUAUUU---- ............(((((((....((((.(((....)))(((((....)))))...............)))).)))))))..........(((((((........))))))).....---- ( -23.40) >DroPer_CAF1 2036 116 - 1 AUCUAUAUAUAAUGCAGCCUUCCCCGCCGUCUAUGGACAGUUUCCACAAGCUUUACCACCUCCCCUAGCGGCGGUUGCACCCACUCAGCGUGAAGGUAAGCUUUUCAUUAUUAUUU---- .............((.((((((.((((((.((((((..(((((....)))))...))).......)))))))))..((.........))..))))))..))...............---- ( -25.11) >consensus AUUUAUAUAUAAUGCAGCCUUUCCCGCCGUCUAUGGACAGUUUCCACAAGCUUUACCACCUCCCCUAGCGGCGGUUGCACCCACUCAGCGUGAAGGUAAGCUUUUCAUUAUUAUUU____ .............((.((((((.((((((.((((((..(((((....)))))...))).......)))))))))..((.........))..))))))..))................... (-23.98 = -23.76 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:34:18 2006