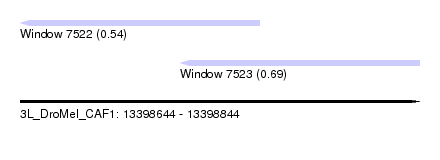
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,398,644 – 13,398,844 |
| Length | 200 |
| Max. P | 0.685787 |

| Location | 13,398,644 – 13,398,764 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.33 |
| Mean single sequence MFE | -37.38 |
| Consensus MFE | -30.34 |
| Energy contribution | -31.14 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.535767 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
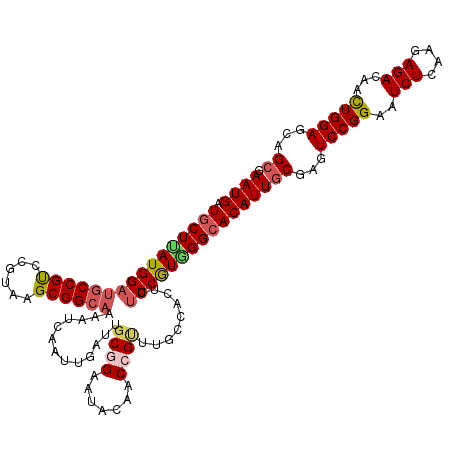
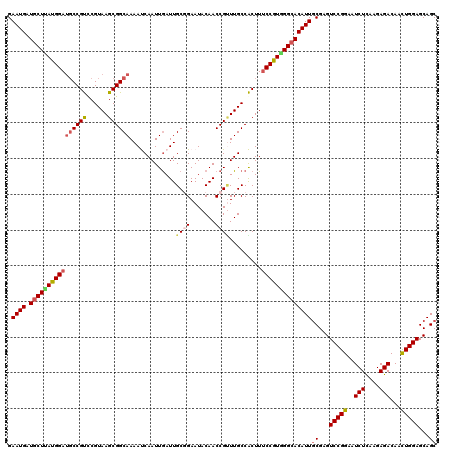
>3L_DroMel_CAF1 13398644 120 - 23771897 GAAUGGUGCUAAUGGAUGCCGUCCGUAAGCGGCAAAAUCAAUUGAUUGCGGAAUACAACCGAUUGCCACUUACCGUGGGCACAUUGCGAGUCCGGAAUCUCAAGAGACAACUGGAGCAGC .....(((((...((((...))))(((((.(((((((((....)))).(((.......))).))))).)))))....))))).((((...(((((..(((....)))...))))))))). ( -36.10) >DroSec_CAF1 14236 120 - 1 GAAUGAUGCUUAUGGAUGCCGUCCGUAAGCGGCAAAAUCAAUUGAUUGCGGAAUACAACCGUUUGCCACUUUCCGUAGGCACAUUACGAGUCCGGAAUCUCAAGAGACAACUGGAGCAGC .((((.((((((((((((((((......)))))).((((....))))((((.......)))).........)))))))))))))).....(((((..(((....)))...)))))..... ( -36.60) >DroSim_CAF1 11215 120 - 1 GAAUGAUGCUUAUGGAUGCCGUUCGUAAGCGGCAAAAUCAAUUGAUUGCGGAAUACAACCGUUUGCCACUUUCCGUGGGCACAUUGCGAGUCCGGAAUCUCAAGAGACAACUGGAGCAGC .((((.((((((((((((((((......)))))).((((....))))((((.......)))).........))))))))))))))((...(((((..(((....)))...)))))...)) ( -37.90) >DroEre_CAF1 11792 120 - 1 GAAUGCUCCUCAUGGAUGCCGCCCGUAAGCGGCAAAACCAAUUGAUUGCCGAAUACAACCGCUUGCCGCUUUCCAUGGGCACAUUGCGAGUCCGGAAUCUCAAGAGACAACUGGAGCAGC ...((((((...((((((..(((((((((((((((......((((((....))).)))....)))))))))...)))))).))))..(((........)))......))...)))))).. ( -37.90) >DroYak_CAF1 11429 120 - 1 GAAUGCUGCUUAUGGAGUCCGCCCGUAAGCGGCAAAACCAAUUAAUUGCCGAAUACAACCGUUUGCCGCUUUCCAUGGGCACAUUGCGAGUCCGGAAUCUUAAGAGACAAUUGGAGCAGC ....(((((((.....((((((((((((((((((((..(((....))).((........))))))))))))...))))))..(((.((....)).))).....).))).....))))))) ( -38.40) >consensus GAAUGAUGCUUAUGGAUGCCGUCCGUAAGCGGCAAAAUCAAUUGAUUGCGGAAUACAACCGUUUGCCACUUUCCGUGGGCACAUUGCGAGUCCGGAAUCUCAAGAGACAACUGGAGCAGC .((((.((((((((((((((((......)))))).............((((.......)))).........))))))))))))))((...(((((..(((....)))...)))))...)) (-30.34 = -31.14 + 0.80)

| Location | 13,398,724 – 13,398,844 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.33 |
| Mean single sequence MFE | -38.28 |
| Consensus MFE | -18.72 |
| Energy contribution | -19.52 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.49 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685787 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
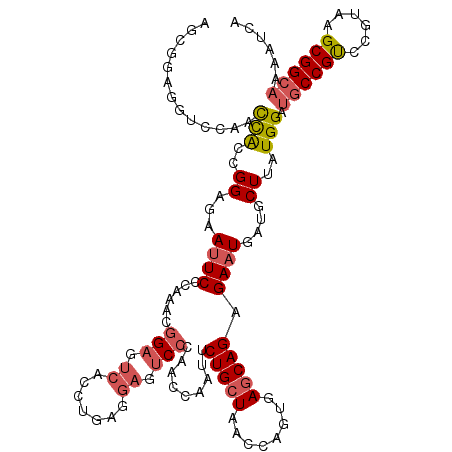
>3L_DroMel_CAF1 13398724 120 - 23771897 AGCGGAAGUCCAACUACCGGAGAAUUCUCAAACGGAUUCACCUGAGGAAUCCCAACCUAUUCUGCUAACCAGUGAGCAGAGAAUGGUGCUAAUGGAUGCCGUCCGUAAGCGGCAAAAUCA (((((.((.....)).))((.((.((((((...((.....)))))))).)))).(((..(((((((........)))))))...))))))......((((((......))))))...... ( -35.50) >DroSec_CAF1 14316 108 - 1 ------------ACCGGCGGAAAAUUCCCAAACGGAGUCACCUGAGGAUUCGCAACCAAUUCUGCUAACCAGUGAACAGAGAAUGAUGCUUAUGGAUGCCGUCCGUAAGCGGCAAAAUCA ------------...(((((((.(((((.....))))).......((........))..)))))))............((...((.(((((((((((...))))))))))).))...)). ( -28.20) >DroSim_CAF1 11295 111 - 1 AGCGGAGGUUCAACCGGCGGAGAAUUCCCAAACGG---------AGGAUUCCCAGCCAAUUCUGCUAAACAGUGAGCAGAGAAUGAUGCUUAUGGAUGCCGUUCGUAAGCGGCAAAAUCA ((((((((.....))(((((.((((((((....))---------.)))))))).)))...)))))).....(((((((........)))))))...((((((......))))))...... ( -41.80) >DroEre_CAF1 11872 120 - 1 AGCUGAGGUCCAACCACAGGAGAACUCCCAGACGGAGUCAGGUGAGGAGUCCCAGCCCAUCCUGCUUACCAGUGAGCAGAGAAUGCUCCUCAUGGAUGCCGCCCGUAAGCGGCAAAACCA ..(((.((.....)).)))..((.((((.....))))))..(((((((((.........(((((((((....))))))).))..)))))))))((.((((((......))))))...)). ( -44.20) >DroYak_CAF1 11509 120 - 1 AGCGGAGAUCCAAUCACCGGAGAAUUCUCAGGCGGAGUCACAUGAGGAGUCCCAACCCAUCCUGCUUUCCAGUGAGCAGAGAAUGCUGCUUAUGGAGUCCGCCCGUAAGCGGCAAAACCA .(.(((..(((..((.((.(((....))).)).))..((....))))).))))......((((((((......)))))).)).((((((((((((.(....)))))))))))))...... ( -41.70) >consensus AGCGGAGGUCCAACCACCGGAGAAUUCCCAAACGGAGUCACCUGAGGAGUCCCAACCAAUUCUGCUAACCAGUGAGCAGAGAAUGAUGCUUAUGGAUGCCGUCCGUAAGCGGCAAAAUCA .............(((..((...((((......(((.((.......)).))).........(((((........))))).))))....))..))).((((((......))))))...... (-18.72 = -19.52 + 0.80)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:33:00 2006