
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,389,308 – 13,389,468 |
| Length | 160 |
| Max. P | 0.696463 |

| Location | 13,389,308 – 13,389,428 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.67 |
| Mean single sequence MFE | -37.10 |
| Consensus MFE | -27.43 |
| Energy contribution | -27.77 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.555144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13389308 120 + 23771897 AUGUGGUGAAGCUUAACUGCCAGCCACGUGGCUUGGCCAUUUUGCGCAACGAGAACAUCAUUGCACUUGCUUGCAUCAAAGAGUUGACUCUCGUGCAGGACCAGAAGAAGAUCUUUUCGC ...((((...((..((..(((((((....))).))))..))..))((((.((.....)).)))).....(((((((...((((....)))).)))))))))))(((((.....))))).. ( -36.80) >DroVir_CAF1 2515 120 + 1 ACGUAGUCAAGCUGAAUUGCCAGCCACGCGGCUUGGCCAUCUUACGGAGUGAGAAUAUCAUUGCCUUGGCUUGCAUUAAGGAGCUUACUUUGGUGCAGGGUCAAAAGAAGAUAUUCUCGC .(((((((((((((....(....)....))))))))).....))))..(((((((((((.((...((((((((((((((((......)))))))))).))))))..)).))))))))))) ( -45.90) >DroGri_CAF1 2370 120 + 1 ACGUGGUCAAGCUCAACUGUCAGCCACGCGGCUUGGCCAUCUUGCGCAACGAGAACAUCAUUGCAUUGGCCUGCAUUAAGGAGCUAACAUUGGUGCAGGAUCAGAAGAAGAUAUUCUCGC ..(((((((((((....(((.....))).))))))))))).........((((((.(((.((...(((((((((((((((.......).)))))))))).))))..)).))).)))))). ( -39.70) >DroWil_CAF1 3274 120 + 1 AUGUGGUUAAAUUGAAUUGCCAGCCACGCGGUUUGGCCAUCUUGCGCAAUGAGAAUAUAAUUGCAUUGGCAUGCAUUAAGGAGUUGACUUUGGUUCAAGAUCAAAAGAAAAUCUUCUCAU ..((((((((((((....(....)....))))))))))))........((((((.......(((((....)))))....(((......(((((((...)))))))......))))))))) ( -23.90) >DroMoj_CAF1 2365 120 + 1 ACGUGGUCAAGUUGAACUGCCAGCCUCGCGGCUUGGCCAUCUUGCGUAGUGAAAACAUAAUCGCUGUGGCGUGCAUUAAGGACAUCACCCUGGUGCAGGGCCAGAAGAAGAUCUUCUCGC ..((((((((((((....(....)....))))))))))))...(((((((((........))))))((((.(((((((.((......)).))))))).).)))((((.....)))).))) ( -39.90) >DroAna_CAF1 10350 120 + 1 ACGUGGUCAAGCUUAACUGUCAGCCGCGCGGUCUGGCCAUCUUGCGCAGCGAAAACAUAAUCGCCCUAGCCUGUAUCAAGGAGGUGACCCUUGUGCAGGAUCAGAAAAAGAUAUUCUCGC .((((((...((......))..)))))).((.....))(((((...(.((((........)))).....((((((.(((((.(....))))))))))))....)...)))))........ ( -36.40) >consensus ACGUGGUCAAGCUGAACUGCCAGCCACGCGGCUUGGCCAUCUUGCGCAACGAGAACAUAAUUGCAUUGGCCUGCAUUAAGGAGCUGACUCUGGUGCAGGAUCAGAAGAAGAUAUUCUCGC ..(((((((((((................)))))))))))........(((((((.((..((...(((((((((((((.((......)).))))))))).))))..))..)).))))))) (-27.43 = -27.77 + 0.34)


| Location | 13,389,348 – 13,389,468 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.17 |
| Mean single sequence MFE | -34.37 |
| Consensus MFE | -22.44 |
| Energy contribution | -23.08 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.696463 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13389348 120 + 23771897 UUUGCGCAACGAGAACAUCAUUGCACUUGCUUGCAUCAAAGAGUUGACUCUCGUGCAGGACCAGAAGAAGAUCUUUUCGCUGCCCAUUAAGUACGAGGCUAGCUCGAUUGCCGUUAAUGC .....(((.((((((.(((.((...((.((((((((...((((....)))).))))))).).))..)).))).)))))).))).(((((((((((((.....))))..)))..)))))). ( -30.90) >DroVir_CAF1 2555 120 + 1 CUUACGGAGUGAGAAUAUCAUUGCCUUGGCUUGCAUUAAGGAGCUUACUUUGGUGCAGGGUCAAAAGAAGAUAUUCUCGCUGCCCAUCAAAUACGAAGCCAGCUCUAUUGCAGCCAAUGC .....((((((((((((((.((...((((((((((((((((......)))))))))).))))))..)).)))))))))))).)).............((..((......)).))...... ( -38.30) >DroGri_CAF1 2410 120 + 1 CUUGCGCAACGAGAACAUCAUUGCAUUGGCCUGCAUUAAGGAGCUAACAUUGGUGCAGGAUCAGAAGAAGAUAUUCUCGCUGCCCAUCAAAUACGAGGCCAGCUCCAUAGCGGCCAAUGC ..((.(((.((((((.(((.((...(((((((((((((((.......).)))))))))).))))..)).))).)))))).))).))..........((((.(((....)))))))..... ( -40.80) >DroWil_CAF1 3314 120 + 1 CUUGCGCAAUGAGAAUAUAAUUGCAUUGGCAUGCAUUAAGGAGUUGACUUUGGUUCAAGAUCAAAAGAAAAUCUUCUCAUUGCCCAUUAAAUACGAGGCCAGUUCGAUAGCCACCAAUGC ..((.(((((((((.......(((((....)))))....(((......(((((((...)))))))......)))))))))))).))..........(((.(......).)))........ ( -24.30) >DroMoj_CAF1 2405 120 + 1 CUUGCGUAGUGAAAACAUAAUCGCUGUGGCGUGCAUUAAGGACAUCACCCUGGUGCAGGGCCAGAAGAAGAUCUUCUCGCUGCCCAUCAAAUACGAGGCCAGCUCCAUAGCCACAAACUC ...(((..(((....)))...)))((((((.(((((((.((......)).)))))))(((..(((((.....))))).((((.((.((......)))).)))))))...))))))..... ( -38.30) >DroAna_CAF1 10390 120 + 1 CUUGCGCAGCGAAAACAUAAUCGCCCUAGCCUGUAUCAAGGAGGUGACCCUUGUGCAGGAUCAGAAAAAGAUAUUCUCGCUUCCAAUCAAGUACGAAGCCAGCUCAAUUGCUGUCAACGC ...(((..((((........)))).((..((((((.(((((.(....))))))))))))...)).....((.......(((((...........)))))((((......))))))..))) ( -33.60) >consensus CUUGCGCAACGAGAACAUAAUUGCAUUGGCCUGCAUUAAGGAGCUGACUCUGGUGCAGGAUCAGAAGAAGAUAUUCUCGCUGCCCAUCAAAUACGAGGCCAGCUCCAUAGCCGCCAAUGC .....((((((((((.((..((...(((((((((((((.((......)).))))))))).))))..))..)).))))))))))..................((......))......... (-22.44 = -23.08 + 0.64)
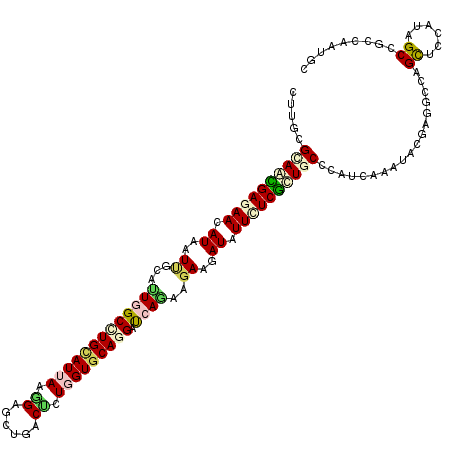


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:50 2006