
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,378,977 – 13,379,071 |
| Length | 94 |
| Max. P | 0.800823 |

| Location | 13,378,977 – 13,379,071 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.94 |
| Mean single sequence MFE | -28.77 |
| Consensus MFE | -21.07 |
| Energy contribution | -22.27 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.732598 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
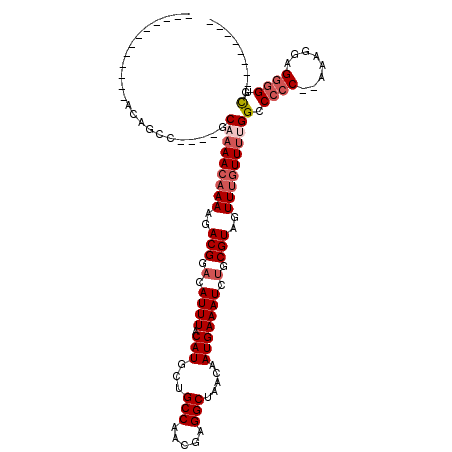
>3L_DroMel_CAF1 13378977 94 + 23771897 ------------ACAGCC----GCAAAACAAAAGACGGACAUUUACAUGCUGCCAACGAGGCUAACAAUGAAAUCAGCGUAGUUUGUUUUGGCCCCC--AUAGGAGGGGUCG-------- ------------......----..((((((((..(((...((((.(((...(((.....))).....)))))))...)))..))))))))(((((((--....).)))))).-------- ( -26.70) >DroSim_CAF1 50436 100 + 1 ------ACAGCCACAGCC----GCAAAACAAAAGACGGACAUUUACAUGCUGCCAACGAGGCUAACAAUGAAAUCUGCGUAGUUUGUUUUGGCCCCC--AAAGGAGGGGUCG-------- ------...((.......----))((((((((..(((.(.((((.(((...(((.....))).....))))))).).)))..))))))))(((((((--....).)))))).-------- ( -29.00) >DroEre_CAF1 50734 96 + 1 ------------GCAGCC----GCAAAACAAAAGACGGACAUUUACAUGCUGCCAACGAGGCUAACAAUGAAAUCUGCGUAGUUUGUUUUGGCCCCCCAAAAGGAGGGGUCG-------- ------------((....----))((((((((..(((.(.((((.(((...(((.....))).....))))))).).)))..))))))))((((((((....)).)))))).-------- ( -33.10) >DroYak_CAF1 52840 108 + 1 ACAGCCACAUCCACAGCC----ACAAAACAAAAGACGGACAUUUACAUGCUGCCAACGAGGCUAACAAUGAAAUCUGCGUAGUUCGUUUUGGCCCCCCAAAAGGAGGGGUCG-------- ..................----.((((((.((..(((.(.((((.(((...(((.....))).....))))))).).)))..)).))))))(((((((....)).)))))..-------- ( -28.20) >DroAna_CAF1 49869 99 + 1 ------ACAUGGUUGCUG----GCCAAACAAAAGACGGACAUUUACAUGCUGCCAUCGAGGCUAACAAUGAAAUCUGCGUAGUUUCUUGUGGCCCCC--AAAG-AGGGGGCG-------- ------...((((.....----)))).((((...(((.(.((((.(((...(((.....))).....))))))).).)))......)))).((((((--....-.)))))).-------- ( -28.20) >DroPer_CAF1 29165 106 + 1 ------------GCAGAAGCAAACGAAACAAAAGACGGACAUUUACAUGCUGCCAACGAGGCUAACAAUGAAAUCUGCGUAGUUUGUUUAGGCCCAC--AGAAUAGGCGGUGGGAGAGCG ------------......((...(.(((((((..(((.(.((((.(((...(((.....))).....))))))).).)))..))))))).).(((((--.(......).)))))...)). ( -27.40) >consensus ____________ACAGCC____GCAAAACAAAAGACGGACAUUUACAUGCUGCCAACGAGGCUAACAAUGAAAUCUGCGUAGUUUGUUUUGGCCCCC__AAAGGAGGGGUCG________ .......................(((((((((..(((.(.((((.(((...(((.....))).....))))))).).)))..)))))))))(.((((........)))).)......... (-21.07 = -22.27 + 1.19)


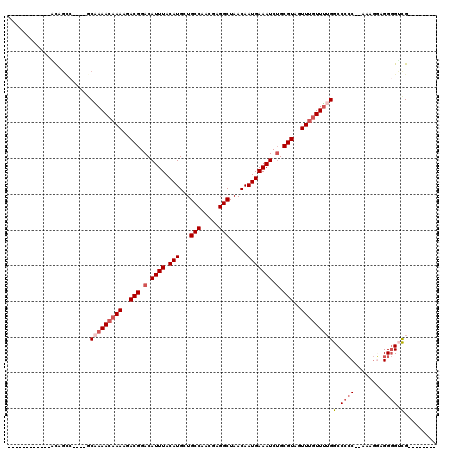
| Location | 13,378,977 – 13,379,071 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.94 |
| Mean single sequence MFE | -30.60 |
| Consensus MFE | -23.23 |
| Energy contribution | -23.73 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.800823 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
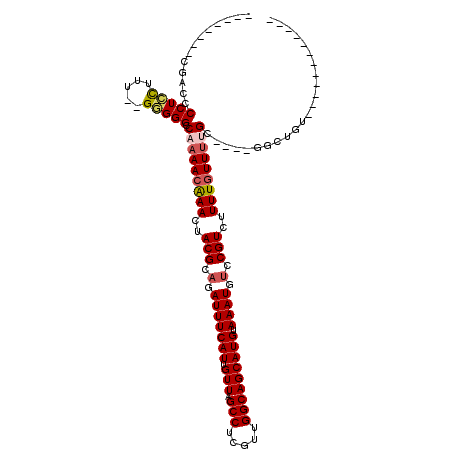
>3L_DroMel_CAF1 13378977 94 - 23771897 --------CGACCCCUCCUAU--GGGGGCCAAAACAAACUACGCUGAUUUCAUUGUUAGCCUCGUUGGCAGCAUGUAAAUGUCCGUCUUUUGUUUUGC----GGCUGU------------ --------...((((......--))))(((((((((((..(((...(((((((.(((.(((.....))))))))).))))...)))..))))))))..----)))...------------ ( -28.90) >DroSim_CAF1 50436 100 - 1 --------CGACCCCUCCUUU--GGGGGCCAAAACAAACUACGCAGAUUUCAUUGUUAGCCUCGUUGGCAGCAUGUAAAUGUCCGUCUUUUGUUUUGC----GGCUGUGGCUGU------ --------...((((......--))))..(((((((((..(((.(.(((((((.(((.(((.....))))))))).)))).).)))..)))))))))(----(((....)))).------ ( -33.00) >DroEre_CAF1 50734 96 - 1 --------CGACCCCUCCUUUUGGGGGGCCAAAACAAACUACGCAGAUUUCAUUGUUAGCCUCGUUGGCAGCAUGUAAAUGUCCGUCUUUUGUUUUGC----GGCUGC------------ --------...((((.......))))((((((((((((..(((.(.(((((((.(((.(((.....))))))))).)))).).)))..))))))))..----))))..------------ ( -31.90) >DroYak_CAF1 52840 108 - 1 --------CGACCCCUCCUUUUGGGGGGCCAAAACGAACUACGCAGAUUUCAUUGUUAGCCUCGUUGGCAGCAUGUAAAUGUCCGUCUUUUGUUUUGU----GGCUGUGGAUGUGGCUGU --------....((((((....)))))).(((((((((..(((.(.(((((((.(((.(((.....))))))))).)))).).)))..))))))))).----(((..(....)..))).. ( -35.40) >DroAna_CAF1 49869 99 - 1 --------CGCCCCCU-CUUU--GGGGGCCACAAGAAACUACGCAGAUUUCAUUGUUAGCCUCGAUGGCAGCAUGUAAAUGUCCGUCUUUUGUUUGGC----CAGCAACCAUGU------ --------.((((((.-....--)))))).(((.((((((....)).)))).(((((.(((.(((.(((.((((....))))..)))..)))...)))----.)))))...)))------ ( -29.90) >DroPer_CAF1 29165 106 - 1 CGCUCUCCCACCGCCUAUUCU--GUGGGCCUAAACAAACUACGCAGAUUUCAUUGUUAGCCUCGUUGGCAGCAUGUAAAUGUCCGUCUUUUGUUUCGUUUGCUUCUGC------------ .((...(((((.(......).--)))))...(((((((..(((.(.(((((((.(((.(((.....))))))))).)))).).)))..))))))).....))......------------ ( -24.50) >consensus ________CGACCCCUCCUUU__GGGGGCCAAAACAAACUACGCAGAUUUCAUUGUUAGCCUCGUUGGCAGCAUGUAAAUGUCCGUCUUUUGUUUUGC____GGCUGU____________ .............(((((.....))))).(((((((((..(((.(.(((((((.(((.(((.....))))))))).)))).).)))..)))))))))....................... (-23.23 = -23.73 + 0.50)


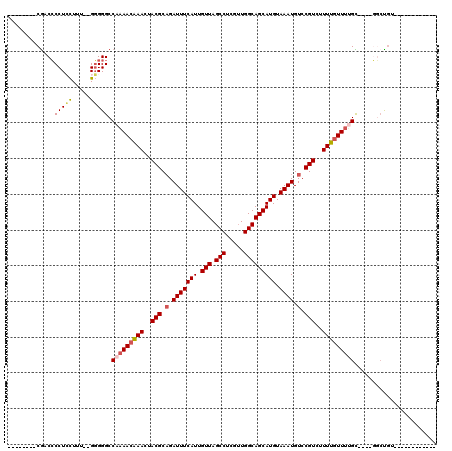
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:37 2006