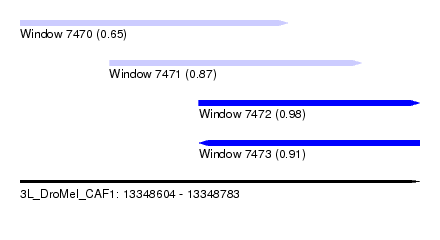

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,348,604 – 13,348,783 |
| Length | 179 |
| Max. P | 0.976499 |

| Location | 13,348,604 – 13,348,724 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.18 |
| Mean single sequence MFE | -20.95 |
| Consensus MFE | -18.01 |
| Energy contribution | -18.40 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.654349 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13348604 120 + 23771897 UGCUGACAUGACAUGAACUGUAGUUUAUUUAUUAUUUCUCUGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGU .....((((.(((.((...(((((......)))))...)))))(((.............)))..........................(((((((((((....))))))))))))))).. ( -21.02) >DroSec_CAF1 20221 120 + 1 UGCUGACAUGACAUGAACUGUAGUUUAUUUAUUAUUUCUCUGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGU .....((((.(((.((...(((((......)))))...)))))(((.............)))..........................(((((((((((....))))))))))))))).. ( -21.02) >DroSim_CAF1 19995 120 + 1 UGCUGAAAUGACAUGAACUGUAGUUUAUUUAUUAUUUCUCUGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUAU (((((((((((.((((((....))))))...............(((.............)))......))))))))............(((((((((((....)))))))))))..))). ( -24.52) >DroEre_CAF1 20496 117 + 1 UGCUGACAUGACAUGAACUGUAGUUUAUUUAUUAUUUCUCUGUGCCUCAUUCUUCUUCUGGCUCCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGCA--- .(((((.((((.((((((....))))))...............(((.............)))......)))).)))............(((((((((((....))))))))))))).--- ( -22.52) >DroYak_CAF1 21285 117 + 1 UGCUGACAUGACAUGAGCUGUAGUUUAUUUAUUAUUUCUCUGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUU--- ...(((.((((((.(((..(((((......)))))..))))))(((.............))).......))).)))............(((((((((((....)))))))))))...--- ( -23.62) >DroAna_CAF1 19271 111 + 1 UACUGACAUGACAUGAACUGUAGUUUAUUUAUUAUUUCUCUGUGCCUCGGUCUCCAUCUUGCUUCCCUCCAUUUUAUUUAUAUCCUUAACGCAUGCAAACAU--UUGUAUGUC------- .(((((....(((.((...(((((......)))))...)))))...))))).......................................((((((((....--)))))))).------- ( -13.00) >consensus UGCUGACAUGACAUGAACUGUAGUUUAUUUAUUAUUUCUCUGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUG___ .((.((.((((.((((((....))))))...)))).))...))(((.............)))..........................(((((((((((....)))))))))))...... (-18.01 = -18.40 + 0.39)


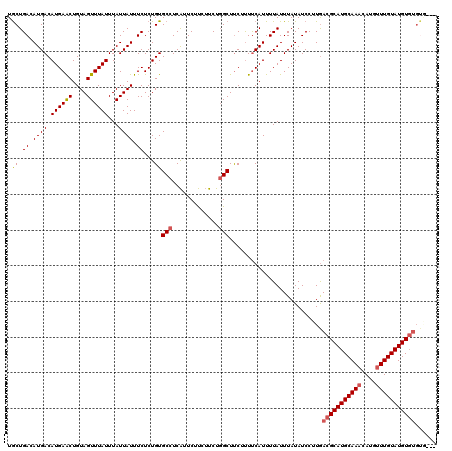
| Location | 13,348,644 – 13,348,757 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 82.54 |
| Mean single sequence MFE | -25.86 |
| Consensus MFE | -16.73 |
| Energy contribution | -16.98 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.870518 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13348644 113 + 23771897 UGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGUGUGCGUGUGCGAAUCGGGUCAUGCACUCACACA-- ...(((.............)))..........................(((((((((((....)))))))))))(((((.((((((((.((...)).).))))))).))))).-- ( -35.92) >DroSec_CAF1 20261 107 + 1 UGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGUG------GGCGAAUCGGGUCAUACACUCACACA-- ...(((.............)))..........................(((((((((((....)))))))))))(((((((------(.(......).)))))))).......-- ( -27.22) >DroSim_CAF1 20035 109 + 1 UGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUAUG------UGCGAAUCGGGUCAUACACUCACACACA ...(((.............)))..........................(((((((((((....)))))))))))(((((((------(.((...)).).)))))))......... ( -24.92) >DroAna_CAF1 19311 99 + 1 UGUGCCUCGGUCUCCAUCUUGCUUCCCUCCAUUUUAUUUAUAUCCUUAACGCAUGCAAACAU--UUGUAUGUC--------------UGCGAAUCAGGUCAUACAUAUAUGUAUG ...((((.((..................))...................((((.(((.((..--..)).))).--------------))))....))))((((((....)))))) ( -15.37) >consensus UGUGCCUCAUUCUUCUUCUGGCUUCUUUUCAUUUUAUUUAUAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGUG______UGCGAAUCGGGUCAUACACUCACACA__ (((((((.((((..........................((......))(((((((((((....)))))))))))................)))).))).))))............ (-16.73 = -16.98 + 0.25)


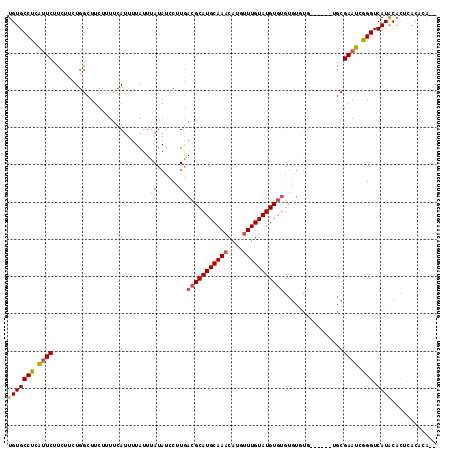
| Location | 13,348,684 – 13,348,783 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 77.48 |
| Mean single sequence MFE | -29.92 |
| Consensus MFE | -13.79 |
| Energy contribution | -14.23 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.15 |
| Mean z-score | -3.27 |
| Structure conservation index | 0.46 |
| SVM decision value | 1.77 |
| SVM RNA-class probability | 0.976499 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13348684 99 + 23771897 UAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGUGUGCGUGUGCGAAUCGGGUCAUGCACUCACACA--CACUCAUUUGUAU---GUCCUGUACUUUA ........(((((((((((....)))))))))))(((((((((.((((..((......))..)))).))))))--)))......((((---.....)))).... ( -36.10) >DroSec_CAF1 20301 93 + 1 UAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGUG------GGCGAAUCGGGUCAUACACUCACACA--CACACAUUUGUAU---GUCCUGUAGUUUA ..........(((((((((((....)))(((((((((((((------((.(.((......)).).))))))))--)))))))))))))---))........... ( -31.50) >DroSim_CAF1 20075 95 + 1 UAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUAUG------UGCGAAUCGGGUCAUACACUCACACACACACACAUUUGUAU---GUCCUGUAGUUUA ..........(((((((((..(((.(((.((((.(((((((------(.((...)).).))))))).)))).))).)))..)))))))---))........... ( -30.00) >DroAna_CAF1 19351 88 + 1 UAUCCUUAACGCAUGCAAACAU--UUGUAUGUC--------------UGCGAAUCAGGUCAUACAUAUAUGUAUGGCGCUGUUAGUACGUCGUCUUGUAGUUUA .........((((.(((.((..--..)).))).--------------))))...(((((((((((....)))))))).)))....((((......))))..... ( -22.10) >consensus UAUCCUUGACGCAUGCAAACAUGUUUGUAUGUGUGUGUGUG______UGCGAAUCGGGUCAUACACUCACACA__CACACAUUUGUAU___GUCCUGUAGUUUA ........(((((((((((....)))))))))))....................((((.(((((....................))))...).))))....... (-13.79 = -14.23 + 0.44)


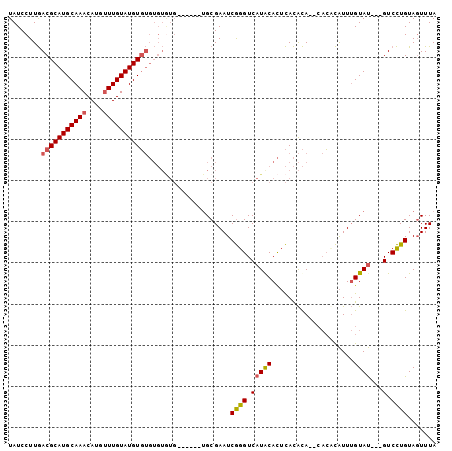
| Location | 13,348,684 – 13,348,783 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 77.48 |
| Mean single sequence MFE | -28.57 |
| Consensus MFE | -10.81 |
| Energy contribution | -12.62 |
| Covariance contribution | 1.81 |
| Combinations/Pair | 1.22 |
| Mean z-score | -3.69 |
| Structure conservation index | 0.38 |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.910795 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13348684 99 - 23771897 UAAAGUACAGGAC---AUACAAAUGAGUG--UGUGUGAGUGCAUGACCCGAUUCGCACACGCACACACACACACAUACAAACAUGUUUGCAUGCGUCAAGGAUA ..........(((---..........(((--((((((.((((.((............)).)))).))))))))).......((((....)))).)))....... ( -28.30) >DroSec_CAF1 20301 93 - 1 UAAACUACAGGAC---AUACAAAUGUGUG--UGUGUGAGUGUAUGACCCGAUUCGCC------CACACACACACAUACAAACAUGUUUGCAUGCGUCAAGGAUA ..........(((---.......((((((--((((((.(((..(((......)))..------))).))))))))))))..((((....)))).)))....... ( -30.70) >DroSim_CAF1 20075 95 - 1 UAAACUACAGGAC---AUACAAAUGUGUGUGUGUGUGAGUGUAUGACCCGAUUCGCA------CAUACACACACAUACAAACAUGUUUGCAUGCGUCAAGGAUA ..........(((---.......((((((((((((((.((((..((......)))))------).))))))))))))))..((((....)))).)))....... ( -31.60) >DroAna_CAF1 19351 88 - 1 UAAACUACAAGACGACGUACUAACAGCGCCAUACAUAUAUGUAUGACCUGAUUCGCA--------------GACAUACAA--AUGUUUGCAUGCGUUAAGGAUA .............((((((....(((.(.((((((....)))))).))))....(((--------------(((((....--)))))))).))))))....... ( -23.70) >consensus UAAACUACAGGAC___AUACAAAUGUGUG__UGUGUGAGUGUAUGACCCGAUUCGCA______CACACACACACAUACAAACAUGUUUGCAUGCGUCAAGGAUA ..........(((...((.(((((((((...((((((.(((..........................))).))))))...))))))))).))..)))....... (-10.81 = -12.62 + 1.81)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:10 2006