
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,111,525 – 13,111,672 |
| Length | 147 |
| Max. P | 0.998744 |

| Location | 13,111,525 – 13,111,622 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 97.53 |
| Mean single sequence MFE | -24.93 |
| Consensus MFE | -22.76 |
| Energy contribution | -23.36 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.21 |
| SVM RNA-class probability | 0.998744 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13111525 97 + 23771897 CCGUCUUUUGUCGCAUCGCAUCAAUUAAAGCAUUACAAUGUCAUAAUCAAAAUGACAAAUAGAAUUACGCAUGAUGGGAAAGGCAGCAGCCGACGGA (((((.....((.((((((..........)).......((((((.......))))))...............)))).))..(((....)))))))). ( -25.60) >DroSec_CAF1 99189 96 + 1 CCGUCUUUUGUCGCAUCGCAUCAAUUAAAGCAU-ACAAUGUCAUAAUCAAAAUGACAAAUAGAAUUACGCAUGAUGGGAAAGGCAGCAGCCGACGGA (((((..((((.((.((.(((((..........-....((((((.......))))))..............))))).))...)).))))..))))). ( -26.09) >DroSim_CAF1 96960 97 + 1 CCGUCUUUUGUCGCAUCGCAUCAAUUAAAGCAUUACAAUGUCAUAAUCAAAAUGACAAAUAGAAUUACGCAUGAUGGGAAAGGCAGCAGCCGACGGA (((((.....((.((((((..........)).......((((((.......))))))...............)))).))..(((....)))))))). ( -25.60) >DroEre_CAF1 95879 97 + 1 UCGCCUUUUGUCGCAUCGCAUCAAUUAAAGCAUUACAAUGUCAUAAUCAAAAUGACAAAUAGAAUUACGCAUGAUGGGAAAGGCGGCAGCCGACACA (((....(((((((.((.(((((...............((((((.......))))))..............))))).))...))))))).))).... ( -21.75) >DroYak_CAF1 99210 97 + 1 CCGUCUUUUGUCGCAUCGCAUCAAUUAAAGCAUUACAAUGUCAUAAUCAAAAUGACAAAUAGAAUUACGCAUGAUGGGAAAGGCAGCAGCCGACGGA (((((.....((.((((((..........)).......((((((.......))))))...............)))).))..(((....)))))))). ( -25.60) >consensus CCGUCUUUUGUCGCAUCGCAUCAAUUAAAGCAUUACAAUGUCAUAAUCAAAAUGACAAAUAGAAUUACGCAUGAUGGGAAAGGCAGCAGCCGACGGA (((((.....((.((((((..........)).......((((((.......))))))...............)))).))..(((....)))))))). (-22.76 = -23.36 + 0.60)

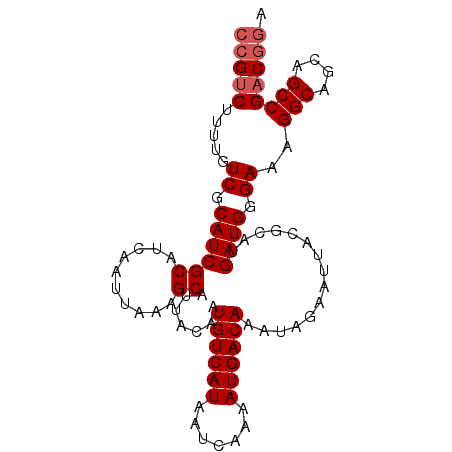

| Location | 13,111,525 – 13,111,622 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 97.53 |
| Mean single sequence MFE | -24.14 |
| Consensus MFE | -23.50 |
| Energy contribution | -23.38 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928217 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13111525 97 - 23771897 UCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUUUGUCAUUUUGAUUAUGACAUUGUAAUGCUUUAAUUGAUGCGAUGCGACAAAAGACGG .((((((((....)))..((.(((((.((((.((......((((((.......))))))..)).))))......))))).))..........))))) ( -24.60) >DroSec_CAF1 99189 96 - 1 UCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUUUGUCAUUUUGAUUAUGACAUUGU-AUGCUUUAAUUGAUGCGAUGCGACAAAAGACGG .((((((((....)))..((.(((((.(((((........((((((.......))))))...)-))))......))))).))..........))))) ( -25.80) >DroSim_CAF1 96960 97 - 1 UCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUUUGUCAUUUUGAUUAUGACAUUGUAAUGCUUUAAUUGAUGCGAUGCGACAAAAGACGG .((((((((....)))..((.(((((.((((.((......((((((.......))))))..)).))))......))))).))..........))))) ( -24.60) >DroEre_CAF1 95879 97 - 1 UGUGUCGGCUGCCGCCUUUCCCAUCAUGCGUAAUUCUAUUUGUCAUUUUGAUUAUGACAUUGUAAUGCUUUAAUUGAUGCGAUGCGACAAAAGGCGA ..((((...((.(((...((.(((((.((((.((......((((((.......))))))..)).))))......))))).)).))).))...)))). ( -21.10) >DroYak_CAF1 99210 97 - 1 UCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUUUGUCAUUUUGAUUAUGACAUUGUAAUGCUUUAAUUGAUGCGAUGCGACAAAAGACGG .((((((((....)))..((.(((((.((((.((......((((((.......))))))..)).))))......))))).))..........))))) ( -24.60) >consensus UCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUUUGUCAUUUUGAUUAUGACAUUGUAAUGCUUUAAUUGAUGCGAUGCGACAAAAGACGG .((((((((....)))..((.(((((.((((.((......((((((.......))))))..)).))))......))))).))..........))))) (-23.50 = -23.38 + -0.12)



| Location | 13,111,582 – 13,111,672 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -30.02 |
| Consensus MFE | -25.46 |
| Energy contribution | -25.94 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724797 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13111582 90 - 23771897 --UACUGGGUGAUGCGGA---UUGCCGGCUAUCAACCACCACAAGAGGAGUU----AGCUCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUU --...((((.((.((((.---..((((((..((...........))((((..----..))))))))))..)))).)).))))................. ( -28.60) >DroSec_CAF1 99245 90 - 1 --UGCUGGGUGAUGCGGA---UUGCCGGCUAUCAACCACCACAAGAGGAGUU----AGCUCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUU --((.((((.((.((((.---..((((((..((...........))((((..----..))))))))))..)))).)).)))).)).............. ( -29.50) >DroSim_CAF1 97017 90 - 1 --UGCUGGGUGAUGCGGA---UUGCCGGCUAUCAACCACCACAAGAGGAGUU----AGCUCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUU --((.((((.((.((((.---..((((((..((...........))((((..----..))))))))))..)))).)).)))).)).............. ( -29.50) >DroEre_CAF1 95936 99 - 1 GGUGCUGGGUGCUGCGGACUAUUGCCGGCUAUCAACCACCACAAGAGGAGUUACAUAGCUGUGUCGGCUGCCGCCUUUCCCAUCAUGCGUAAUUCUAUU .(((.((((....((((.(....((((((.........((......))((((....))))..)))))).)))))....)))).)))............. ( -28.30) >DroYak_CAF1 99267 95 - 1 AGUGAUGGGUGAUGCGGACUAUUGCCGGGUAUCAACCACCACAAGAGGAGUU----AGCUCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUU .((((((((.((.((((.(....(((((((....))).........((((..----..))))..)))).))))).)).))))))))............. ( -34.20) >consensus __UGCUGGGUGAUGCGGA___UUGCCGGCUAUCAACCACCACAAGAGGAGUU____AGCUCCGUCGGCUGCUGCCUUUCCCAUCAUGCGUAAUUCUAUU ..((.((((.((.((((......((((((..((...........))(((((......)))))))))))..)))).)).)))).)).............. (-25.46 = -25.94 + 0.48)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:26:33 2006