
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 13,015,589 – 13,015,793 |
| Length | 204 |
| Max. P | 0.721628 |

| Location | 13,015,589 – 13,015,695 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 83.33 |
| Mean single sequence MFE | -35.40 |
| Consensus MFE | -26.02 |
| Energy contribution | -25.77 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.721628 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
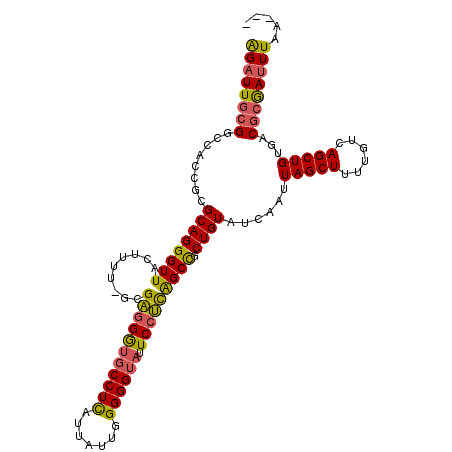
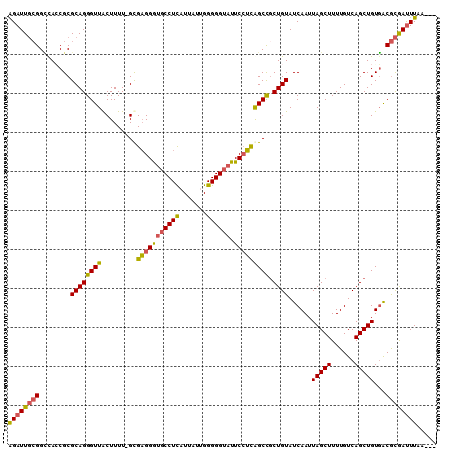
>3L_DroMel_CAF1 13015589 106 - 23771897 GGAUUGCGGUUACCGCGCAGGGUUACUUUU-GCGAGGAUGCCUUAUUAUCGGGGGUAUUCCUCAGCCCCUGUAUCAAUUAGCUUUUGUCAGCUGUGACGCGAUUUAA--- ((((((((.((((.((((((((........-(((((((((((((.......)))))).))))).))))))))..(((.......)))...)).))))))))))))..--- ( -36.40) >DroSec_CAF1 6658 106 - 1 AGAUUGCGGCCAGCGCGCAGGGUUACUUUU-GCGAGGGUUCCUCAUUAUUGGGGGUAUUCCUCAGCCGCUGUAUCAAUUAGCUUUUGUCAGCUGUGACGCGAUUUAA--- ((((((((..((((((((((((....))))-))(((((.((((((....))))))...))))).).)))))..(((..(((((......))))))))))))))))..--- ( -39.40) >DroSim_CAF1 6676 106 - 1 AGAUUGCGGCCACCGCGCAGGGUUACUUUU-GUGAGGGCGCCUCAUUAUUGGGGGUAUUCCUCAGCCGCUGUAUCAAUUAGCUUUUGUCAGCUGUGACGCGAUUUAU--- ((((((((..(((.((((((((((((....-))((((((.(((((....)))))))...)))))))).))))..(((.......)))...)).))).))))))))..--- ( -34.90) >DroYak_CAF1 6641 110 - 1 AGAUUCGGGGUAACGCGCAGGGUUGCUUUUUGUGUGGGUGCCUUUUUGUUGGGGGUAUUCAAUGGCUGCUGUAUCAAUUAGCUUUUGUCAGCUGUUGCAUAACUUAGACA ......((((((.(((((((((.....)))))))))..))))))..((((..((.(((.((((((((((.((........))....).))))))))).))).))..)))) ( -30.90) >consensus AGAUUGCGGCCACCGCGCAGGGUUACUUUU_GCGAGGGUGCCUCAUUAUUGGGGGUAUUCCUCAGCCGCUGUAUCAAUUAGCUUUUGUCAGCUGUGACGCGAUUUAA___ ((((((((........((((((((.........(((((((((((.......)))))).))))))))).))))......(((((......)))))...))))))))..... (-26.02 = -25.77 + -0.25)

| Location | 13,015,695 – 13,015,793 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 88.77 |
| Mean single sequence MFE | -31.07 |
| Consensus MFE | -21.98 |
| Energy contribution | -22.70 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.560176 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 13015695 98 - 23771897 UACACAUACCGUCCGUCGGGCAAUACAACAGGAAAAGGAAAAGGCAGAUUUGC--GGGCGGUGUGGCCAGCGGGCAGUUGGCGUCGAAUAACUAAAUACU ....((((((((((((...((.......(.......)......))......))--))))))))))((((((.....)))))).................. ( -31.02) >DroSec_CAF1 6764 98 - 1 UACACAUACCGUCCGUCGGGCAAUACAACAGGAAAAGGAAAAGGCAGAUUUGC--GGGCGGUGUGGCCAGCGGGCAGUUGGCGCCGAAUAACUAAAUACU ....((((((((((((...((.......(.......)......))......))--))))))))))((((((.....)))))).................. ( -31.02) >DroSim_CAF1 6782 100 - 1 UACAACUGCUAGGCCUCGGGCAAUACAACAGGAAAAGGAAAAGGCAGACUUGGAGGGGGGGUGUGGCCAGCGGGCAGUUGGCGUCGAAUAACUAAAUACU ..((((((((..((....(((.((((..(.....(((...........)))......)..)))).))).)).)))))))).................... ( -25.30) >DroEre_CAF1 9086 98 - 1 UACACAUACCGUCCGUCGGGCAAUACAACAGGAAAAGGAAAAGGCCGAUUUUC--GGACGGUGUGGCCAGCAGGCGGUUGGCGCGAAAUGAUUAAAUACU ....(((((((((((((((.(.....................).))))....)--))))))))))((((((.....)))))).................. ( -34.20) >DroYak_CAF1 6751 98 - 1 UACACAUACCGUCCGUCGGGCAAUACAACAGGAAAAGGAAAAGGCCGAUUUUC--GGACGGUGUGGCCAGCCGGCAGUUGGCGCGAAAUGAUUAAAUACU ....(((((((((((((((.(.....................).))))....)--))))))))))((((((.....)))))).................. ( -33.80) >consensus UACACAUACCGUCCGUCGGGCAAUACAACAGGAAAAGGAAAAGGCAGAUUUGC__GGGCGGUGUGGCCAGCGGGCAGUUGGCGCCGAAUAACUAAAUACU ....(((((((((((((..((.......(.......)......)).)))......))))))))))((((((.....)))))).................. (-21.98 = -22.70 + 0.72)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:25:33 2006