| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
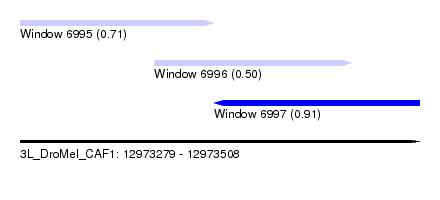
| Location | 12,973,279 – 12,973,508 |
| Length | 229 |
| Max. P | 0.906086 |

| Location | 12,973,279 – 12,973,390 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.35 |
| Mean single sequence MFE | -35.42 |
| Consensus MFE | -33.90 |
| Energy contribution | -33.66 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.713923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12973279 111 + 23771897 AAACCGAAGGAUAGUCGGACAUACCGCCGGCGGUUUUGCUCUGAAGUGCGCUUUAAUAGUUGGCUGUCUGCAGCUGG---------CGCAGCUGGGAAAUCGGAAAAGCGAAAAAGCGAA ...((((.(((((((((..((.((((....))))..))..)(((((....)))))......)))))))).((((((.---------..)))))).....))))....((......))... ( -35.40) >DroSec_CAF1 90663 103 + 1 AAACCGAAGGAUAGUCGGACAUACCGCCGGCGGUUUUGCUCUGAAGUGCGCUUUAAUAGUUGGCUGUCUGCAGCUGG---------CGCAGCUGGGAAAUCGGAAAAG--------CGAA ...((((.(((((((((..((.((((....))))..))..)(((((....)))))......)))))))).((((((.---------..)))))).....)))).....--------.... ( -33.30) >DroSim_CAF1 100356 103 + 1 AAACCGAAGGAUAGUCGGACAUACCGCCGGCGGUUUUGCUCUGAAGUGCGCUUUAAUAGUUGGCUGUCUGCAGCUGG---------CGCAGCUGGGAAAUCGGAAAAG--------CGAA ...((((.(((((((((..((.((((....))))..))..)(((((....)))))......)))))))).((((((.---------..)))))).....)))).....--------.... ( -33.30) >DroEre_CAF1 96034 112 + 1 AAACCGAAGGAUAGUCAGACAUACCGCCGGCGGUUUUGCUCUGAAGUGCGCUUUAAUAGUUGGCUGUCUGCAGCUGCCGCAGCUGCCGCAGCUGGGAAAUCGGAAAAG--------CGAA ...((((.(((((((((((((.((((....))))..)).))(((((....))))).....))))))))).(((((((.((....)).))))))).....)))).....--------.... ( -41.70) >DroYak_CAF1 94598 103 + 1 AAACCGAAGGAUAGUCAGACAUACCGCCGGCGGUUUUGCUCUGAAGUGCGCUUUAAUAGUUGGCUGUCUGCAGCUGC---------CGCAGCUGGGAAAUCGGAAAAG--------CGCA ...((((.(((((((((((((.((((....))))..)).))(((((....))))).....))))))))).((((((.---------..)))))).....)))).....--------.... ( -33.40) >consensus AAACCGAAGGAUAGUCGGACAUACCGCCGGCGGUUUUGCUCUGAAGUGCGCUUUAAUAGUUGGCUGUCUGCAGCUGG_________CGCAGCUGGGAAAUCGGAAAAG________CGAA ...((((.(((((((((((((.((((....))))..)).))(((((....))))).....))))))))).((((((............)))))).....))))................. (-33.90 = -33.66 + -0.24)



| Location | 12,973,356 – 12,973,469 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.02 |
| Mean single sequence MFE | -33.31 |
| Consensus MFE | -27.20 |
| Energy contribution | -27.44 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12973356 113 + 23771897 ------CGCAGCUGGGAAAUCGGAAAAGCGAAAAAGCGAAA-AGCAACAGCACCGUGGCUGCGGCAUUUGAAUGUGGCGUGUCGGCAGGCCAGAGUGCUGUUUAAUAAAAUGGUGGGGAA ------(((..((((....))))....))).....((....-.))((((((((..(((((((.((((....)))).))((....)).)))))..)))))))).................. ( -33.40) >DroSec_CAF1 90740 105 + 1 ------CGCAGCUGGGAAAUCGGAAAAG--------CGAAA-AGCAACAGCACCGUGGCUGCGGCAUUUGAAUGUGGCGUGUCGGCAGGCCAGAGUGCUGUUUAAUAAAAUGGUGGGGCA ------.((.(((..............(--------(....-.))((((((((..(((((((.((((....)))).))((....)).)))))..)))))))).........)))...)). ( -32.70) >DroSim_CAF1 100433 105 + 1 ------CGCAGCUGGGAAAUCGGAAAAG--------CGAAA-AGCAACAGCACCGUGGCUGCGGCAUUUGAAUGUGGCGUGUCGGCAGGCCAGAGUGCUGUUUAAUAAAAUGGUGGGGCA ------.((.(((..............(--------(....-.))((((((((..(((((((.((((....)))).))((....)).)))))..)))))))).........)))...)). ( -32.70) >DroEre_CAF1 96114 108 + 1 AGCUGCCGCAGCUGGGAAAUCGGAAAAG--------CGAAAAUGCAACAGCACCGU----GCGGCAUUUGAAUGUGGCGUGUCGGCAGACCAGAGUGCUGUUUAAUAAAAUGGUGGGGCA .(((.((((..((((....))))....(--------(......))(((((((((..----((.((((....)))).))(.(((....)))).).))))))))..........))))))). ( -32.50) >DroYak_CAF1 94675 106 + 1 ------CGCAGCUGGGAAAUCGGAAAAG--------CGCAAAUGCAACAGCACCGUGGCUGCGGCAUUUGAAUGUGGCGUGUCGGCAGGCCAAAGUGCUGUUUAAUAAAAUGGUGGGGCA ------.((.((((........(((.((--------(((((((((..((((......))))..)))))).....((((.((....)).))))..))))).))).......))))...)). ( -35.26) >consensus ______CGCAGCUGGGAAAUCGGAAAAG________CGAAA_AGCAACAGCACCGUGGCUGCGGCAUUUGAAUGUGGCGUGUCGGCAGGCCAGAGUGCUGUUUAAUAAAAUGGUGGGGCA .......((..((((....))))...........................((((((....(((((((((.....((((.((....)).)))))))))))))........))))))..)). (-27.20 = -27.44 + 0.24)



| Location | 12,973,390 – 12,973,508 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.47 |
| Mean single sequence MFE | -31.48 |
| Consensus MFE | -28.52 |
| Energy contribution | -29.32 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.906086 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12973390 118 - 23771897 CAGUUCGUGUUGCAGAUCCAAUGUGCCAUGACUUCUGUG-UUCCCCACCAUUUUAUUAAACAGCACUCUGGCCUGCCGACACGCCACAUUCAAAUGCCGCAGCCACGGUGCUGUUGCU-U (((.(((((..(((.......)))..)))))...)))..-..................(((((((((.((((.(((.(.((.............))).))))))).)))))))))...-. ( -31.92) >DroSec_CAF1 90766 118 - 1 CAGUUCGUGUUGCAGAUCCAAUGUGCCAUGACUUCUGUU-UGCCCCACCAUUUUAUUAAACAGCACUCUGGCCUGCCGACACGCCACAUUCAAAUGCCGCAGCCACGGUGCUGUUGCU-U (((.(((((..(((.......)))..)))))...)))..-..................(((((((((.((((.(((.(.((.............))).))))))).)))))))))...-. ( -31.92) >DroSim_CAF1 100459 118 - 1 CAGUUCGUGUUGCAGAUCCAAUGUGCCAUGACUUCUGUU-UGCCCCACCAUUUUAUUAAACAGCACUCUGGCCUGCCGACACGCCACAUUCAAAUGCCGCAGCCACGGUGCUGUUGCU-U (((.(((((..(((.......)))..)))))...)))..-..................(((((((((.((((.(((.(.((.............))).))))))).)))))))))...-. ( -31.92) >DroEre_CAF1 96146 116 - 1 CAGUUCGUGUUGCAGCUCCAAUGUGCCAUGACUUCUGUUUUGCCCCACCAUUUUAUUAAACAGCACUCUGGUCUGCCGACACGCCACAUUCAAAUGCCGC----ACGGUGCUGUUGCAUU ..((.((((((((((..(((..((((((.(((....))).))........(((....)))..))))..))).))).)))))))..)).....(((((.((----(......))).))))) ( -27.60) >DroYak_CAF1 94701 119 - 1 CAGUUCGUGUUGCAGCUCCAAUGUGCCAUGACUUCUGUU-UGCCCCACCAUUUUAUUAAACAGCACUUUGGCCUGCCGACACGCCACAUUCAAAUGCCGCAGCCACGGUGCUGUUGCAUU ..((.((((((((((..((((.((((.........((((-((..............)))))))))).)))).))).)))))))..)).....(((((.(((((......))))).))))) ( -34.04) >consensus CAGUUCGUGUUGCAGAUCCAAUGUGCCAUGACUUCUGUU_UGCCCCACCAUUUUAUUAAACAGCACUCUGGCCUGCCGACACGCCACAUUCAAAUGCCGCAGCCACGGUGCUGUUGCU_U (((.(((((..(((.......)))..)))))...))).....................(((((((((.((((.(((.(.((.............))).))))))).)))))))))..... (-28.52 = -29.32 + 0.80)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:24:32 2006