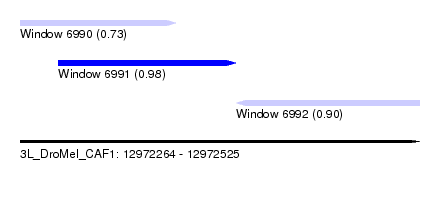

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 12,972,264 – 12,972,525 |
| Length | 261 |
| Max. P | 0.984856 |

| Location | 12,972,264 – 12,972,366 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.17 |
| Mean single sequence MFE | -30.25 |
| Consensus MFE | -23.38 |
| Energy contribution | -23.62 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.731949 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12972264 102 + 23771897 AUUGCGUUGGUGUUUCUGGCUUU---------------UGCCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUCGAA---ACAACAAAUUGAAUGUAAG .(((((((.(((((((((((...---------------.)))).((((.....((....))(((((((......)))))))))))..........)))---)))......).))))))). ( -30.80) >DroSec_CAF1 89643 120 + 1 AUUGCGUUGGCGUUUUUGGCUUUUCGAGGUUUGCUGCUUGCCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUCGAAACAACAACAAAUUGAAUGUAAG .....((((.((((((((((((((....(((((((..........)))))))..)))))).(((((((......)))))))......)))))).))...))))................. ( -34.80) >DroSim_CAF1 98755 119 + 1 AUUGCGUUGGCGUUUUUGGCUUUUCGAGGUUUGCUGCUUGCCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUC-AAACAACAACAAAUUGAAUGUAAG .((((((((((.......((((((....(((((((..........)))))))..)))))).(((((((......))))))))))...........-......(((....)))))))))). ( -34.00) >DroEre_CAF1 95104 114 + 1 AUUGCGUUGGCGUUUCUGGCUUUACGAGGUUUG---CUUGCCAAUGGCAAACAUGAAAGCAGCUCUGCAUGUGAGUAGAACACCAAACAAAAAUCGAA---ACAACAAAUUGAAUGUAAG .(((((((...(((((..(((((.....(((((---((.......)))))))...))))).(.(((((......))))).)..............)))---))..(.....)))))))). ( -26.40) >DroYak_CAF1 93666 114 + 1 AUUGCGUUGGCGUUUCUGGCUUUACGAGGUUUG---CUUGCCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGUCGAACACCAAACAAAAAUCGAA---ACAACAAAUUGAAUGUAAG .(((((((.(((...(((.((((.....(((((---((.......)))))))...)))))))...))).(((...((((..............)))).---)))........))))))). ( -25.24) >consensus AUUGCGUUGGCGUUUCUGGCUUUACGAGGUUUG___CUUGCCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUCGAA___ACAACAAAUUGAAUGUAAG .((((((((((((((...((((...............(((((...)))))(((((((((...)))).))))))))).)))))))..................(((....)))))))))). (-23.38 = -23.62 + 0.24)


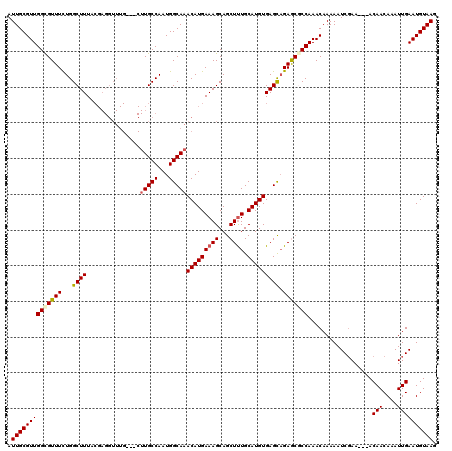
| Location | 12,972,289 – 12,972,405 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.11 |
| Mean single sequence MFE | -27.70 |
| Consensus MFE | -25.57 |
| Energy contribution | -25.90 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.09 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.99 |
| SVM RNA-class probability | 0.984856 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12972289 116 + 23771897 CCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUCGAA---ACAACAAAUUGAAUGUAAGCCAAAGCAACAAUAU-GUAUGUAUGUGGCAACAACAGGUA ((..((((.....((....))(((((((......))))))))))).............---..................((((..(((((.....-)).)))...)))).......)).. ( -25.50) >DroSec_CAF1 89683 120 + 1 CCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUCGAAACAACAACAAAUUGAAUGUAAGCCACAGCAACAAUACCAUAUGUAUGUGGCAACUACAGGUA ((..((((.....((....))(((((((......))))))))))).............................((((.((((((...(((........))).))))))...)))))).. ( -29.20) >DroSim_CAF1 98795 113 + 1 CCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUC-AAACAACAACAAAUUGAAUGUAAGCCACAGCAA-----CCAUAUGUAUGUGGCAACAACAGGC- ((..((((.....((....))(((((((......)))))))))))..........-.......................(((((((((.-----.....))).)))))).......)).- ( -28.40) >consensus CCAAUGGCAAACAUGAAAGCAGCUUUGCAUGUGAGCAGAGCGCCAAACAAAAAUCGAAACAACAACAAAUUGAAUGUAAGCCACAGCAACAAUACCAUAUGUAUGUGGCAACAACAGGUA ((..((((.....((....))(((((((......)))))))))))..................................(((((((((...........))).)))))).......)).. (-25.57 = -25.90 + 0.33)

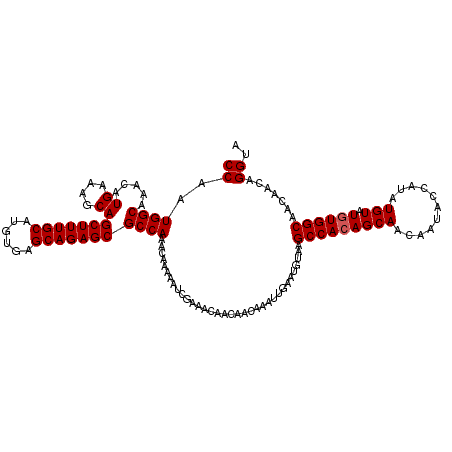

| Location | 12,972,405 – 12,972,525 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -34.64 |
| Consensus MFE | -29.92 |
| Energy contribution | -31.20 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.897449 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12972405 120 - 23771897 UCCGAGCAAGUAACGUUAGAGGCUAGAGGCCAUGUUAAAGUUUUGUCCUCGUCGCCUUCAUCAAGUGUUUCUACACGCCCCGUGCAAUUAAGGUAAUUAGCUUUACGUUGCCAUUGCUCU ...(((((((((((((..(((((((((((.((...........)).))))...(((((......((((....))))((.....))....)))))...))))))))))))))..)))))). ( -37.80) >DroSec_CAF1 89803 120 - 1 UCCGAGCAAGUAACGUUAGAGGCUAGAGGCCAAGUUAAAGUUUUGUCCUCGUCGCCUUCAUCAAGUGUUUCUACACGCCCCGUGCAAUUAAGGUAAUUAGCGUUACGUUGCCAUUGCUCU ...(((((((((((((..((.((((((((.((((.......)))).))))...(((((......((((....))))((.....))....)))))...)))).)))))))))..)))))). ( -38.30) >DroSim_CAF1 98908 120 - 1 UCCGAGCAAGUUACGUUUGGAGGUUGAGGCCAAGUUAAAGUUUUGUGCUCGUCGCCUUCAUCAAGUGUUUCUACACGCCCCGUGCAAAUAAGGUAAUUAGCGUUACGUUGCCAUUGCUCU ...((((((((.(((..(((((((((((..((((.......))))..))))..)))))))....((((....))))....)))))......((((((.........)))))).)))))). ( -32.70) >DroEre_CAF1 95249 120 - 1 UACGAGCCAGUAAUGUUAGAGGCUAAAGGCCAAGUUAAAGUUUUGUCCUCGUCGCCUUCAUCAAGUGUUCAUACACGCCCAGUGCAAUUAAGGUAAUUAGCGUUACGUUGCCAUUGCUCU ...((((..(((((((..((.((((((((.((((.......)))).)))....(((((......((((....))))((.....))....)))))..))))).)))))))))....)))). ( -30.40) >DroYak_CAF1 93811 120 - 1 UCCGAGCAAGUAACAUUAGAGGCUAGAGGCCAAGCUAAAGUUUUGUCCUCGUCGCCUUCAUCAAGUGUUCCUACACGCCCAGUGCAAUUAAGGUAAUUAGCGUUACGUUGCCAUUGCUCU ...(((((((((((....((.(((((((((.((((....)))).).))))...(((((......((((....))))((.....))....)))))...)))).))..)))))..)))))). ( -34.00) >consensus UCCGAGCAAGUAACGUUAGAGGCUAGAGGCCAAGUUAAAGUUUUGUCCUCGUCGCCUUCAUCAAGUGUUUCUACACGCCCCGUGCAAUUAAGGUAAUUAGCGUUACGUUGCCAUUGCUCU ...(((((((((((((..((.((((((((.((((.......)))).))))...(((((......((((....))))((.....))....)))))...)))).)))))))))..)))))). (-29.92 = -31.20 + 1.28)


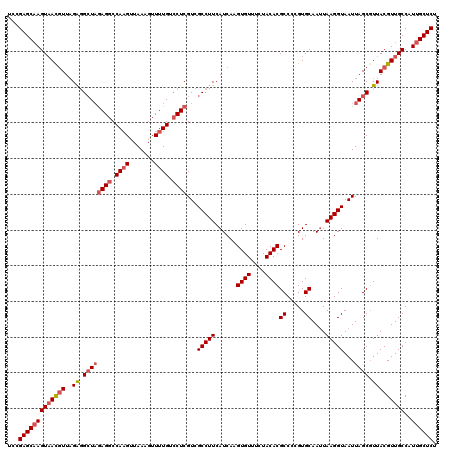
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:24:27 2006