
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 12,949,331 – 12,949,636 |
| Length | 305 |
| Max. P | 0.981561 |

| Location | 12,949,331 – 12,949,438 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.64 |
| Mean single sequence MFE | -33.68 |
| Consensus MFE | -25.04 |
| Energy contribution | -27.04 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928538 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
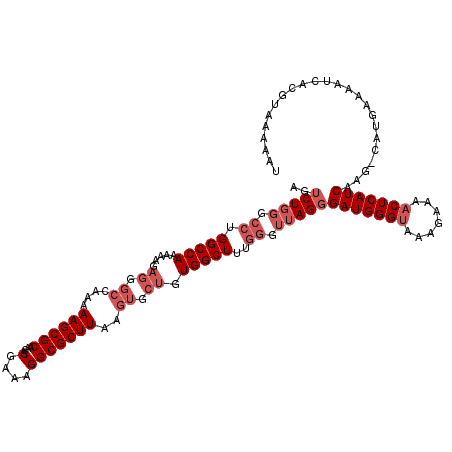
>3L_DroMel_CAF1 12949331 107 - 23771897 AGUCUGGGCCUGGCCA------------AAAAGCGCAACACGAAAGGCGCUUAAGUGCUGUGGCUUUGGGUUAGGGAUGGGUAAAGAAAACUCAUCAAG-CAUGAAAAUCACGUAAAGAA ..(((...(((((((.------------.((((((((...(....)((((....))))))).))))).)))))))(((((((.......)))))))..(-(.((.....)).))..))). ( -31.70) >DroSec_CAF1 66527 119 - 1 AGUCUGGGCCUGGCCAAAAAGAGGGCCAAAAAGCGCAACACGAAAGGCGCUUAAGUGCUGUGGCUUUGGGUUAGGGAUGGGUAAAGAAAACUCAUCAAG-CAUGAAAAUCACGUAAAAAU ..(((((.(((((((........))))).((((((((...(....)((((....))))))).))))))).)))))(((((((.......)))))))..(-(.((.....)).))...... ( -34.80) >DroSim_CAF1 71354 119 - 1 AGUCUGGGCCUGGCCAAAAAGAGGGCCAAAAAGCGCAACACGAAAGGCGCUUAAGUGCUGUGGCUUUGGGUUAGGGAUGGGUAAAGAAAACUCAUCAAG-CAUGAAAAUCACGUAAAAAU ..(((((.(((((((........))))).((((((((...(....)((((....))))))).))))))).)))))(((((((.......)))))))..(-(.((.....)).))...... ( -34.80) >DroEre_CAF1 72559 119 - 1 AGUCUGGGCCUGGCCAAAAAGAGGGCCAA-AAGCGCAACACGAAAGGCGCUUAAGUGCUGUGGCUUUGGAUUAGGGAUGGGCAAAGAAAACUCAUCAGGCCAUGAAUAUCACGUAAAUAU ......(((((((((........)))).(-(((((((...(....)((((....))))))).)))))........((((((.........)))))))))))................... ( -34.20) >DroYak_CAF1 70435 114 - 1 AGUCU------GGCCAAAAAGAGGGCCAAAAAGCGCAACACGAAAGGCGCUUAAGUGCUGUGGCUUUGGAUUAGUGAUGGGUAACGAAGACUCAUCAAGCCAUGAAAAUCACGUAAAUAU ....(------((((........)))))..((((((....(....)))))))..(((.(((((((.........((((((((.......))))))))))))))).....)))........ ( -32.90) >consensus AGUCUGGGCCUGGCCAAAAAGAGGGCCAAAAAGCGCAACACGAAAGGCGCUUAAGUGCUGUGGCUUUGGGUUAGGGAUGGGUAAAGAAAACUCAUCAAG_CAUGAAAAUCACGUAAAAAU ..(((((.((.(((((.....((.((....((((((....(....)))))))..)).)).)))))..)).)))))(((((((.......)))))))........................ (-25.04 = -27.04 + 2.00)


| Location | 12,949,410 – 12,949,516 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.46 |
| Mean single sequence MFE | -33.80 |
| Consensus MFE | -24.56 |
| Energy contribution | -25.32 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.613366 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12949410 106 - 23771897 AAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUC--GCUGCUCUAACCACAGUCUGGGCCUGGCCA------------AAAAGCGCAACA .(((...((....(((((...)))))....))..)))....((((((((((....))((((((.--...((((..((....))..))))..)))))------------).)))))))).. ( -25.60) >DroSec_CAF1 66606 120 - 1 AAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUGGGGCUGCUCUAACCACAGUCUGGGCCUGGCCAAAAAGAGGGCCAAAAAGCGCAACA .(((...((....(((((...)))))....))..)))....((((((((..........((((.((((((.........)))))).))))(((((........)))))..)))))))).. ( -38.60) >DroSim_CAF1 71433 120 - 1 AAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUGGGGCUGCUCUAACCACAGUCUGGGCCUGGCCAAAAAGAGGGCCAAAAAGCGCAACA .(((...((....(((((...)))))....))..)))....((((((((..........((((.((((((.........)))))).))))(((((........)))))..)))))))).. ( -38.60) >DroEre_CAF1 72639 117 - 1 AAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGAUG--GCUGCUCUAACCGCAGUCUGGGCCUGGCCAAAAAGAGGGCCAA-AAGCGCAACA .(((...((....(((((...)))))....))..)))....((((((((..........((..(--(((((.......))))))....))(((((........))))).-)))))))).. ( -33.60) >DroYak_CAF1 70515 112 - 1 AAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUG--GCUGCUCUAACCGCAGUCU------GGCCAAAAAGAGGGCCAAAAAGCGCAACA ......((.....(((((((((((.....(..(((....)))..)(((.((....))..)))))--))))))...)))((..(.(------((((........))))).)..)))).... ( -32.60) >consensus AAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUG__GCUGCUCUAACCACAGUCUGGGCCUGGCCAAAAAGAGGGCCAAAAAGCGCAACA ..(((.((....((((((((((.......(..(((....)))..)(((.((....))..)))....))))))...))))............((((........)))).....)).))).. (-24.56 = -25.32 + 0.76)



| Location | 12,949,438 – 12,949,556 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.04 |
| Mean single sequence MFE | -26.60 |
| Consensus MFE | -23.78 |
| Energy contribution | -23.74 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.573270 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12949438 118 - 23771897 AAAAAAAAAAAGGAAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUC--GCUGCUCUAACCAC ...........(....)......(((..((..(....)...))..)))....((((((((((.......(..(((....)))..)(((.((....))..)))..--))))))...)))). ( -26.60) >DroSec_CAF1 66646 116 - 1 AAAA----AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUGGGGCUGCUCUAACCAC ....----...(....)......(((..((..(....)...))..)))....((((((((((..(....(..(((....)))..)(((.((....))..))).)..))))))...)))). ( -26.40) >DroSim_CAF1 71473 116 - 1 AAAA----AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUGGGGCUGCUCUAACCAC ....----...(....)......(((..((..(....)...))..)))....((((((((((..(....(..(((....)))..)(((.((....))..))).)..))))))...)))). ( -26.40) >DroEre_CAF1 72678 113 - 1 AAA-----AAAGGGAACGAGGGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGAUG--GCUGCUCUAACCGC ...-----...(....)..((..(((..((..(....)...))..)))....(((.((((((((..(((((..........((((....))))..)))))..))--))))))))).)).. ( -26.70) >DroYak_CAF1 70549 113 - 1 AAA-----AAAGGGAACGAGGGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUG--GCUGCUCUAACCGC ...-----...(....)..((..(((..((..(....)...))..)))....(((.((((((((.....(..(((....)))..)(((.((....))..)))))--))))))))).)).. ( -26.90) >consensus AAAA____AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUUUUUGCGCUUGUAAUAACUUGGCUG__GCUGCUCUAACCAC ...........(....)......(((..((..(....)...))..)))....((((((((((.......(..(((....)))..)(((.((....))..)))....))))))...)))). (-23.78 = -23.74 + -0.04)



| Location | 12,949,476 – 12,949,596 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.43 |
| Mean single sequence MFE | -20.92 |
| Consensus MFE | -18.36 |
| Energy contribution | -18.56 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.930739 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12949476 120 - 23771897 ACUGAAAGCCAAACAUUACACGAAAAUUUUGUGUUAAAUUAAAAAAAAAAAGGAAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUU .......((((..((((((((((.....)))))).................(....)......(((..((..(....)...))..)))..)))).))))..................... ( -20.00) >DroSec_CAF1 66686 116 - 1 ACUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAAA----AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUU .(((...((((.....(((((((...)))))))...........----...............(((..((..(....)...))..)))....))))..)))................... ( -19.30) >DroSim_CAF1 71513 116 - 1 ACUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAAA----AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUU .(((...((((.....(((((((...)))))))...........----...............(((..((..(....)...))..)))....))))..)))................... ( -19.30) >DroEre_CAF1 72716 115 - 1 GCUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAA-----AAAGGGAACGAGGGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUU ((((...((((.....(((((((...)))))))..........-----...............(((..((..(....)...))..)))....))))..)))).................. ( -23.00) >DroYak_CAF1 70587 115 - 1 GCUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAA-----AAAGGGAACGAGGGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUU ((((...((((.....(((((((...)))))))..........-----...............(((..((..(....)...))..)))....))))..)))).................. ( -23.00) >consensus ACUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAAA____AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUAAAUUGAGCAAAAUGGUGGCAGCUACUUAAGUGAAAAUAUU .(((...((((.....(((((((...)))))))..................(....)......(((..((..(....)...))..)))....))))..)))................... (-18.36 = -18.56 + 0.20)



| Location | 12,949,516 – 12,949,636 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.96 |
| Mean single sequence MFE | -23.52 |
| Consensus MFE | -21.38 |
| Energy contribution | -21.22 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.981561 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12949516 120 - 23771897 CCCGAAGAUUGUAAAAUGUUCUUGCUCAAAAGAGCAGCGAACUGAAAGCCAAACAUUACACGAAAAUUUUGUGUUAAAUUAAAAAAAAAAAGGAAACGAGAGAGCUUGAUGUUGGAAAUA .((((..(((.......((((((((((....)))))).))))...(((((.......((((((.....)))))).................(....)....).))))))).))))..... ( -23.90) >DroSec_CAF1 66726 116 - 1 UCCGAAGAUUGUAAAAUGUUCUUGCUCAAAAGAGCAGCGAACUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAAA----AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUA (((((..(((.......((((((((((....)))))).))))...(((((......(((((((...)))))))...........----...(....)....).))))))).))))).... ( -24.30) >DroSim_CAF1 71553 116 - 1 UCCGAAGAUUGUAAAAUGUUCUUGUUCAAAAGAGCAGCGAACUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAAA----AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUA (((((..(((.......((((((((((....)))))).))))...(((((......(((((((...)))))))...........----...(....)....).))))))).))))).... ( -21.90) >DroEre_CAF1 72756 115 - 1 UACGAAGAUUGUGAAAUGUUCUUGCUCAAAAGGGCAGCGAGCUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAA-----AAAGGGAACGAGGGAGCUUGAUGUUGGAAAUA .................((((((((((....)))))).))))......((((.((((((((((...)))))))..........-----........((((....)))))))))))..... ( -23.20) >DroYak_CAF1 70627 115 - 1 UUCGAAGAUUGUGAAAUGUUCUCGCUCAAAAGAGCAGCGAGCUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAA-----AAAGGGAACGAGGGAGCUUGAUGUUGGAAAUA (((.((.(((.......((((((((((....))))((((.((.....))).......((((((...)))))))))........-----...))))))(((....)))))).)).)))... ( -24.30) >consensus UCCGAAGAUUGUAAAAUGUUCUUGCUCAAAAGAGCAGCGAACUGAAAGCCAAACAUUACACGAAAAUUGUGUGUUAAAUUAAAA____AAAGGGAACGAGAGAGCUUGAUGUUGGAAAUA .................((((((((((....)))))).))))......((((.((((((((((...))))))).......................((((....)))))))))))..... (-21.38 = -21.22 + -0.16)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:23:10 2006