
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 12,914,732 – 12,914,904 |
| Length | 172 |
| Max. P | 0.962542 |

| Location | 12,914,732 – 12,914,840 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.84 |
| Mean single sequence MFE | -18.71 |
| Consensus MFE | -10.38 |
| Energy contribution | -11.05 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.748023 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12914732 108 + 23771897 AUGCGGCCACCA----UUAUAAGCCUUCUUU---UUUUUUUGUGCAU----AACCAUAAACCAGGCUUAUGGGCAAAAAA-AAAAAUCAAACUAAAAUGUAGCCAAACAAUGAAUAAUAU ....(((...((----((..........(((---(((((((((.(((----((((........)).))))).))))))))-))))..........))))..)))................ ( -22.55) >DroSec_CAF1 35487 110 + 1 AUGCGGCCACCAUUCAUUAUAAGCCUGCUUU------UUUUGUGCAU----AACCAUAAACCAGGCUUAUGGGCAAAAAAAAAAAAUCAAACUAAAAUGUAGCCAAACAAUGAAUAAUAU ...........((((((((((((((((....------.((((((...----...)))))).))))))))).(((...........................)))....)))))))..... ( -20.83) >DroSim_CAF1 34523 106 + 1 AUGCGGCCACCA----UUAUAAGCCUGCUUU------UUUUGUGCAU----AACCAUAAACCAGGCUUAUGGGCAAAAAAAAAAAAACAAACUAAAAUGUAGCCAAACAAUGAAUAAUAU ....(((.....----(((((((((((....------.((((((...----...)))))).)))))))))))..............(((........))).)))................ ( -19.90) >DroEre_CAF1 40792 97 + 1 AUGCGGCCACCA----UUAUAAGCCUGAUAUU------UUUUUGAAU----AACCAUAAACCAGGCUUAAGGGUAAAAAGAA---------CUAAAUUGUAGCCAAACAAUGAAUAAUAU ....(((.(((.----...((((((((.....------.........----..........))))))))..))).......(---------(......)).)))................ ( -17.06) >DroYak_CAF1 38241 109 + 1 AUGCGACCACCA----UUAUAAGCCAGAUUUUUUUUUUUUUUUGAAU----AACCAUAAACCAGGCUUAUGGGCAAAAAAAA---AACUAGCUAAAAUGUAGCCAAACAAUGAAUAAUAU ..........((----((...............(((((((((((...----..((((((.......)))))).)))))))))---))...((((.....)))).....))))........ ( -15.80) >DroPer_CAF1 39753 102 + 1 AAGCGGCUACCA----UGAAAGGCCUGCUUU------UU--GUCCAGCAGCCACAAUAAACCAGGCUUACUA------AGAAAAAAGUGAACUAAAAAGUAACCAAAUGAUGAAUAAUAU ....((.(((..----.....(((.((((..------..--....)))))))...........((.(((((.------.......))))).)).....))).))................ ( -16.10) >consensus AUGCGGCCACCA____UUAUAAGCCUGCUUU______UUUUGUGCAU____AACCAUAAACCAGGCUUAUGGGCAAAAAAAAAAAAACAAACUAAAAUGUAGCCAAACAAUGAAUAAUAU ....(((.........(((((((((((..................................)))))))))))..................((......)).)))................ (-10.38 = -11.05 + 0.67)


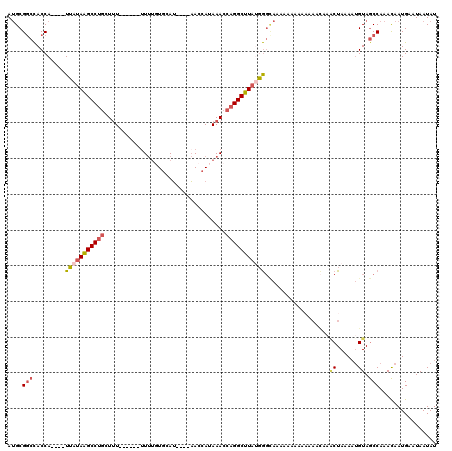
| Location | 12,914,732 – 12,914,840 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.84 |
| Mean single sequence MFE | -26.75 |
| Consensus MFE | -13.70 |
| Energy contribution | -14.59 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.86 |
| Structure conservation index | 0.51 |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.962542 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12914732 108 - 23771897 AUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGAUUUUU-UUUUUUGCCCAUAAGCCUGGUUUAUGGUU----AUGCACAAAAAAA---AAAGAAGGCUUAUAA----UGGUGGCCGCAU ................(((((((((..(((((..(((((-((((((((((((((((...))))))))..----..))).)))))))---)))..)))))...))----).)))))).... ( -30.10) >DroSec_CAF1 35487 110 - 1 AUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGAUUUUUUUUUUUUGCCCAUAAGCCUGGUUUAUGGUU----AUGCACAAAA------AAAGCAGGCUUAUAAUGAAUGGUGGCCGCAU .(((((((((((((..((((.((........))...((((((((.(((((((((((...))))))))..----..))).))))------))))..)))).)))))))))))))....... ( -27.90) >DroSim_CAF1 34523 106 - 1 AUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGUUUUUUUUUUUUUGCCCAUAAGCCUGGUUUAUGGUU----AUGCACAAAA------AAAGCAGGCUUAUAA----UGGUGGCCGCAU ................(((((((((..((((((((((((((....(((((((((((...))))))))..----..))).))))------))))))))))...))----).)))))).... ( -31.30) >DroEre_CAF1 40792 97 - 1 AUAUUAUUCAUUGUUUGGCUACAAUUUAG---------UUCUUUUUACCCUUAAGCCUGGUUUAUGGUU----AUUCAAAAA------AAUAUCAGGCUUAUAA----UGGUGGCCGCAU ................((((((......(---------(.......))...((((((((((...(((..----..)))....------...))))))))))...----..)))))).... ( -21.20) >DroYak_CAF1 38241 109 - 1 AUAUUAUUCAUUGUUUGGCUACAUUUUAGCUAGUU---UUUUUUUUGCCCAUAAGCCUGGUUUAUGGUU----AUUCAAAAAAAAAAAAAAAUCUGGCUUAUAA----UGGUGGUCGCAU ................(((((((((..((((((((---(((((((((.((((((((...))))))))..----...)))))))))))......))))))...))----).)))))).... ( -27.10) >DroPer_CAF1 39753 102 - 1 AUAUUAUUCAUCAUUUGGUUACUUUUUAGUUCACUUUUUUCU------UAGUAAGCCUGGUUUAUUGUGGCUGCUGGAC--AA------AAAGCAGGCCUUUCA----UGGUAGCCGCUU ................(((((((....((..(.((((((..(------(((((.(((...........)))))))))..--))------))))..)..))....----.))))))).... ( -22.90) >consensus AUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGAUUUUUUUUUUUUGCCCAUAAGCCUGGUUUAUGGUU____AUGCACAAAA______AAAGCAGGCUUAUAA____UGGUGGCCGCAU ................((((((......((................))..(((((((((.(((..........................))).)))))))))........)))))).... (-13.70 = -14.59 + 0.89)

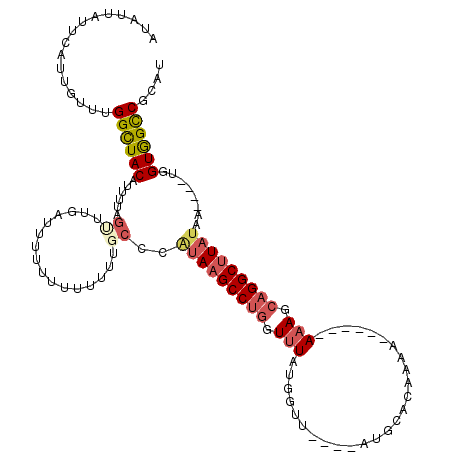
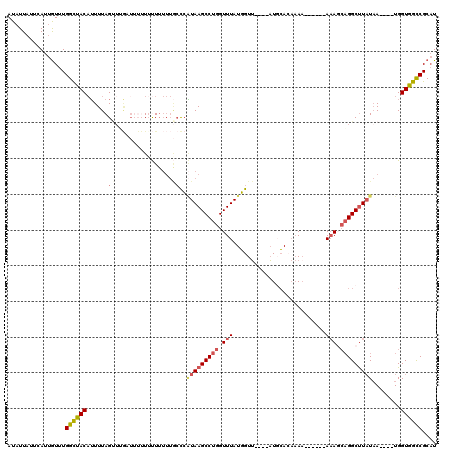
| Location | 12,914,765 – 12,914,873 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 86.54 |
| Mean single sequence MFE | -20.17 |
| Consensus MFE | -15.63 |
| Energy contribution | -16.32 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.77 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.532774 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12914765 108 - 23771897 UCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGAUUUUU-UUUUUUGCCCAUAAGCCUGGUUUAUGGUUAUGCACA .......(((((((...((((((((..(((..............))).)))))))).....)))))))....-.....(((((((((((...))))))))....))).. ( -23.24) >DroSec_CAF1 35521 109 - 1 UCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGAUUUUUUUUUUUUGCCCAUAAGCCUGGUUUAUGGUUAUGCACA .......(((((((...((((((((..(((..............))).)))))))).....)))))))..........(((((((((((...))))))))....))).. ( -23.24) >DroSim_CAF1 34553 109 - 1 UCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGUUUUUUUUUUUUUGCCCAUAAGCCUGGUUUAUGGUUAUGCACA ........(((((....((((((((..(((..............))).)))))))).)))))................(((((((((((...))))))))....))).. ( -21.24) >DroEre_CAF1 40822 100 - 1 UCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAAUUUAG---------UUCUUUUUACCCUUAAGCCUGGUUUAUGGUUAUUCAAA ................(((((((((..(((..............))).))))))))).....---------...............((((........))))....... ( -16.74) >DroYak_CAF1 38277 106 - 1 UCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGCUAGUU---UUUUUUUUGCCCAUAAGCCUGGUUUAUGGUUAUUCAAA ..(((..((((((....((((((((..(((..............))).)))))))).)))))).))).---..........((((((((...))))))))......... ( -24.44) >DroAna_CAF1 61288 96 - 1 UUACUUUGUUAAAUCCUUGUGGCCAUUGAAUAAAAAUUAUUUAUAGUUUGGUUACACUUUA-----------UUUUUUCGUCCAUAAGCCUCUUUUAUUGUGUUGA--A .................((((((((.((((((.....)))))).....)))))))).....-----------....((((..(((((..........))))).)))--) ( -12.10) >consensus UCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUG_U___UUUUUUUUGCCCAUAAGCCUGGUUUAUGGUUAUGCACA .................((((((((..(((..............))).)))))))).........................((((((((...))))))))......... (-15.63 = -16.32 + 0.70)

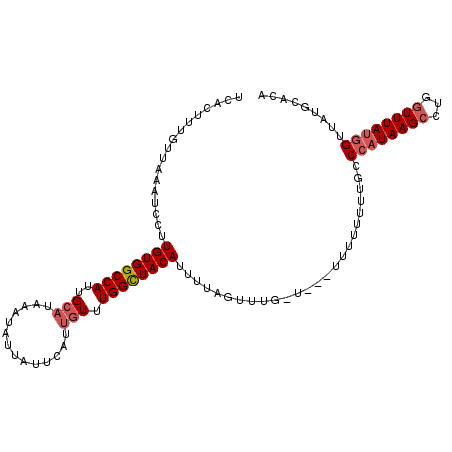
| Location | 12,914,801 – 12,914,904 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 86.86 |
| Mean single sequence MFE | -16.40 |
| Consensus MFE | -13.50 |
| Energy contribution | -14.06 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.738689 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12914801 103 - 23771897 UUAUAACUUUAAUACUUUUGUAUCUUUGCAAUCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGAUUUUU- .................(((((....))))).......(((((((...((((((((..(((..............))).)))))))).....)))))))....- ( -17.34) >DroSec_CAF1 35557 104 - 1 UUAUAACUUUAAUACUUUUGUAUCUUUGCAAUCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGAUUUUUU .................(((((....))))).......(((((((...((((((((..(((..............))).)))))))).....)))))))..... ( -17.34) >DroSim_CAF1 34589 104 - 1 UUAUAACUUUAAUACUUUUGUAUCUUUGCAAUCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUGUUUUUUU .......(((((((...(((((....)))))......)))))))....((((((((..(((..............))).))))))))................. ( -16.64) >DroEre_CAF1 40858 95 - 1 UUAUAGUUUUAAUACUUUUGUAUCUUUGCAAUCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAAUUUAG---------UU .......(((((((...(((((....)))))......)))))))...(((((((((..(((..............))).))))))))).....---------.. ( -17.54) >DroYak_CAF1 38313 101 - 1 UUAUAAUUUUAAUACUUUUGUAGCUUUGCAAUCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGCUAGUU---UU .................(((((....)))))..(((..((((((....((((((((..(((..............))).)))))))).)))))).))).---.. ( -18.44) >DroAna_CAF1 61322 92 - 1 UAAUCAUUUAAAUACC-UUGCAUUUUAGCCAUUACUUUGUUAAAUCCUUGUGGCCAUUGAAUAAAAAUUAUUUAUAGUUUGGUUACACUUUA-----------U ................-......((((((.........))))))....((((((((.((((((.....)))))).....)))))))).....-----------. ( -11.10) >consensus UUAUAACUUUAAUACUUUUGUAUCUUUGCAAUCACUUUGUUAAAUCCUUGUGGCCAUUGCAUAAAUAUUAUUCAUUGUUUGGCUACAUUUUAGUUUG_U___UU .......(((((((...(((((....)))))......)))))))....((((((((..(((..............))).))))))))................. (-13.50 = -14.06 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:22:17 2006