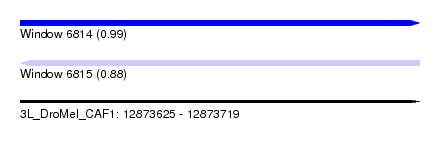

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 12,873,625 – 12,873,719 |
| Length | 94 |
| Max. P | 0.992393 |

| Location | 12,873,625 – 12,873,719 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 78.62 |
| Mean single sequence MFE | -31.61 |
| Consensus MFE | -25.47 |
| Energy contribution | -25.81 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.81 |
| SVM decision value | 2.33 |
| SVM RNA-class probability | 0.992393 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12873625 94 + 23771897 GGAUGGGGGCAUUUUGGGUUUCUGCGUUUCCGAAGUGACAAAAUGGGAAAAGCAUUUGCCCGAGACACUUGAAGUGUAAAUUUCAUGGCCUCA-------------------A ...(((((.((((((((((...(((.(((((.(..(....)..).))))).)))...))))))))....((((((....)))))))).)))))-------------------. ( -31.60) >DroSim_CAF1 7087 89 + 1 G-----GGGCACUUUGGGUUUCUGCGUUUCCGAAGUGACAAAAUAGGAAAAGCAUUUGCCCGAGACACUUGAAGUGUAAAUUUCAUGGCCUCA-------------------A (-----((((..(((((((...(((.(((((..............))))).)))...)))))))...(.((((((....)))))).)))))).-------------------. ( -30.34) >DroYak_CAF1 7119 110 + 1 ---UGGGGGGACAUUGGGCUUCUGGGUUUCCGCAGUGACAAAAUAGGAAAAGCAUUUGCCCGAGACACUUGAAGUGUAAACUUUAUGGCCUCAAGGCGUUCGUACAAGUGCAA ---((.((..((.(..(....)..)))..)).)).................(((((((..((((.(.(.((((((....)))))).)(((....))))))))..))))))).. ( -32.90) >consensus G__UGGGGGCACUUUGGGUUUCUGCGUUUCCGAAGUGACAAAAUAGGAAAAGCAUUUGCCCGAGACACUUGAAGUGUAAAUUUCAUGGCCUCA___________________A .....(((((...((((((...(((.(((((..............))))).)))...))))))....(.((((((....)))))).))))))..................... (-25.47 = -25.81 + 0.34)


| Location | 12,873,625 – 12,873,719 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 78.62 |
| Mean single sequence MFE | -23.41 |
| Consensus MFE | -18.61 |
| Energy contribution | -19.27 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.882618 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12873625 94 - 23771897 U-------------------UGAGGCCAUGAAAUUUACACUUCAAGUGUCUCGGGCAAAUGCUUUUCCCAUUUUGUCACUUCGGAAACGCAGAAACCCAAAAUGCCCCCAUCC .-------------------.((((((.((((........)))).).)))))(((((..(((.(((((..............))))).)))(.....)....)))))...... ( -22.44) >DroSim_CAF1 7087 89 - 1 U-------------------UGAGGCCAUGAAAUUUACACUUCAAGUGUCUCGGGCAAAUGCUUUUCCUAUUUUGUCACUUCGGAAACGCAGAAACCCAAAGUGCCC-----C .-------------------.((((((.((((........)))).).)))))(((((..(((.(((((..............))))).)))...........)))))-----. ( -23.56) >DroYak_CAF1 7119 110 - 1 UUGCACUUGUACGAACGCCUUGAGGCCAUAAAGUUUACACUUCAAGUGUCUCGGGCAAAUGCUUUUCCUAUUUUGUCACUGCGGAAACCCAGAAGCCCAAUGUCCCCCCA--- ..(((((((.......(((....)))....((((....)))))))))))...((((...((..(((((..............)))))..))...))))............--- ( -24.24) >consensus U___________________UGAGGCCAUGAAAUUUACACUUCAAGUGUCUCGGGCAAAUGCUUUUCCUAUUUUGUCACUUCGGAAACGCAGAAACCCAAAGUGCCCCCA__C .....................((((((.((((........)))).).)))))(((....(((.(((((..............))))).)))....)))............... (-18.61 = -19.27 + 0.67)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:21:35 2006