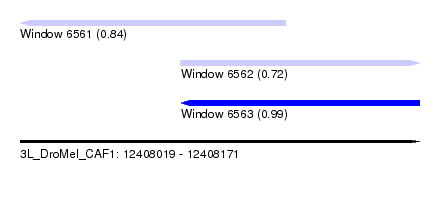

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 12,408,019 – 12,408,171 |
| Length | 152 |
| Max. P | 0.989416 |

| Location | 12,408,019 – 12,408,120 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 78.22 |
| Mean single sequence MFE | -25.42 |
| Consensus MFE | -18.50 |
| Energy contribution | -17.98 |
| Covariance contribution | -0.52 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.838764 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12408019 101 - 23771897 AGUGCCGCUGCAGUGGGGAGUUGCAAUAACAACACCUCUGGCUGUGCAAAAAUCAUUUCACGUUAAAGGGCCCAA----AUACACAUGCAUGCAUA------------UGUACAUAC .((((.((((((((((((.((((......)))).(((.((((.(((.(((.....)))))))))).))).)))..----...))).)))).))...------------.)))).... ( -25.80) >DroSec_CAF1 10617 97 - 1 AGCGCGGCUGCAAUGAGGAGUUGCAAUAACAACACCUCUGGCUGUGCAAAAAUCACUUCACUUUAAAGGGCCCAA----ACACACAU----ACAUG------------UGUACAUAC .((((((((.....((((.((((......)))).)))).))))))))............................----..(((((.----...))------------)))...... ( -26.30) >DroSim_CAF1 10593 101 - 1 AGCGCGGUUGCAAUGGGGAGUUGCAAUAACAACACCUCUGGCUGUGCAAAAAUCACUUCACGUUAAAGGGCCCAA----ACACACAUACAUACAUG------------UGUACAUAC .((((((((.....((((.((((......)))).)))).))))))))............................----..((((((......)))------------)))...... ( -24.20) >DroEre_CAF1 10930 85 - 1 AGCGCGGCAGCAAUGGGGAGUUGCAACAACAACACCUCUGGUUGUGCAAAAAUCAUUUCAUGUUAAAGGGCCCAG----GCACA----------------------------CAUAC .(((((((......((((.((((......)))).))))..)))))))............((((......((....----))..)----------------------------))).. ( -21.20) >DroYak_CAF1 11465 113 - 1 AGCGCGGCAGCAAUGGGGAGUUGCAAAAACAACACCUCUGGUUGUGCAAAAAUCAUUUCAACUUAAAAGGGCCAAACAAGCACA----CAUGCAUAUGUAUGUAUGUAUGUACAUAC .(((((((......((((.((((......)))).))))..)))))))............................(((.(((.(----(((((....)))))).))).)))...... ( -29.60) >consensus AGCGCGGCUGCAAUGGGGAGUUGCAAUAACAACACCUCUGGCUGUGCAAAAAUCAUUUCACGUUAAAGGGCCCAA____ACACACAU_CAUACAUA____________UGUACAUAC .(((((((......((((.((((......)))).))))..)))))))...................................................................... (-18.50 = -17.98 + -0.52)


| Location | 12,408,080 – 12,408,171 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 88.69 |
| Mean single sequence MFE | -34.62 |
| Consensus MFE | -24.20 |
| Energy contribution | -24.36 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.716931 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12408080 91 + 23771897 CAGAGGUGUUGUUAUUGCAACUCCCCACUGCAGCGGCACUUCUCUAUCCCCCCUUUCU-UUUCCAGUGGGUGUGACAGAGAGGCGGCGAACG .(((((((((((...((((.........)))))))))))))))...((((.(((((((-.(..((.....))..).))))))).)).))... ( -28.80) >DroSec_CAF1 10674 90 + 1 CAGAGGUGUUGUUAUUGCAACUCCUCAUUGCAGCCGCGCUUCUCUAUCCCC-CGUUCU-UUUUUAGUGGGUGUGACAGAGAGGCGGCGAACG ..((((.(((((....))))).))))......(((((.((.((((.(((((-((..((-.....)))))).).)).)))))))))))..... ( -35.80) >DroSim_CAF1 10654 91 + 1 CAGAGGUGUUGUUAUUGCAACUCCCCAUUGCAACCGCGCUUCUCUAUCCCCCCUUUCU-UUUUUAGUGGGUGUGACAGAGAGGCGGCGAACG ..(.((.(((((....))))).)).)........(((((((((((.((((((((....-.....)).))).).)).)))))))).))).... ( -31.30) >DroEre_CAF1 10975 91 + 1 CAGAGGUGUUGUUGUUGCAACUCCCCAUUGCUGCCGCGCUUUUCUAUGCCC-CGCUUUGAUUCCAGGGGGUGUGGCAGAGAGGCGGCGAACG ..(.((.(((((....))))).)).).((((((((...((((.((((((((-(.((........))))))))))).)))).))))))))... ( -40.10) >DroYak_CAF1 11538 90 + 1 CAGAGGUGUUGUUUUUGCAACUCCCCAUUGCUGCCGCGCUUUUCUAUGCCC-CUCGUU-UUUCCAGGGGGUGUGACAGAGAGGCGGCGAACG ..(.((.(((((....))))).)).).((((((((...((((..(((((((-(..(..-....)..))))))))..)))).))))))))... ( -37.10) >consensus CAGAGGUGUUGUUAUUGCAACUCCCCAUUGCAGCCGCGCUUCUCUAUCCCC_CUUUCU_UUUCCAGUGGGUGUGACAGAGAGGCGGCGAACG ..(.((.(((((....))))).)).)........((((((((((((..(((.(............).)))..)...)))))))).))).... (-24.20 = -24.36 + 0.16)


| Location | 12,408,080 – 12,408,171 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 88.69 |
| Mean single sequence MFE | -33.46 |
| Consensus MFE | -26.31 |
| Energy contribution | -27.19 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.79 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989416 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 12408080 91 - 23771897 CGUUCGCCGCCUCUCUGUCACACCCACUGGAAA-AGAAAGGGGGGAUAGAGAAGUGCCGCUGCAGUGGGGAGUUGCAAUAACAACACCUCUG .((.((.(((.((((((((...(((.((.....-....))))).)))))))).))).))..))...((((.((((......)))).)))).. ( -32.30) >DroSec_CAF1 10674 90 - 1 CGUUCGCCGCCUCUCUGUCACACCCACUAAAAA-AGAACG-GGGGAUAGAGAAGCGCGGCUGCAAUGAGGAGUUGCAAUAACAACACCUCUG .((..((((((((((((((...(((........-.....)-)).))))))).)).))))).))...((((.((((......)))).)))).. ( -36.42) >DroSim_CAF1 10654 91 - 1 CGUUCGCCGCCUCUCUGUCACACCCACUAAAAA-AGAAAGGGGGGAUAGAGAAGCGCGGUUGCAAUGGGGAGUUGCAAUAACAACACCUCUG .((..((((((((((((((...(((.((.....-....))))).))))))).)).))))).))...((((.((((......)))).)))).. ( -34.90) >DroEre_CAF1 10975 91 - 1 CGUUCGCCGCCUCUCUGCCACACCCCCUGGAAUCAAAGCG-GGGCAUAGAAAAGCGCGGCAGCAAUGGGGAGUUGCAACAACAACACCUCUG .(((.(((((((.((((.....((((((........)).)-)))..))))..)).))))))))...((((.((((......)))).)))).. ( -30.70) >DroYak_CAF1 11538 90 - 1 CGUUCGCCGCCUCUCUGUCACACCCCCUGGAAA-AACGAG-GGGCAUAGAAAAGCGCGGCAGCAAUGGGGAGUUGCAAAAACAACACCUCUG .(((.(((((((.(((((.....(((((.(...-..).))-))).)))))..)).))))))))...((((.((((......)))).)))).. ( -33.00) >consensus CGUUCGCCGCCUCUCUGUCACACCCACUGGAAA_AGAAAG_GGGGAUAGAGAAGCGCGGCUGCAAUGGGGAGUUGCAAUAACAACACCUCUG .....((((((((((((((...(((................)))))))))).)).)))))......((((.((((......)))).)))).. (-26.31 = -27.19 + 0.88)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:17:33 2006