
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 12,388,649 – 12,388,763 |
| Length | 114 |
| Max. P | 0.999604 |

| Location | 12,388,649 – 12,388,763 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 93.88 |
| Mean single sequence MFE | -31.69 |
| Consensus MFE | -29.53 |
| Energy contribution | -28.49 |
| Covariance contribution | -1.04 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.93 |
| SVM decision value | 3.77 |
| SVM RNA-class probability | 0.999604 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
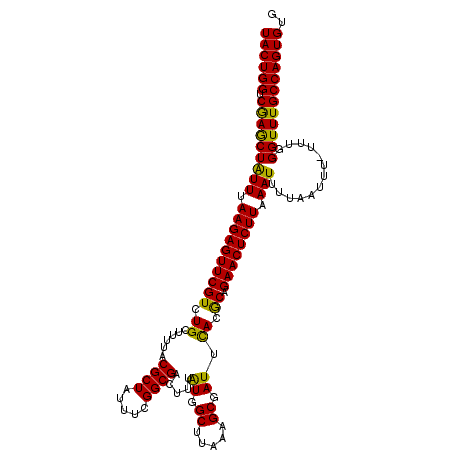
>3L_DroMel_CAF1 12388649 114 + 23771897 UACUGGUCGAGCUAUUUAAGAGUUCGUCUGCUUUUACGCUAUUUCGGCGACUUUGUGGCUUAAAGCGAUUCACGCAGAACUCUUCAAUUUUAAUUU-UUUGGGUUUGCCAGUGUG ((((((.((((((....((((((((.......((((.(((((...(....)...))))).))))(((.....))).))))))))(((.........-.))))))))))))))).. ( -33.30) >DroSec_CAF1 15102 114 + 1 UACUGGUCAAACUAUUUAAGAGUUCGUCUGCUUUUACGCUAAUUCGGCGACUUUAUGGCUUAAAGCGAUUUACGCAGAACUCUUUAAUUUUAAUUU-UUUGGGUUUGCCAGUGUG ((((((.((((((....(((((((((((........((((.....)))).......))).....(((.....))).))))))))............-....)))))))))))).. ( -31.61) >DroSim_CAF1 15228 114 + 1 UACUGGUCGAGCUAUUUAAGAGUUCGUCUGCUUUUACGCUAUUUCGGCGACUUUAUGGCUUAAAGCGAUUCACGCAGAACUCUUUAAUUUUAAUUU-UUUGGGUUUGCCAGUGUG ((((((.((((((....((((((((.......((((.(((((...(....)...))))).))))(((.....))).))))))))............-....)))))))))))).. ( -32.35) >DroEre_CAF1 15055 115 + 1 UACUGGUCGAGCUAUUCAAGAGUUCGUCUGGUUUUACGCUAUUGCGGCGACUUUAUGGCUUAACGCGAUUCACACAGAACUCUUAAAUUUUAAUUUUUUUGGGUUUGCCAGUGUG ((((((.((((((....((((((((((..((((...(((....)))..))))..)).((.....))..........))))))))......(((.....))))))))))))))).. ( -30.10) >DroYak_CAF1 15306 114 + 1 UACUGGUCGAGCUGUUCAAGAGUUCGUCUGGUUUUACGCUAUUUCGGCGGCUUUAUGGCUUUAAGCGAUUCACACAGAACUCUUAAAUUUUAAUUU-UUUGGGUUUGCCAGUGUG ((((((.(((((((((.((((((((((.(((.....((((.....))))((((.........))))...))).)).)))))))).)))........-....)))))))))))).. ( -31.10) >consensus UACUGGUCGAGCUAUUUAAGAGUUCGUCUGCUUUUACGCUAUUUCGGCGACUUUAUGGCUUAAAGCGAUUCACGCAGAACUCUUAAAUUUUAAUUU_UUUGGGUUUGCCAGUGUG ((((((.(((((((((.((((((((((.((......((((.....)))).....((.((.....)).)).)).)).)))))))).))).............)))))))))))).. (-29.53 = -28.49 + -1.04)


| Location | 12,388,649 – 12,388,763 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 93.88 |
| Mean single sequence MFE | -24.21 |
| Consensus MFE | -18.60 |
| Energy contribution | -18.36 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.788675 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
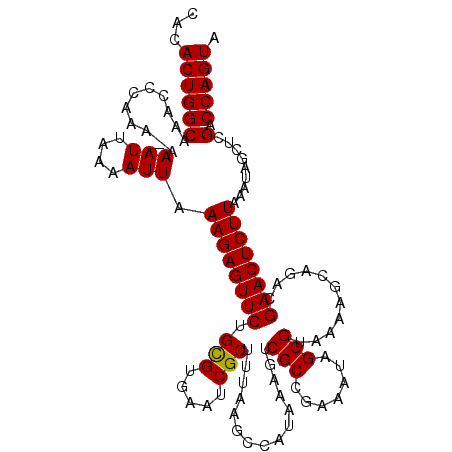
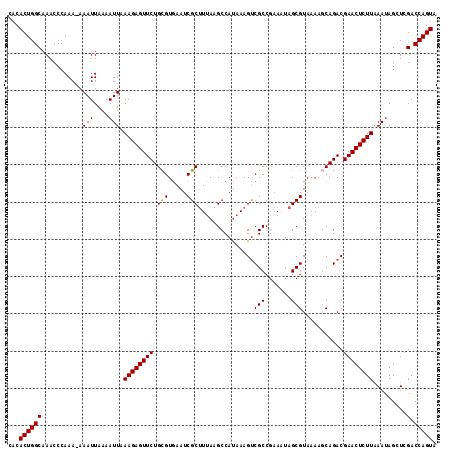
>3L_DroMel_CAF1 12388649 114 - 23771897 CACACUGGCAAACCCAAA-AAAUUAAAAUUGAAGAGUUCUGCGUGAAUCGCUUUAAGCCACAAAGUCGCCGAAAUAGCGUAAAAGCAGACGAACUCUUAAAUAGCUCGACCAGUA ...((((((.....(((.-.........)))(((((((((((......((((....((.((...)).))......)))).....)))))...)))))).........).))))). ( -23.40) >DroSec_CAF1 15102 114 - 1 CACACUGGCAAACCCAAA-AAAUUAAAAUUAAAGAGUUCUGCGUAAAUCGCUUUAAGCCAUAAAGUCGCCGAAUUAGCGUAAAAGCAGACGAACUCUUAAAUAGUUUGACCAGUA ...((((((((((.....-.(((....))).(((((((((((.......((((((.....))))))(((.......))).....)))))...)))))).....))))).))))). ( -26.70) >DroSim_CAF1 15228 114 - 1 CACACUGGCAAACCCAAA-AAAUUAAAAUUAAAGAGUUCUGCGUGAAUCGCUUUAAGCCAUAAAGUCGCCGAAAUAGCGUAAAAGCAGACGAACUCUUAAAUAGCUCGACCAGUA ...((((((.........-.(((....))).(((((((((((.......((((((.....))))))(((.......))).....)))))...)))))).........).))))). ( -21.60) >DroEre_CAF1 15055 115 - 1 CACACUGGCAAACCCAAAAAAAUUAAAAUUUAAGAGUUCUGUGUGAAUCGCGUUAAGCCAUAAAGUCGCCGCAAUAGCGUAAAACCAGACGAACUCUUGAAUAGCUCGACCAGUA ...((((((.......((....))...((((((((((((.((.((....(((..............)))(((....)))......)).)))))))))))))).....).))))). ( -25.34) >DroYak_CAF1 15306 114 - 1 CACACUGGCAAACCCAAA-AAAUUAAAAUUUAAGAGUUCUGUGUGAAUCGCUUAAAGCCAUAAAGCCGCCGAAAUAGCGUAAAACCAGACGAACUCUUGAACAGCUCGACCAGUA ...(((((..........-.........(((((((((((.((.((....((((.........))))(((.......)))......)).)))))))))))))(.....).))))). ( -24.00) >consensus CACACUGGCAAACCCAAA_AAAUUAAAAUUAAAGAGUUCUGCGUGAAUCGCUUUAAGCCAUAAAGUCGCCGAAAUAGCGUAAAAGCAGACGAACUCUUAAAUAGCUCGACCAGUA ...((((((...........(((....))).((((((((.(((.....)))...............(((.......)))...........)))))))).........).))))). (-18.60 = -18.36 + -0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:17:24 2006