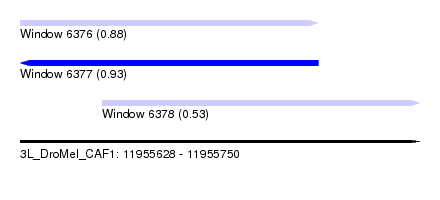
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 11,955,628 – 11,955,750 |
| Length | 122 |
| Max. P | 0.927579 |

| Location | 11,955,628 – 11,955,719 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 80.07 |
| Mean single sequence MFE | -37.68 |
| Consensus MFE | -27.38 |
| Energy contribution | -28.33 |
| Covariance contribution | 0.95 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.883808 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11955628 91 + 23771897 UGCGGGGCAGUGUAUCCUGCCGGCGGAGCGGAGUGGA-------AACUGCUCCAACGGAGAGAGUAGAUCCGUGGUGGGCAGUUGCCACAGCUACGAA .....(((((......)))))..((.(((...((((.-------(((((((((.(((((.........))))).).)))))))).)))).))).)).. ( -38.90) >DroVir_CAF1 8922 97 + 1 UGCCAAUCAGUGCCGCCUGAACGUU-AGUCAAGUGGAAUCCACUGGCUGCUUGCAUGGACAAAAACGCGCCUUGGCCACCAGCUGCCACUUAUACGAA ......((((......)))).(((.-....((((((......((((..(((.((.((........)).))...)))..))))...))))))..))).. ( -22.00) >DroSec_CAF1 8488 91 + 1 UGCGGGGCAGUGUAUCCUGCCGGCGGAGCGUAGUGGA-------AGCUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAA .....(((((......)))))..((.(((...((((.-------(((((((((.(((((.........))))).).)))))))).)))).))).)).. ( -41.30) >DroSim_CAF1 8436 91 + 1 UGCGGGGCAGUGUAUCCUGCCGGCGGAGCGGAGUGGA-------AACUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAA .((..((((((....((..((...(((((((......-------..))))))).(((((.........)))))))..))..))))))...))...... ( -41.10) >DroEre_CAF1 8799 91 + 1 UGCGGGGAUGUGCUUCCUGCCGGCAGAGCGGAGAGGA-------AACUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAA .((..((((.((((..((.(((((...))((((((..-------..)).))))..)))))..)))).))))((((..(....)..)))).))...... ( -42.50) >DroYak_CAF1 6778 91 + 1 UGCGGGGCAGUGUGUCCUGCCGGCGGAGCGGAGUGAA-------AGCUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGUUGCCACAGCUACGAC ..((..((.(((.((.(((((.(((((((((......-------..))))))).(((((.........))))).)).)))))..))))).))..)).. ( -40.30) >consensus UGCGGGGCAGUGUAUCCUGCCGGCGGAGCGGAGUGGA_______AACUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAA ..((.(((.(((....(((((.(((((((((...............))))))).(((((.........))))).)).)))))....))).))).)).. (-27.38 = -28.33 + 0.95)



| Location | 11,955,628 – 11,955,719 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 80.07 |
| Mean single sequence MFE | -35.02 |
| Consensus MFE | -23.30 |
| Energy contribution | -23.72 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.67 |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.927579 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11955628 91 - 23771897 UUCGUAGCUGUGGCAACUGCCCACCACGGAUCUACUCUCUCCGUUGGAGCAGUU-------UCCACUCCGCUCCGCCGGCAGGAUACACUGCCCCGCA ...(.(((.((((.(((((((((..(((((.........)))))))).))))))-------.))))...))).)((.(((((......)))))..)). ( -37.70) >DroVir_CAF1 8922 97 - 1 UUCGUAUAAGUGGCAGCUGGUGGCCAAGGCGCGUUUUUGUCCAUGCAAGCAGCCAGUGGAUUCCACUUGACU-AACGUUCAGGCGGCACUGAUUGGCA ...((...((((.(.(((.((((.(((((.....))))).))))...))).((((((((...)))))((((.-...).))))))).)))).....)). ( -28.40) >DroSec_CAF1 8488 91 - 1 UUCGUAGCUGUGGCAGCUGCCCACCACGGAUCUGCUCUCUCCGUUGGAGCAGCU-------UCCACUACGCUCCGCCGGCAGGAUACACUGCCCCGCA ...(.(((.((((.(((((((((..(((((.........)))))))).))))))-------.))))...))).)((.(((((......)))))..)). ( -40.10) >DroSim_CAF1 8436 91 - 1 UUCGUAGCUGUGGCAGCUGCCCACCACGGAUCUGCUCUCUCCGUUGGAGCAGUU-------UCCACUCCGCUCCGCCGGCAGGAUACACUGCCCCGCA ...(.(((.((((.(((((((((..(((((.........)))))))).))))))-------.))))...))).)((.(((((......)))))..)). ( -37.70) >DroEre_CAF1 8799 91 - 1 UUCGUAGCUGUGGCAGCUGCCCACCACGGAUCUGCUCUCUCCGUUGGAGCAGUU-------UCCUCUCCGCUCUGCCGGCAGGAAGCACAUCCCCGCA ...(((((((...))))))).......((((.((((..(((((..(((((.(..-------......).)))))..))).))..)))).))))..... ( -35.00) >DroYak_CAF1 6778 91 - 1 GUCGUAGCUGUGGCAACUGCCCACCACGGAUCUGCUCUCUCCGUUGGAGCAGCU-------UUCACUCCGCUCCGCCGGCAGGACACACUGCCCCGCA ...(.(((.((((...(((((((..(((((.........)))))))).))))..-------.))))...))).)((.(((((......)))))..)). ( -31.20) >consensus UUCGUAGCUGUGGCAGCUGCCCACCACGGAUCUGCUCUCUCCGUUGGAGCAGCU_______UCCACUCCGCUCCGCCGGCAGGAUACACUGCCCCGCA .........((((.(((((((((..(((((.........)))))))).)))))).......................(((((......))))))))). (-23.30 = -23.72 + 0.42)



| Location | 11,955,653 – 11,955,750 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 82.43 |
| Mean single sequence MFE | -33.42 |
| Consensus MFE | -23.14 |
| Energy contribution | -24.00 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.533773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11955653 97 + 23771897 GAGCGGAGUGGAAACUGCUCCAACGGAGAGAGUAGAUCCGUGGUGGGCAGUUGCCACAGCUACGAAGUGUGCUCGGAUAAUGGCCAGUGGAUAAGGA (((((..((((.(((((((((.(((((.........))))).).)))))))).)))).(((....))).)))))((.......))............ ( -33.70) >DroSec_CAF1 8513 97 + 1 GAGCGUAGUGGAAGCUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAAUUGUGCUCGGAUAAUGGCCAGUGGAUGAGGA ((((...((((.(((((((((.(((((.........))))).).)))))))).)))).((........))))))((.......))............ ( -35.30) >DroSim_CAF1 8461 97 + 1 GAGCGGAGUGGAAACUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAACUGUGCUCGGAUAAUGGCCAGUGGAUGAGGC (((((.((((((..(((((((......).)))))).)))(((((.(((....)))...)))))..))).)))))........(((.........))) ( -36.80) >DroEre_CAF1 8824 94 + 1 GAGCGGAGAGGAAACUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAACUGUGCUCGGG---UGGCCAGUGGAGGAGGA (((((.((.(((..(((((((......).)))))).)))(((((.(((....)))...)))))...)).)))))((.---...))............ ( -35.20) >DroYak_CAF1 6803 94 + 1 GAGCGGAGUGAAAGCUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGUUGCCACAGCUACGACCUGUGUGCGGA---UGGCCAGUGGAUGAGGA ..(((((.......(((((((......).)))))).)))))(.(((.((.(((((((((.......))))).)))).---)).))).)......... ( -31.80) >DroAna_CAF1 8711 94 + 1 GACGAGAAAGCAAACUGUUCGGCCGGAGAUAAUCGUGCUCUGGUGGGCAGCUGCCAUAGCUAUGAGCUGUGCCAGGA---GGAAAAAUGGAUCACAA ...((.........(((((((.((((((.........)))))))))))))(((.((((((.....)))))).)))..---...........)).... ( -27.70) >consensus GAGCGGAGUGGAAACUGCUCCAACGGAGAGAGCAGAUCCGUGGUGGGCAGCUGCCACAGCUACGAACUGUGCUCGGA___UGGCCAGUGGAUGAGGA (((((.((......(((((((......).))))))...((((((.(((....)))...))))))..)).)))))....................... (-23.14 = -24.00 + 0.86)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:14:27 2006