
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 11,797,326 – 11,797,481 |
| Length | 155 |
| Max. P | 0.982574 |

| Location | 11,797,326 – 11,797,441 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.38 |
| Mean single sequence MFE | -41.42 |
| Consensus MFE | -26.90 |
| Energy contribution | -27.88 |
| Covariance contribution | 0.98 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.92 |
| SVM RNA-class probability | 0.982574 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11797326 115 + 23771897 ACGUGCUUGGAUAGAAGCUCCCAUGGGAGUUGAGUUCAUCACGUCAUCAGGUAGUCGAGGACAUGUGGCUGUCACAUC--UCGACGGACCUGCACUCAGGGUCUUUAAAA---UUUGAGU (((((..(((((...((((((....))))))..))))).)))))...(((((.((((((...(((((.....))))))--)))))..))))).(((((((..........---))))))) ( -42.30) >DroPse_CAF1 10896 110 + 1 CAGUGAGU--ACGCCUGCUCCC-CGCGAGUAGUGUG--CCACAGCGUUAGGAAGUCGAU--CAUGUGG---UAACAUUGAUCGGCGGACCUGCACUCAGGGUCUUAAAAACAAAUUGAGU ..(((.((--((((.(((((..-...))))))))))--))))...((.(((..((((((--(((((..---..))).))))))))...)))))((((((.((.......))...)))))) ( -39.10) >DroSec_CAF1 6864 115 + 1 ACGUGCUUGGAGAGAAGCUCCCAUGGGAGUUGAGUCCAUCACGUCAUCAGGUAGUCGAGCACAUGUGGCUGUCACAUC--UCGACGGACCUGCACUCAGGGUCUUUAAAA---UUUGAGU (((((..((((....((((((....))))))...)))).)))))...(((((.((((((...(((((.....))))))--)))))..))))).(((((((..........---))))))) ( -43.40) >DroSim_CAF1 6460 115 + 1 ACGUGCUUGGAGAGAAGCUCCCAUGGGAGUUGAGUCCAUCACGACAUCAGGUAGUCGAGGACAUGUGGCUGUCACAUC--UCGACGGACCUGCACUCAGGGUCUUUAAAA---UUUGAGU .((((..((((....((((((....))))))...)))).))))....(((((.((((((...(((((.....))))))--)))))..))))).(((((((..........---))))))) ( -43.20) >DroPer_CAF1 11413 110 + 1 CAGUGAGU--ACGCCUGCUCCC-CGCGAGUAGUGUG--CCACAGCGUUAGGAAGUCGAU--CAUGUGG---UAACAUUGAUCGGCGGACCUGCACUCAGGGUCUUAAAAACAAAUUGAGU ..(((.((--((((.(((((..-...))))))))))--))))...((.(((..((((((--(((((..---..))).))))))))...)))))((((((.((.......))...)))))) ( -39.10) >consensus ACGUGCUUGGACAGAAGCUCCCAUGGGAGUUGAGUCCAUCACGGCAUCAGGUAGUCGAG_ACAUGUGGCUGUCACAUC__UCGACGGACCUGCACUCAGGGUCUUUAAAA___UUUGAGU ..(((..(((((...((((((....))))))..))))).)))....((((...(((((....(((((.....)))))...)))))((((((.......))))))..........)))).. (-26.90 = -27.88 + 0.98)


| Location | 11,797,366 – 11,797,481 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.55 |
| Mean single sequence MFE | -38.32 |
| Consensus MFE | -25.22 |
| Energy contribution | -26.84 |
| Covariance contribution | 1.62 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.84 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968099 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
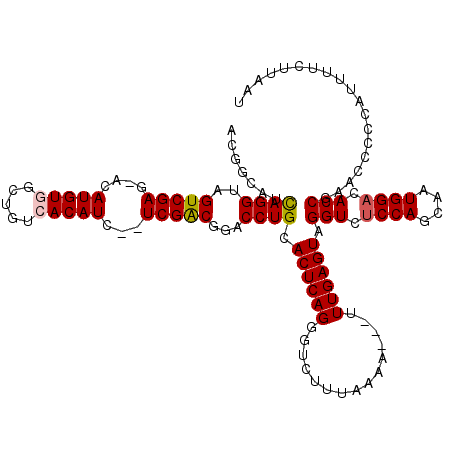
>3L_DroMel_CAF1 11797366 115 + 23771897 ACGUCAUCAGGUAGUCGAGGACAUGUGGCUGUCACAUC--UCGACGGACCUGCACUCAGGGUCUUUAAAA---UUUGAGUAGGUCUCCAGCAAUGGACACCGAACCCCCAUUUUCUUAAU .......(((((.((((((...(((((.....))))))--)))))..))))).(((((((..........---))))))).(((.((((....)))).)))................... ( -35.20) >DroPse_CAF1 10931 114 + 1 ACAGCGUUAGGAAGUCGAU--CAUGUGG---UAACAUUGAUCGGCGGACCUGCACUCAGGGUCUUAAAAACAAAUUGAGUAGGUCCCCAGCAAUGGAGACCGACCCCUGUAGUUAUUAA- ((((.((((((..((((((--(((((..---..))).))))))))...)))).))...(((((......((.......)).((((.(((....))).))))))))))))).........- ( -40.20) >DroSec_CAF1 6904 115 + 1 ACGUCAUCAGGUAGUCGAGCACAUGUGGCUGUCACAUC--UCGACGGACCUGCACUCAGGGUCUUUAAAA---UUUGAGUAGGUCUCCAGUAAUGGACACCGAACCCCCAUUUUCUUAAU .......(((((.((((((...(((((.....))))))--)))))..))))).(((((((..........---))))))).(((.((((....)))).)))................... ( -35.20) >DroSim_CAF1 6500 115 + 1 ACGACAUCAGGUAGUCGAGGACAUGUGGCUGUCACAUC--UCGACGGACCUGCACUCAGGGUCUUUAAAA---UUUGAGUAGGUCUCCAGUAAUGGACACCGAACCCCCAUUUUCUUAAU ..((...(((((.((((((...(((((.....))))))--)))))..)))))...)).(((((((.....---...)))..(((.((((....)))).)))..))))............. ( -36.00) >DroPer_CAF1 11448 114 + 1 ACAGCGUUAGGAAGUCGAU--CAUGUGG---UAACAUUGAUCGGCGGACCUGCACUCAGGGUCUUAAAAACAAAUUGAGUAGGUCUCCAGCAAUGGAGACCGACCCCUGUAGUUAUUAA- ((((.((((((..((((((--(((((..---..))).))))))))...)))).))...(((((......((.......)).((((((((....))))))))))))))))).........- ( -45.00) >consensus ACGGCAUCAGGUAGUCGAG_ACAUGUGGCUGUCACAUC__UCGACGGACCUGCACUCAGGGUCUUUAAAA___UUUGAGUAGGUCUCCAGCAAUGGACACCGAACCCCCAUUUUCUUAAU .......((((..(((((....(((((.....)))))...)))))...)))).((((((...............)))))).(((.((((....)))).)))................... (-25.22 = -26.84 + 1.62)


| Location | 11,797,366 – 11,797,481 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.55 |
| Mean single sequence MFE | -36.12 |
| Consensus MFE | -24.48 |
| Energy contribution | -24.64 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.768728 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11797366 115 - 23771897 AUUAAGAAAAUGGGGGUUCGGUGUCCAUUGCUGGAGACCUACUCAAA---UUUUAAAGACCCUGAGUGCAGGUCCGUCGA--GAUGUGACAGCCACAUGUCCUCGACUACCUGAUGACGU ......(((((..((((..(((.((((....)))).))).))))..)---))))...........((.(((((..(((((--((((((.....)))))...)))))).)))))...)).. ( -35.80) >DroPse_CAF1 10931 114 - 1 -UUAAUAACUACAGGGGUCGGUCUCCAUUGCUGGGGACCUACUCAAUUUGUUUUUAAGACCCUGAGUGCAGGUCCGCCGAUCAAUGUUA---CCACAUG--AUCGACUUCCUAACGCUGU -.........(((((((((((((((((....)))))))).((.......))......)))))...((..(((...(.((((((.(((..---..)))))--)))).)..))).)).)))) ( -35.80) >DroSec_CAF1 6904 115 - 1 AUUAAGAAAAUGGGGGUUCGGUGUCCAUUACUGGAGACCUACUCAAA---UUUUAAAGACCCUGAGUGCAGGUCCGUCGA--GAUGUGACAGCCACAUGUGCUCGACUACCUGAUGACGU ......(((((..((((..(((.((((....)))).))).))))..)---))))...........((.(((((..(((((--((((((.....)))))...)))))).)))))...)).. ( -35.80) >DroSim_CAF1 6500 115 - 1 AUUAAGAAAAUGGGGGUUCGGUGUCCAUUACUGGAGACCUACUCAAA---UUUUAAAGACCCUGAGUGCAGGUCCGUCGA--GAUGUGACAGCCACAUGUCCUCGACUACCUGAUGUCGU ......(((((..((((..(((.((((....)))).))).))))..)---)))).........((...(((((..(((((--((((((.....)))))...)))))).)))))...)).. ( -36.50) >DroPer_CAF1 11448 114 - 1 -UUAAUAACUACAGGGGUCGGUCUCCAUUGCUGGAGACCUACUCAAUUUGUUUUUAAGACCCUGAGUGCAGGUCCGCCGAUCAAUGUUA---CCACAUG--AUCGACUUCCUAACGCUGU -.........(((((((((((((((((....)))))))).((.......))......)))))...((..(((...(.((((((.(((..---..)))))--)))).)..))).)).)))) ( -36.70) >consensus AUUAAGAAAAUGGGGGUUCGGUGUCCAUUGCUGGAGACCUACUCAAA___UUUUAAAGACCCUGAGUGCAGGUCCGUCGA__GAUGUGACAGCCACAUGU_CUCGACUACCUGAUGACGU .............((((..(((.((((....)))).)))..................(..((((....))))..)(((((...(((((.....)))))....))))).))))........ (-24.48 = -24.64 + 0.16)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:13:18 2006