
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 11,694,747 – 11,694,876 |
| Length | 129 |
| Max. P | 0.999428 |

| Location | 11,694,747 – 11,694,840 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 77.74 |
| Mean single sequence MFE | -23.20 |
| Consensus MFE | -14.28 |
| Energy contribution | -14.12 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.885766 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11694747 93 + 23771897 U---------UUUCUGAAAUGUUUUCUUUUCAAUUCAAAUUUGGCGGGAAAGCCAUUUCCCGCUGUUAAGUGUGACCAAAAUCAAUUGCGAGAUGUAACAAU .---------.........((((.((((..(((((....((((((((((((....)))))))).((((....))))))))...))))).))))...)))).. ( -21.00) >DroSec_CAF1 12849 93 + 1 U---------UUUCAAAAAUGUUUUCUUUUCAAUUCAAAUUUAGCGGGAAAUCCAUUUCCCGCUGUUAAGUGUGACCAAAAAAAAUUGAGAGAUCCAUCGAU .---------........(((...(((((.(((((.......(((((((((....)))))))))((((....)))).......))))))))))..))).... ( -19.90) >DroSim_CAF1 8313 93 + 1 A---------UUUCUAAAAUGUUUUCUUUUCAAUUCAAAUUUGGCGGGAAAUCCAUUUCCCGCUGUUAAGUGCGACCAAAAAGAAUUGAGAGAUGUAUCGAU .---------................((((((((((...((((((((((((....))))))((........))).)))))..)))))))))).......... ( -23.20) >DroEre_CAF1 8551 86 + 1 ---------------GAUAUGC-UUCUUUGCAAUUCAAAUUUGGCGGGAAAUCCUUUUCCCGCUGUACAGUGUGACCAAAGGGGUGCGCGAGAUUUGUCGGU ---------------....(((-......)))...((((((((((((((((....))))))))).....((((..((....))..)))).)))))))..... ( -27.90) >DroYak_CAF1 8627 102 + 1 UUGUAAAAUUUAUCUGAAAUGUUUUCCGUUCUGUUCAAAUUUGGCGGGAAAUCCAUUUCCCGCUGUAUAGUGUGACCAAAAGAAAUUGUGAUAUUUCUCGAU ...........(((.((((((((.....((((..(((.(((.(((((((((....)))))))))....))).))).....)))).....))))))))..))) ( -24.00) >consensus U_________UUUCUGAAAUGUUUUCUUUUCAAUUCAAAUUUGGCGGGAAAUCCAUUUCCCGCUGUUAAGUGUGACCAAAAAAAAUUGAGAGAUGUAUCGAU ..........................................(((((((((....)))))))))...................................... (-14.28 = -14.12 + -0.16)


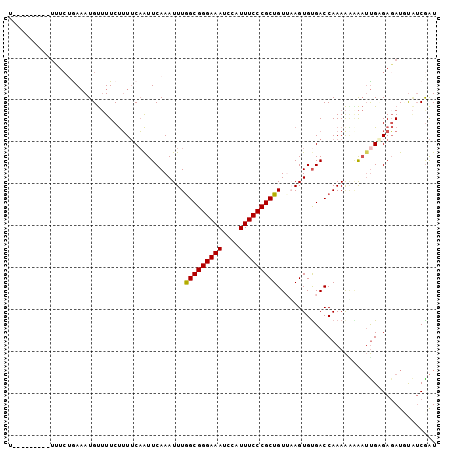
| Location | 11,694,747 – 11,694,840 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 77.74 |
| Mean single sequence MFE | -19.63 |
| Consensus MFE | -14.30 |
| Energy contribution | -15.30 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.831056 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11694747 93 - 23771897 AUUGUUACAUCUCGCAAUUGAUUUUGGUCACACUUAACAGCGGGAAAUGGCUUUCCCGCCAAAUUUGAAUUGAAAAGAAAACAUUUCAGAAA---------A ..((((...(((..(((((.(.((((((........)).((((((((....))))))))))))..).)))))...))).)))).........---------. ( -18.40) >DroSec_CAF1 12849 93 - 1 AUCGAUGGAUCUCUCAAUUUUUUUUGGUCACACUUAACAGCGGGAAAUGGAUUUCCCGCUAAAUUUGAAUUGAAAAGAAAACAUUUUUGAAA---------A ...((((..(((.(((((((....((....))......(((((((((....)))))))))......)))))))..)))...)))).......---------. ( -20.10) >DroSim_CAF1 8313 93 - 1 AUCGAUACAUCUCUCAAUUCUUUUUGGUCGCACUUAACAGCGGGAAAUGGAUUUCCCGCCAAAUUUGAAUUGAAAAGAAAACAUUUUAGAAA---------U .........(((.(((((((..((((((.((........))((((((....))))))))))))...)))))))..)))..............---------. ( -23.30) >DroEre_CAF1 8551 86 - 1 ACCGACAAAUCUCGCGCACCCCUUUGGUCACACUGUACAGCGGGAAAAGGAUUUCCCGCCAAAUUUGAAUUGCAAAGAA-GCAUAUC--------------- .....(((((...(.(.(((.....)))).)........((((((((....))))))))...)))))...(((......-)))....--------------- ( -17.80) >DroYak_CAF1 8627 102 - 1 AUCGAGAAAUAUCACAAUUUCUUUUGGUCACACUAUACAGCGGGAAAUGGAUUUCCCGCCAAAUUUGAACAGAACGGAAAACAUUUCAGAUAAAUUUUACAA (((..(((((.......(((((((((.(((.........((((((((....))))))))......))).))))).))))...))))).)))........... ( -18.56) >consensus AUCGAUAAAUCUCGCAAUUCCUUUUGGUCACACUUAACAGCGGGAAAUGGAUUUCCCGCCAAAUUUGAAUUGAAAAGAAAACAUUUCAGAAA_________A .............(((((((..((((((........)).((((((((....))))))))))))...)))))))............................. (-14.30 = -15.30 + 1.00)



| Location | 11,694,775 – 11,694,876 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 84.36 |
| Mean single sequence MFE | -35.96 |
| Consensus MFE | -23.44 |
| Energy contribution | -24.12 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.17 |
| Mean z-score | -4.25 |
| Structure conservation index | 0.65 |
| SVM decision value | 2.61 |
| SVM RNA-class probability | 0.995726 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11694775 101 + 23771897 AAUUUGGCGGGAAAGCCAUUUCCCGCUGUUAAGUGUGACCAAAAUCAAUUGCGAGAUGUAACAAUAUUUCGCGUGCAAAAAGUUGCGACCACACUGAACUA .....(((((((((....)))))))))(((.((((((.............(((((((((....))))))))).(((((....)))))..)))))).))).. ( -37.90) >DroSec_CAF1 12877 101 + 1 AAUUUAGCGGGAAAUCCAUUUCCCGCUGUUAAGUGUGACCAAAAAAAAUUGAGAGAUCCAUCGAUAUUUCGGUUGCAAAGAGUUGCGACCACACUGAAUUG ....((((((((((....))))))))))...(((((..............((((.(((....))).))))((((((((....)))))))))))))...... ( -34.10) >DroSim_CAF1 8341 101 + 1 AAUUUGGCGGGAAAUCCAUUUCCCGCUGUUAAGUGCGACCAAAAAGAAUUGAGAGAUGUAUCGAUAUUUCGGUUGCAAAGAGUUGCGACCACACUAAAUUA .....(((((((((....)))))))))....((((.................(((((((....)))))))((((((((....)))))))).))))...... ( -33.00) >DroEre_CAF1 8572 101 + 1 AAUUUGGCGGGAAAUCCUUUUCCCGCUGUACAGUGUGACCAAAGGGGUGCGCGAGAUUUGUCGGUAUUCCGGGUGCAAAAAGUUGCGACCACACUGAAUAA .....(((((((((....)))))))))...((((((((((...(((((((.(((......)))))))))).)))((((....))))...)))))))..... ( -43.50) >DroYak_CAF1 8664 101 + 1 AAUUUGGCGGGAAAUCCAUUUCCCGCUGUAUAGUGUGACCAAAAGAAAUUGUGAUAUUUCUCGAUAUUCCGGGAGAGAAAAGUUGCGACCACACUGAAUUA .....(((((((((....)))))))))...(((((((.....((....))(..((.((((((............)))))).))..)...)))))))..... ( -31.30) >consensus AAUUUGGCGGGAAAUCCAUUUCCCGCUGUUAAGUGUGACCAAAAAAAAUUGAGAGAUGUAUCGAUAUUUCGGGUGCAAAAAGUUGCGACCACACUGAAUUA .....(((((((((....)))))))))....((((((......................(((((....))))).((((....))))...))))))...... (-23.44 = -24.12 + 0.68)



| Location | 11,694,775 – 11,694,876 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 84.36 |
| Mean single sequence MFE | -30.62 |
| Consensus MFE | -26.68 |
| Energy contribution | -26.36 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.18 |
| Mean z-score | -3.37 |
| Structure conservation index | 0.87 |
| SVM decision value | 3.59 |
| SVM RNA-class probability | 0.999428 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11694775 101 - 23771897 UAGUUCAGUGUGGUCGCAACUUUUUGCACGCGAAAUAUUGUUACAUCUCGCAAUUGAUUUUGGUCACACUUAACAGCGGGAAAUGGCUUUCCCGCCAAAUU ..(((.((((((..(((((....))))..((((.((........)).))))..........)..)))))).))).((((((((....))))))))...... ( -34.80) >DroSec_CAF1 12877 101 - 1 CAAUUCAGUGUGGUCGCAACUCUUUGCAACCGAAAUAUCGAUGGAUCUCUCAAUUUUUUUUGGUCACACUUAACAGCGGGAAAUGGAUUUCCCGCUAAAUU ......((((((..(((((....))))...(((....))).....................)..))))))....(((((((((....)))))))))..... ( -29.60) >DroSim_CAF1 8341 101 - 1 UAAUUUAGUGUGGUCGCAACUCUUUGCAACCGAAAUAUCGAUACAUCUCUCAAUUCUUUUUGGUCGCACUUAACAGCGGGAAAUGGAUUUCCCGCCAAAUU ......((((((..(((((....))))...(((....))).....................)..)))))).....((((((((....))))))))...... ( -27.50) >DroEre_CAF1 8572 101 - 1 UUAUUCAGUGUGGUCGCAACUUUUUGCACCCGGAAUACCGACAAAUCUCGCGCACCCCUUUGGUCACACUGUACAGCGGGAAAAGGAUUUCCCGCCAAAUU .....(((((((..(((((....))))...(((....))).....................)..)))))))....((((((((....))))))))...... ( -32.60) >DroYak_CAF1 8664 101 - 1 UAAUUCAGUGUGGUCGCAACUUUUCUCUCCCGGAAUAUCGAGAAAUAUCACAAUUUCUUUUGGUCACACUAUACAGCGGGAAAUGGAUUUCCCGCCAAAUU ......((((((..(.......((((.....))))....(((((((......)))))))..)..)))))).....((((((((....))))))))...... ( -28.60) >consensus UAAUUCAGUGUGGUCGCAACUUUUUGCACCCGAAAUAUCGAUAAAUCUCGCAAUUCCUUUUGGUCACACUUAACAGCGGGAAAUGGAUUUCCCGCCAAAUU ......((((((..(((((....))))...(((....))).....................)..)))))).....((((((((....))))))))...... (-26.68 = -26.36 + -0.32)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:16 2006