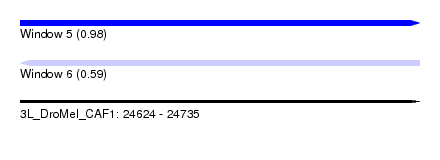

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 24,624 – 24,735 |
| Length | 111 |
| Max. P | 0.979444 |

| Location | 24,624 – 24,735 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.84 |
| Mean single sequence MFE | -20.97 |
| Consensus MFE | -13.84 |
| Energy contribution | -13.40 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.84 |
| SVM RNA-class probability | 0.979444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
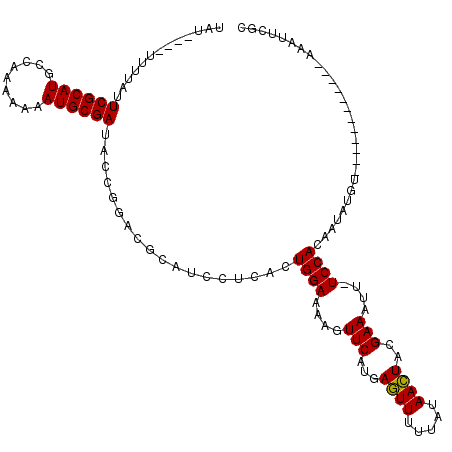
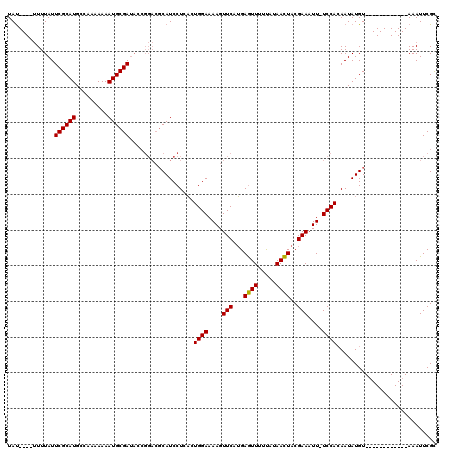
>3L_DroMel_CAF1 24624 111 + 23771897 UACAUAAAUUUAUUCGCAUGUCACAAAAAUGCGA---------CAUCCUCUCUGGAAAAGUUCAAGAAUUUCUAUAAUUACGAAAUUUUCCAGAAUAUGUACAUAUGUAUAUAAAUUCGC .............((((((.........))))))---------.......(((((((((.(((...((((.....))))..))).))))))))).(((((((....)))))))....... ( -24.90) >DroSec_CAF1 14200 103 + 1 UAU----UUUUAUUCGCAUGCCAAAAAAAUGCGAUACCGGACGCAUCCUCACUGGAAAAGUUCAUGAGUUUUCAUAACUACGAAAUU-UCCACAAUAUGU------------AAAUUCGC ...----......((((((.........))))))....((((...(((.....)))...))))..((((((.((((.....((....-)).....)))).------------)))))).. ( -19.40) >DroSim_CAF1 15750 103 + 1 UAU----UUUUAUUCGCAUGCCAAAAAAAUGCGAUACCGGACGCAUCCUCACUGGAAAAGUUCAUGAGUUUUUAUAACUACGAAAUU-UCCACAAUAUGU------------AAAUUCGU ...----......((((((.........))))))....((((...(((.....)))...))))((((((((.((((.....((....-)).....)))).------------)))))))) ( -18.60) >consensus UAU____UUUUAUUCGCAUGCCAAAAAAAUGCGAUACCGGACGCAUCCUCACUGGAAAAGUUCAUGAGUUUUUAUAACUACGAAAUU_UCCACAAUAUGU____________AAAUUCGC .............((((((.........))))))..................((((....(((...((((.....))))..)))....))))............................ (-13.84 = -13.40 + -0.44)

| Location | 24,624 – 24,735 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.84 |
| Mean single sequence MFE | -25.00 |
| Consensus MFE | -14.34 |
| Energy contribution | -14.10 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.588448 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
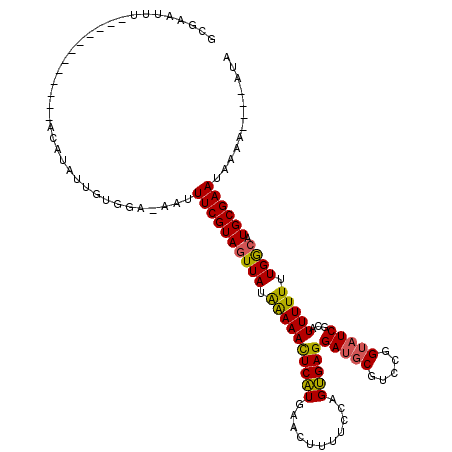
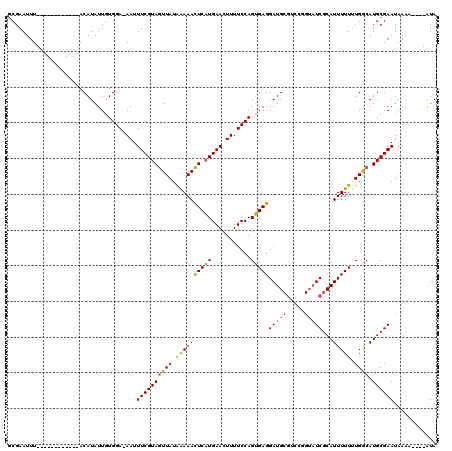
>3L_DroMel_CAF1 24624 111 - 23771897 GCGAAUUUAUAUACAUAUGUACAUAUUCUGGAAAAUUUCGUAAUUAUAGAAAUUCUUGAACUUUUCCAGAGAGGAUG---------UCGCAUUUUUGUGACAUGCGAAUAAAUUUAUGUA ..((((((((...............((((((((((.((((.((((.....))))..)))).)))))))))).(.(((---------(((((....)))))))).)..))))))))..... ( -30.30) >DroSec_CAF1 14200 103 - 1 GCGAAUUU------------ACAUAUUGUGGA-AAUUUCGUAGUUAUGAAAACUCAUGAACUUUUCCAGUGAGGAUGCGUCCGGUAUCGCAUUUUUUUGGCAUGCGAAUAAAA----AUA .....(((------------((.....)))))-...((((((.((((((....))))))......((((.(((((((((........)))))))))))))..)))))).....----... ( -22.40) >DroSim_CAF1 15750 103 - 1 ACGAAUUU------------ACAUAUUGUGGA-AAUUUCGUAGUUAUAAAAACUCAUGAACUUUUCCAGUGAGGAUGCGUCCGGUAUCGCAUUUUUUUGGCAUGCGAAUAAAA----AUA ....(((.------------.(((....((((-((.(((((((((.....)))).)))))..))))))(((((((((((........)))))))))...)))))..)))....----... ( -22.30) >consensus GCGAAUUU____________ACAUAUUGUGGA_AAUUUCGUAGUUAUAAAAACUCAUGAACUUUUCCAGUGAGGAUGCGUCCGGUAUCGCAUUUUUUUGGCAUGCGAAUAAAA____AUA ....................................((((((((((.((((((((((...........)))))(((((.....)))))...))))).)))).))))))............ (-14.34 = -14.10 + -0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:29:42 2006