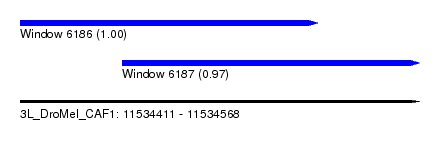

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 11,534,411 – 11,534,568 |
| Length | 157 |
| Max. P | 0.999787 |

| Location | 11,534,411 – 11,534,528 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 70.46 |
| Mean single sequence MFE | -26.44 |
| Consensus MFE | -15.89 |
| Energy contribution | -18.01 |
| Covariance contribution | 2.12 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.60 |
| SVM decision value | 4.08 |
| SVM RNA-class probability | 0.999787 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11534411 117 + 23771897 ACAAAAAGGUUAAAGUCGAAAAUUAACUAAAAAGGUUACCUGGG---UAGCCAGGGUGGAAAUCUUAAUGUUGUUUCCAAAACUCCCAGAAAGGAACAAUUUGGGGAUUAUCAACCACCA .......((((((.........)))))).....((((((....)---)))))..(((((.((((((...((((((((...............))))))))...)))))).....))))). ( -28.56) >DroSec_CAF1 40801 117 + 1 ACAAAAAGGUUGAAGUCGCAAAUUAACUAAAAAGGUUACCUGGG---UAACCAGGGUGGAAAUCUUAAAGUUCUUUCCACAACUCACAUAAAGAAACAAUUUGGGGAUUUUCAACAACCA .......(((((...........(((((.....)))))(((((.---...))))).(((((((((..(((((.((((...............)))).)))))..))))))))).))))). ( -30.46) >DroEre_CAF1 35118 110 + 1 ACAAAAAGGGUGCAGUAGC------ACCAAAAAGGUUACCUGGAAGUUCUACAGGGUGGAAUUCUUAAAGUUGUUUCCCAAACUCCUAAAAGGGAACAUUUUGGAGAUAGUCAACC---- ........(((((....))------))).....((((.((((.........))))...((..((((.(((.(((((((.............))))))).))).))))...))))))---- ( -28.02) >DroYak_CAF1 39679 95 + 1 A-------------------------CCAAAAAGGUUACCUAAAAGUCCUACAGGGUGGAAAUCUCAAAAUUGUUUCCAAAACUUCCAAAAAGAAUCAUUUUGGAGAUAGUCAACCACCA (-------------------------((.....)))..................(((((..........(((((((((((((.(((......)))...)))))))))))))...))))). ( -18.72) >consensus ACAAAAAGGUUGAAGUCGC______ACCAAAAAGGUUACCUGGA___UAUACAGGGUGGAAAUCUUAAAGUUGUUUCCAAAACUCCCAAAAAGAAACAAUUUGGAGAUAGUCAACCACCA .................................((((.(((((.......)))))..((.((((((.((((((((((...............)))))))))).)))))).)))))).... (-15.89 = -18.01 + 2.12)

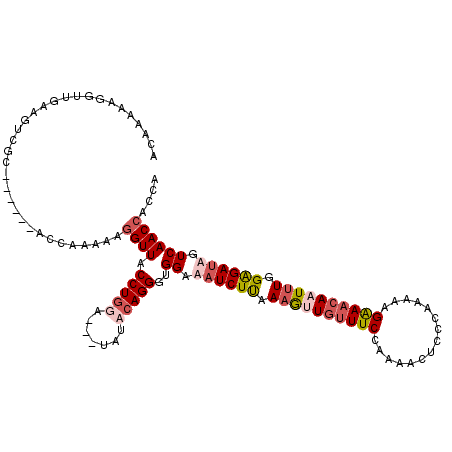
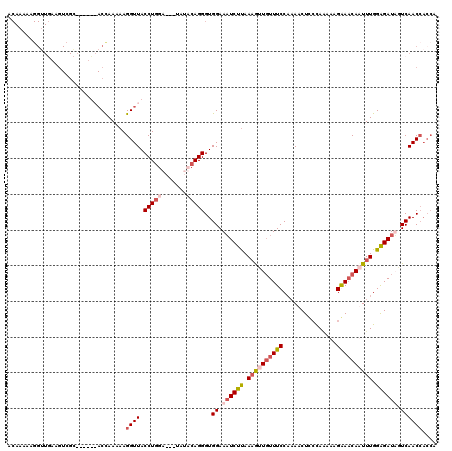
| Location | 11,534,451 – 11,534,568 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.03 |
| Mean single sequence MFE | -26.80 |
| Consensus MFE | -18.41 |
| Energy contribution | -19.04 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969560 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11534451 117 + 23771897 UGGG---UAGCCAGGGUGGAAAUCUUAAUGUUGUUUCCAAAACUCCCAGAAAGGAACAAUUUGGGGAUUAUCAACCACCACCCCAACAUCUUGACCGGUUUUUACCUUCACAACCCUUAU .(((---(.(..((((((((((((.....((((((((...............))))))))((((((..............))))))..........))))))))))))..).)))).... ( -31.10) >DroSec_CAF1 40841 117 + 1 UGGG---UAACCAGGGUGGAAAUCUUAAAGUUCUUUCCACAACUCACAUAAAGAAACAAUUUGGGGAUUUUCAACAACCACCCCAACAUCUUGACCGGUUUUUACCUUCACAACCCUUAU .(((---(....((((((((((((......((((((.............)))))).....((((((..............))))))..........))))))))))))....)))).... ( -27.16) >DroEre_CAF1 35152 114 + 1 UGGAAGUUCUACAGGGUGGAAUUCUUAAAGUUGUUUCCCAAACUCCUAAAAGGGAACAUUUUGGAGAUAGUCAACC------CCAACUUCUUGUCCGGUUUUUACCUUCACAACCCUUAU .(((((((.....((((.((..((((.(((.(((((((.............))))))).))).))))...)).)))------).)))))))(((..(((....)))...)))........ ( -25.82) >DroYak_CAF1 39694 120 + 1 UAAAAGUCCUACAGGGUGGAAAUCUCAAAAUUGUUUCCAAAACUUCCAAAAAGAAUCAUUUUGGAGAUAGUCAACCACCAACCCAACAGCUUGUCCGGUUUUUACCUUCACAACCCUUAA ............(((((.(((........(((((((((((((.(((......)))...)))))))))))))........((((..((.....))..))))......)))...)))))... ( -23.10) >consensus UGGA___UAUACAGGGUGGAAAUCUUAAAGUUGUUUCCAAAACUCCCAAAAAGAAACAAUUUGGAGAUAGUCAACCACCACCCCAACAUCUUGACCGGUUUUUACCUUCACAACCCUUAU ............(((((.((((((((.((((((((((...............)))))))))).)))))............................(((....))))))...)))))... (-18.41 = -19.04 + 0.63)


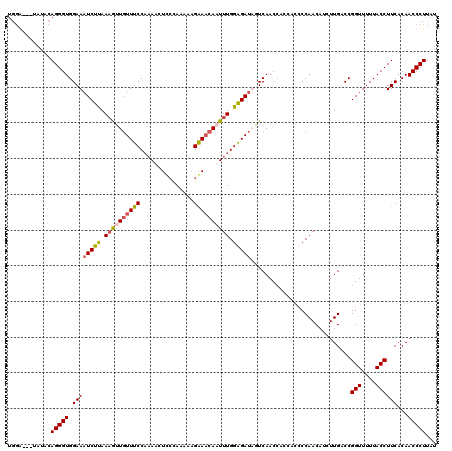
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:11:21 2006