
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 11,526,384 – 11,526,520 |
| Length | 136 |
| Max. P | 0.995118 |

| Location | 11,526,384 – 11,526,487 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 83.15 |
| Mean single sequence MFE | -42.05 |
| Consensus MFE | -31.17 |
| Energy contribution | -32.92 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.06 |
| Structure conservation index | 0.74 |
| SVM decision value | 2.54 |
| SVM RNA-class probability | 0.995118 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11526384 103 + 23771897 GACGC------UGGAGUAGCAGCUUUGCUGCAGCGUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGA (((((------((.(((((.....))))).))))))).(((((((.(((((((..........((((((.(.((((....))))))))))))))))))...))))))). ( -45.90) >DroSec_CAF1 32556 98 + 1 GCCGCUGCUGCUGGAGUAGCA-----------GCAUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGA ...((((((((....))))))-----------))....(((((((.(((((((..........((((((.(.((((....))))))))))))))))))...))))))). ( -45.10) >DroSim_CAF1 31928 98 + 1 GCCGCUGCUGCUGGAGUAGCA-----------GCAUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGA ...((((((((....))))))-----------))....(((((((.(((((((..........((((((.(.((((....))))))))))))))))))...))))))). ( -45.10) >DroYak_CAF1 31074 86 + 1 GCUGC-----------------------AGCAGUGUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGA ...((-----------------------(((...)))))((((((.(((((((..........((((((.(.((((....))))))))))))))))))...)))))).. ( -32.10) >consensus GCCGC______UGGAGUAGCA___________GCAUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGA ...((((((.....))))))..................(((((((.(((((((..........((((((.(.((((....))))))))))))))))))...))))))). (-31.17 = -32.92 + 1.75)
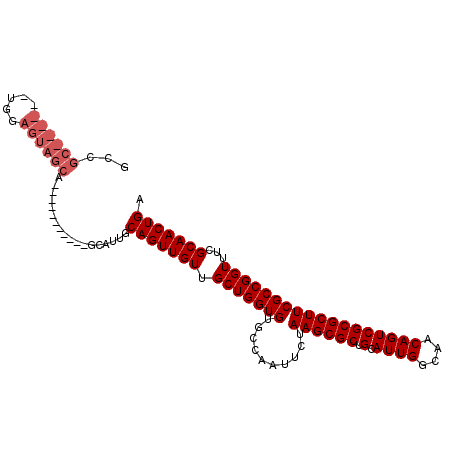



| Location | 11,526,410 – 11,526,520 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 98.19 |
| Mean single sequence MFE | -38.23 |
| Consensus MFE | -37.17 |
| Energy contribution | -37.50 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.832338 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11526410 110 + 23771897 GCGUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGAACUCUCAGCUGCUAUUGCUCCUCUGCAGUUAGU- ..(((.(((((((.(((((((..........((((((.(.((((....))))))))))))))))))...))))))))))....((((((............))))))...- ( -38.80) >DroSec_CAF1 32577 110 + 1 GCAUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGAACUCUCAGCUGCUAUUGCUCCUCUGCAGUUAGU- ..(((((((..(..(((((((..........((((((.(.((((....))))))))))))))))))..)(((.((((....)))).)))..........)))))))....- ( -37.10) >DroYak_CAF1 31083 111 + 1 GUGUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGAACUCUCAGCUGCUAUUGCUCCUCUGCAGUUAGUU (.(((.(((((((.(((((((..........((((((.(.((((....))))))))))))))))))...)))))))))).)..((((((............)))))).... ( -38.80) >consensus GCGUUGCAGUUGUUGCUGGUGUGCCAAUUCUAAGCGCUGCAUUGGCAACAGUCGCGCUUCGCCGGUUUCGCAACUGAACUCUCAGCUGCUAUUGCUCCUCUGCAGUUAGU_ (.(((.(((((((.(((((((..........((((((.(.((((....))))))))))))))))))...)))))))))).)..((((((............)))))).... (-37.17 = -37.50 + 0.33)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:11:19 2006