
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 11,402,463 – 11,402,596 |
| Length | 133 |
| Max. P | 0.953374 |

| Location | 11,402,463 – 11,402,565 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 81.22 |
| Mean single sequence MFE | -26.10 |
| Consensus MFE | -18.98 |
| Energy contribution | -19.10 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.953374 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11402463 102 + 23771897 AG-ACAUAGUUCUCGAGCGCUGAGAACUAAGAGAUCAGCUUUCUCUUGAGUGAUCGCGGCAAGCUCAAGAGUGUCAAAAAAUUGAAAAAAAAGAGAGCAACAA ..-.....((((((((((((((.(((((((((((.......)))))).)))..)).))))..)))).......((((....)))).......))))))..... ( -29.40) >DroSec_CAF1 122674 101 + 1 AG-ACAUAGUUCUUGAGCGCUGAGAACUAAGAGAUCGGCUUUCUCUUGAGUGAUCGCCGCUAACUCAAAAGAGUCAAAAAAUUUAAA-AAAAGAGAGCAACAA ..-.....((((((.((((.(((..(((((((((.......)))))).)))..))).)))).((((....)))).............-....))))))..... ( -26.80) >DroSim_CAF1 134791 101 + 1 AG-ACAUAGUUCUUGAGCGCUGAGAACUAAGAGAUCGGCUUUCUCUUGAGUGAUCGCGGCAAACUCAAGAGAGUCAAAAAAUUGUAA-AAAAGAGAGCAACAA ..-.....((((((....(((((...........))))).(((((((((((...........)))))))))))..............-....))))))..... ( -26.40) >DroEre_CAF1 129571 98 + 1 AGCACAUAGUUCUUGAGCUCUGAGAACUAAGAGUUCAGGUUUCUCUUGAGUGAUCGCGG----CUUAAGAGUGUCGGUAAACCAAGU-GUAAGAGAGCCACAC .((((......((((((((((........))))))))))...(((((((((.......)----))))))))....((....))..))-))............. ( -28.70) >DroYak_CAF1 120357 98 + 1 GGCACAUAGUUCUUGAGUUCUGAGAAAUAAGAGUUCAGCUUUCUCUUGAGUGAUCGCGU----UUUAAGAGAGUCAAAAAACCGAGU-UAAAGAGAUUCAAUU (((.........((((..(((........)))..))))...(((((((((.........----))))))))))))........((((-(.....))))).... ( -19.20) >consensus AG_ACAUAGUUCUUGAGCGCUGAGAACUAAGAGAUCAGCUUUCUCUUGAGUGAUCGCGGC_A_CUCAAGAGAGUCAAAAAAUUGAAA_AAAAGAGAGCAACAA ........((((((....(((((...........))))).(((((((((((...........)))))))))))...................))))))..... (-18.98 = -19.10 + 0.12)



| Location | 11,402,499 – 11,402,596 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 81.05 |
| Mean single sequence MFE | -22.39 |
| Consensus MFE | -15.70 |
| Energy contribution | -16.70 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.593444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11402499 97 + 23771897 GCUUUCUCUUGAGUGAUCGCGGCAAGCUCAAGAGUGUCAAAAAAUUGAAAAAAAAGAGAGCAACAAUUUCCACUCAAUUGCUCUUAAAU--UGGCGUGA ((((((((((((((...........))))))))...((((....)))).......)))))).........(((((((((.......)))--))).))). ( -24.00) >DroSec_CAF1 122710 96 + 1 GCUUUCUCUUGAGUGAUCGCCGCUAACUCAAAAGAGUCAAAAAAUUUAAA-AAAAGAGAGCAACAAUUUUCACUCAAUUGCUCUUAAAU--UGGCGUGA (((((((....((((.....)))).((((....)))).............-...)))))))........((((((((((.......)))--))).)))) ( -20.60) >DroSim_CAF1 134827 96 + 1 GCUUUCUCUUGAGUGAUCGCGGCAAACUCAAGAGAGUCAAAAAAUUGUAA-AAAAGAGAGCAACAAAUUCCACUCAAUUGCUCUUAAAU--UGGCGUGA ((((((((((((((...........)))))))))))..............-....((((((((..............))))))))....--.))).... ( -24.84) >DroEre_CAF1 129608 94 + 1 GGUUUCUCUUGAGUGAUCGCGG----CUUAAGAGUGUCGGUAAACCAAGU-GUAAGAGAGCCACACUUUUCACUCAAUUGCUCUUAAAUUGUGGCGUGA ((((.((....)).))))((.(----(((((((((...((....)).(((-(.(((((.......))))))))).....))))))))...)).)).... ( -22.30) >DroYak_CAF1 120394 93 + 1 GCUUUCUCUUGAGUGAUCGCGU----UUUAAGAGAGUCAAAAAACCGAGU-UAAAGAGAUUCAAUUUUCUCACUCAAUUGCUCUUAAAC-AUAGCGUGA ((((......))))..((((((----((((((((.((......)).((((-....((((........)))))))).....)))))))).-...)))))) ( -20.20) >consensus GCUUUCUCUUGAGUGAUCGCGGC_A_CUCAAGAGAGUCAAAAAAUUGAAA_AAAAGAGAGCAACAAUUUUCACUCAAUUGCUCUUAAAU__UGGCGUGA ((((((((((((((...........)))))))))))...................((((((((..............)))))))).......))).... (-15.70 = -16.70 + 1.00)

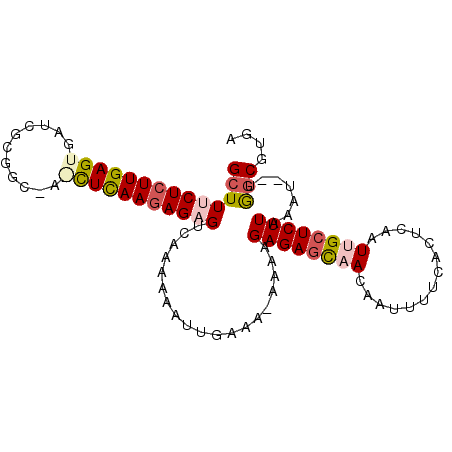

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:10:14 2006