
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 11,246,918 – 11,247,027 |
| Length | 109 |
| Max. P | 0.751796 |

| Location | 11,246,918 – 11,247,027 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 87.92 |
| Mean single sequence MFE | -27.45 |
| Consensus MFE | -19.99 |
| Energy contribution | -20.30 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751796 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 11246918 109 + 23771897 AUAAGAUGCUAAUGUUUUGACCAUAGCUGUGCAGACUUUAAAAUAGAACGCACUUAAAUUGAGUGCGAAUUGUUAGAGGCACUCUAACAGCCGGCAAACCUAUUUUCAA .((((((......))))))......((((.((................(((((((.....)))))))...((((((((...)))))))))))))).............. ( -28.00) >DroSec_CAF1 48827 109 + 1 AUUCGAUUCUAAUGUUUUGGCCAUAGCUGUGCAGACUUUAAAAUAGAACGCACUUAAAUUGAGUGCGAAUUGUUAGAGGCACUUCUACUGCCGGUAAACCUAUUUUCGA ..((((............(((..(((..((((...((((((.......(((((((.....))))))).....))))))))))..)))..)))((....)).....)))) ( -29.20) >DroSim_CAF1 50492 109 + 1 AUACGAUUCUAAGUUCUUGACCAUAGCUGUGCAGACUUUAAAAUAGAACGCACUUAAAUUGAGUGCGAAUUGUUAGAGGCACUCUAACUGCCGGCAAACCUAUUUUCGA ...(((......(((...)))....((((.((((..............(((((((.....))))))).....((((((...))))))))))))))..........))). ( -26.60) >DroYak_CAF1 51848 107 + 1 GUAAGAUGCUAAUUUUUCAGCCAUUGCUGUGCAGACUUUAAAAUAGAACGCACUUAAGUUGAGUGCGAAUUGU-AGAG-CUCCAUUUCUGCCGGCAAACCAAUUUUCGA (.((((((((........)))..((((((.((((((((((.(((....(((((((.....))))))).))).)-))))-.......)))))))))))....)))))).. ( -26.01) >consensus AUAAGAUGCUAAUGUUUUGACCAUAGCUGUGCAGACUUUAAAAUAGAACGCACUUAAAUUGAGUGCGAAUUGUUAGAGGCACUCUAACUGCCGGCAAACCUAUUUUCGA .........................((((.((((.((((((.......(((((((.....))))))).....)))))).........)))))))).............. (-19.99 = -20.30 + 0.31)


| Location | 11,246,918 – 11,247,027 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 87.92 |
| Mean single sequence MFE | -28.00 |
| Consensus MFE | -18.86 |
| Energy contribution | -20.30 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.737876 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
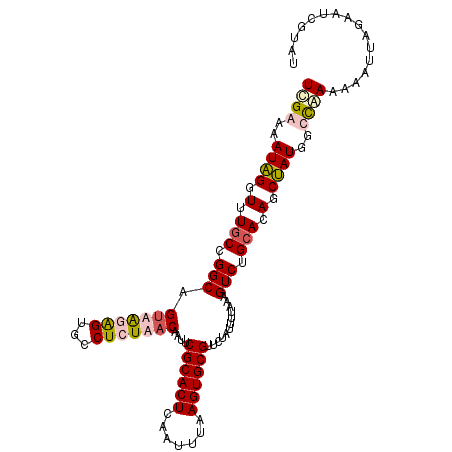
>3L_DroMel_CAF1 11246918 109 - 23771897 UUGAAAAUAGGUUUGCCGGCUGUUAGAGUGCCUCUAACAAUUCGCACUCAAUUUAAGUGCGUUCUAUUUUAAAGUCUGCACAGCUAUGGUCAAAACAUUAGCAUCUUAU ((((..((((.(.(((.(((((((((((...)))))))....((((((.......))))))...........)))).))).).))))..))))................ ( -30.30) >DroSec_CAF1 48827 109 - 1 UCGAAAAUAGGUUUACCGGCAGUAGAAGUGCCUCUAACAAUUCGCACUCAAUUUAAGUGCGUUCUAUUUUAAAGUCUGCACAGCUAUGGCCAAAACAUUAGAAUCGAAU ((((.....((....))(((.((((..((((((.(((.(((.((((((.......))))))....)))))).))...))))..)))).)))............)))).. ( -27.10) >DroSim_CAF1 50492 109 - 1 UCGAAAAUAGGUUUGCCGGCAGUUAGAGUGCCUCUAACAAUUCGCACUCAAUUUAAGUGCGUUCUAUUUUAAAGUCUGCACAGCUAUGGUCAAGAACUUAGAAUCGUAU .(((...(((((((((((..((((.((((((............)))))).......(((((..((.......))..))))))))).))))...)))))))...)))... ( -28.80) >DroYak_CAF1 51848 107 - 1 UCGAAAAUUGGUUUGCCGGCAGAAAUGGAG-CUCU-ACAAUUCGCACUCAACUUAAGUGCGUUCUAUUUUAAAGUCUGCACAGCAAUGGCUGAAAAAUUAGCAUCUUAC ......((((.(.(((.(((.(((((((((-(...-.......(((((.......)))))))))))))))...))).))).).)))).(((((....)))))....... ( -25.80) >consensus UCGAAAAUAGGUUUGCCGGCAGUAAGAGUGCCUCUAACAAUUCGCACUCAAUUUAAGUGCGUUCUAUUUUAAAGUCUGCACAGCUAUGGCCAAAAAAUUAGAAUCGUAU ((((..((((.(.(((.(((.(((((((...)))))))....((((((.......))))))............))).))).).))))..))))................ (-18.86 = -20.30 + 1.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:09:04 2006