
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,957,291 – 10,957,396 |
| Length | 105 |
| Max. P | 0.608980 |

| Location | 10,957,291 – 10,957,396 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.09 |
| Mean single sequence MFE | -32.29 |
| Consensus MFE | -20.97 |
| Energy contribution | -21.58 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.608980 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
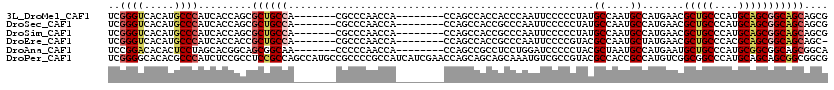
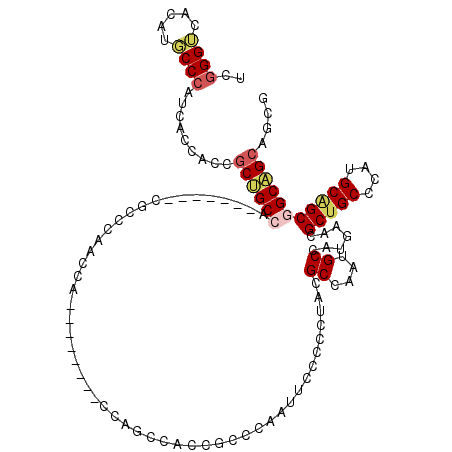
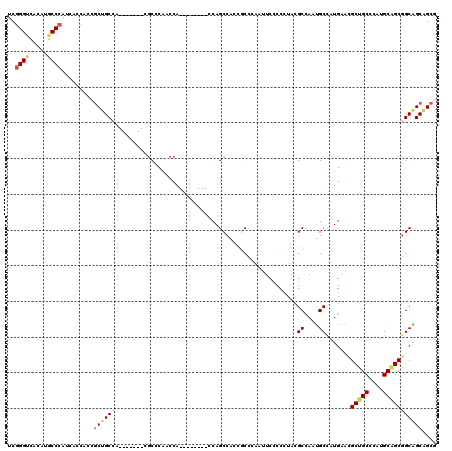
>3L_DroMel_CAF1 10957291 105 - 23771897 UCGGGUCACAUGCCCAUCACCAGCGCUGCCA-------CGCCCAACCA--------CCAGCCACCACCCAAUUCCCCCUAUGCCAAUGCCAUGAACGCUGCCCAUGCAGCGGCAGCAGCG ..((((.....)))).......((((((((.-------..........--------.........................((....)).......(((((....))))))))))).)). ( -28.70) >DroSec_CAF1 89553 105 - 1 UCGGGUCACAUGCCCAUCACCAGCGCUGCCA-------CGCCCAACCA--------CCAGCCACCGCCCAAUUCCCCCUAUGCCAAUGCCAUGAACGCUGCCCAUGCAGCGGCAGCAGCG ..((((.....)))).......((((((((.-------..........--------...((....)).............................(((((....))))))))))).)). ( -29.00) >DroSim_CAF1 86989 105 - 1 UCGGGUCACAUGCCCAUCACCAGCGCUGCCA-------CGCCCAACCA--------CCAGCCACCGCCCAAUUCCCCCUAUGCCAAUGCCAUGAACGCUGCCCAUGCAGCGGCAGCAGCG ..((((.....)))).......((((((((.-------..........--------...((....)).............................(((((....))))))))))).)). ( -29.00) >DroEre_CAF1 90021 104 - 1 UCGGGUCACAUGCCCAUCACCACCGCUGCCA-------CGCCCAACCA--------CCAGCCACCGCCCAAUUCCCCGUACGCCAAUGCUAUGAACGCUGCCCACGCAGCGGCAGCAGC- ..((((.....)))).........((((((.-------..........--------....................((((.((....))))))...(((((....)))))))))))...- ( -29.90) >DroAna_CAF1 84067 105 - 1 UCCGGACACACUCCUAGCACGGCAGCGGCAA-------CCCCCAACCA--------CCAGCCGCCUCCUGGAUCCCCCUACGCUAAUGCCAUGAAUGCUGCCCAUGCGGCGGCAGCGGCA ...(((.....)))((((..((..(((((..-------..........--------...)))))..)).((.....))...)))).((((.....(((((((.....)))))))..)))) ( -33.86) >DroPer_CAF1 81383 120 - 1 UCGGGGCACACGCCCAUCUCCGCCUCCGCCAGCCAUGCCGCCCCGCCAUCAUCGAACCAGCAGCAGCAAAUGUCGCCGUACGCCACCGCCAUGUCGGCGGCCCAUGCAGCAGCGGCGGCG ...((((....)))).....((((.((((..((..(((.((..................)).)))(((...(((((((.((((....))...)))))))))...))).)).)))).)))) ( -43.27) >consensus UCGGGUCACAUGCCCAUCACCACCGCUGCCA_______CGCCCAACCA________CCAGCCACCGCCCAAUUCCCCCUACGCCAAUGCCAUGAACGCUGCCCAUGCAGCGGCAGCAGCG ..((((.....)))).........((((((...................................................((....)).......(((((....))))))))))).... (-20.97 = -21.58 + 0.61)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:52 2006