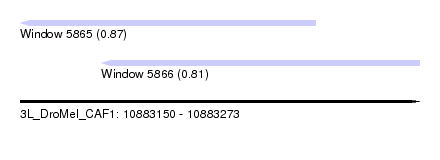

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,883,150 – 10,883,273 |
| Length | 123 |
| Max. P | 0.867411 |

| Location | 10,883,150 – 10,883,241 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 76.97 |
| Mean single sequence MFE | -24.53 |
| Consensus MFE | -10.52 |
| Energy contribution | -11.83 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.43 |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.867411 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
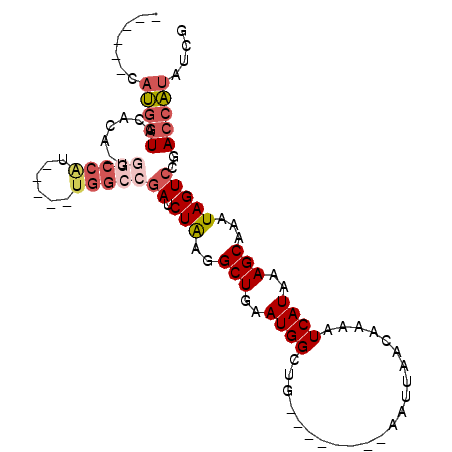
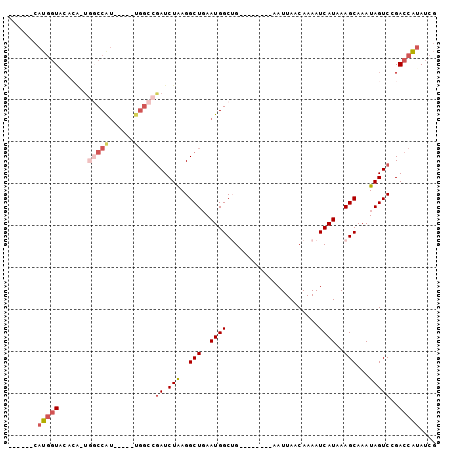
>3L_DroMel_CAF1 10883150 91 - 23771897 ------CAUGGUACCCGUUGGCCAU-----UGGCCGAUCUAAGGCUGAAUGGCUGAAUGGUUGAAUUAACAAAAUCAUAAAGCAAAUAGUCCGACCAUAUCG ------.(((((....(((((((..-----.)))))))....(((((....(((..((((((..........))))))..)))...)))))..))))).... ( -25.00) >DroSec_CAF1 15508 90 - 1 ------CAUGGUACACA-UGGCCAU-----UGGCCGAUCUAAGGCUGAAUGGCUGAAUGGUUGAAUUAACAAAAUCAUAAAGCAAAUAGUCCGACCAUAUCG ------((((.....))-))(((((-----(((((.......)))).)))))).(((((((((.((((..................)))).))))))).)). ( -25.67) >DroSim_CAF1 11491 90 - 1 ------CAUGGUACCCA-UGGCCAU-----UGGCCGAUCUAAGGCUGAAUGGCUGAAUGGUUGAAUUAACAAAAUCAUAAAGCAAAUAGUCCGACCAUAUCG ------(((((...)))-))(((((-----(((((.......)))).)))))).(((((((((.((((..................)))).))))))).)). ( -26.47) >DroEre_CAF1 10793 80 - 1 ------CAUGGUC--------CCGUGGCCAUGGCCGAUCUAAGGCUCAAUGGCAG--------AAUUAACAAAAUCAUAAAGCAAAUAGUCGGACCAUAUCG ------.((((((--------(....(((((((((.......))))..))))).(--------(.((....)).))...............))))))).... ( -22.70) >DroYak_CAF1 12407 93 - 1 GAUAGCCAUGGUCCGCA-UGGCCAUGUCCAUGGCCAAUCUAAGGCUCAAUGGCAG--------AAUUAACAAAAUCAUAAAGCAAAUAGUCGGACCAUAUCG ((((....((((((((.-(((((((....))))))).......(((..((((...--------...........))))..))).....).))))))))))). ( -28.84) >DroAna_CAF1 5138 83 - 1 ------CAGACUCCAAG-U-CCCAGCUCCGACUCCGAUCUGAGGCUGAAUGGCUG--------AAUUAACAAAAUCAUAAAGCAAAUAGUCGGACCUGA--- ------..(((.....)-)-).(((.(((((((.(.......)(((..(((..(.--------.........)..)))..)))....))))))).))).--- ( -18.50) >consensus ______CAUGGUACACA_UGGCCAU_____UGGCCGAUCUAAGGCUGAAUGGCUG________AAUUAACAAAAUCAUAAAGCAAAUAGUCCGACCAUAUCG .......(((((.......(((((......)))))((.(((..(((..((((......................))))..)))...)))))..))))).... (-10.52 = -11.83 + 1.31)

| Location | 10,883,175 – 10,883,273 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 77.03 |
| Mean single sequence MFE | -26.57 |
| Consensus MFE | -11.89 |
| Energy contribution | -13.58 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.808553 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
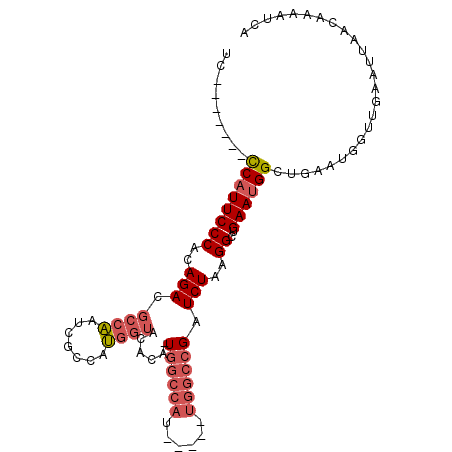
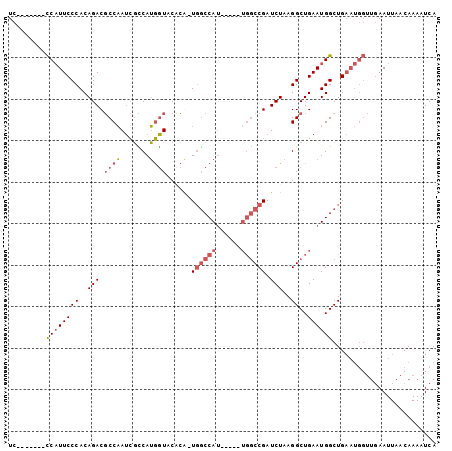
>3L_DroMel_CAF1 10883175 98 - 23771897 UCCCAUAUCCCAUUCCCACAGACGCCAAUCGCCAUGGUACCCGUUGGCCAU-----UGGCCGAUCUAAGGCUGAAUGGCUGAAUGGUUGAAUUAACAAAAUCA ..((((.((((((((((..((((((((((.(((((((...))).)))).))-----))).)).)))..))..))))))..))))))................. ( -29.00) >DroSec_CAF1 15533 90 - 1 UC-------CCAUUCCCACAGACGCCAAUCGUCAUGGUACACA-UGGCCAU-----UGGCCGAUCUAAGGCUGAAUGGCUGAAUGGUUGAAUUAACAAAAUCA ..-------((((((....((((((((((.((((((.....))-)))).))-----))).)).)))..((((....))))))))))................. ( -27.90) >DroSim_CAF1 11516 90 - 1 UC-------CCAUUCCCACAGACGCCAAUCGCCAUGGUACCCA-UGGCCAU-----UGGCCGAUCUAAGGCUGAAUGGCUGAAUGGUUGAAUUAACAAAAUCA ..-------((((((....((((((((((.(((((((...)))-)))).))-----))).)).)))..((((....))))))))))................. ( -31.40) >DroAna_CAF1 5160 86 - 1 UC-------UCUUUCCCACAGACUCCCAGUUGCAGACUCCAAG-U-CCCAGCUCCGACUCCGAUCUGAGGCUGAAUGGCUG--------AAUUAACAAAAUCA ..-------.........(((.((..((((..((((.((..((-(-(........))))..))))))..))))...)))))--------.............. ( -18.00) >consensus UC_______CCAUUCCCACAGACGCCAAUCGCCAUGGUACACA_UGGCCAU_____UGGCCGAUCUAAGGCUGAAUGGCUGAAUGGUUGAAUUAACAAAAUCA .........((((((((..(((.((((.......))))......((((((......)))))).)))..))..))))))......................... (-11.89 = -13.58 + 1.69)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:00 2006