
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,759,304 – 10,759,442 |
| Length | 138 |
| Max. P | 0.698277 |

| Location | 10,759,304 – 10,759,410 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 85.31 |
| Mean single sequence MFE | -20.90 |
| Consensus MFE | -13.68 |
| Energy contribution | -13.80 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.698277 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10759304 106 + 23771897 AGUAAAAAAAUGAUAAUAAAAUCUCUUGGGUAACAAAAUUUGUUGUUUGUUGUUGCAAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACAGAAGAAAA--AA .....................(((..((..((((((....(((.((((((....)))))))))....))))))((((....)))).........))..)))...--.. ( -17.80) >DroSec_CAF1 93040 104 + 1 AGUAAAAAAAUGAUAAUAAAAUCUCUUGGGUAACACAAUUUGUUGUUUGUUGUUGCUAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACCGAAAGAAA---- .....................(((.(((((((((((((........))).))))))...........((((..((((....))))..))))...)))).)))..---- ( -21.90) >DroSim_CAF1 95853 104 + 1 AGUAAAAAAAUGAUAAUAAAAUCUCUUGGGUAACACAAUUUGUUGUUUGUUGUUGCUAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACCGAAAGAAA---- .....................(((.(((((((((((((........))).))))))...........((((..((((....))))..))))...)))).)))..---- ( -21.90) >DroEre_CAF1 97738 93 + 1 -----------AAUAUUAAAAUCCCUUGGGUAACAAAAUUUGUUGUCUGCUGUUGCUAAUACAAUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACAGAAG----AAAA -----------.........(((((((..(((((((...(((((((..((....))....)))))))...)))))))..))).)))).............----.... ( -22.00) >consensus AGUAAAAAAAUGAUAAUAAAAUCUCUUGGGUAACAAAAUUUGUUGUUUGUUGUUGCUAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACAGAAAGAAA____ ....................(((((((..(((((((((((...(((..((....))....)))..))))).))))))..))).))))..................... (-13.68 = -13.80 + 0.12)


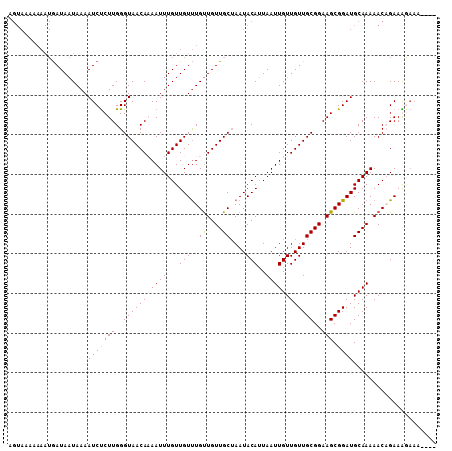
| Location | 10,759,339 – 10,759,442 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 82.62 |
| Mean single sequence MFE | -18.48 |
| Consensus MFE | -16.08 |
| Energy contribution | -15.60 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.696253 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
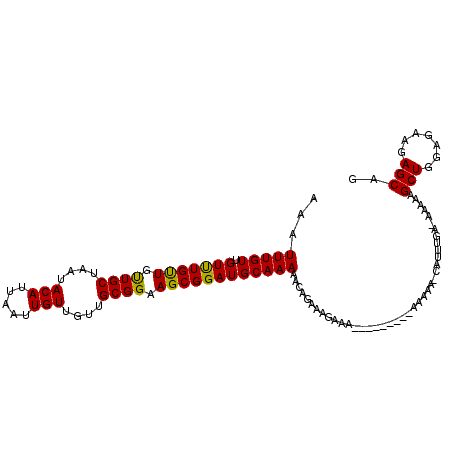
>3L_DroMel_CAF1 10759339 103 + 23771897 AAAUUUGUUGUUUGUUGUUGCAAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACAGAAGAAAA---------AAAAAACAUUUGAAAAAAAGCUGGAGAAGAGCAG ...(((....((((((.(((((...(((.....)))..((((....)))).))))).))))))....))---------)..................(((.......))).. ( -18.00) >DroSec_CAF1 93075 89 + 1 CAAUUUGUUGUUUGUUGUUGCUAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACCGAAAGAAA-----------------------AAAAAGCUGGAGAAGAGCAG .....((((.(((......(((..........((((..((((....))))..))))....(....)...-----------------------....)))....))).)))). ( -16.80) >DroSim_CAF1 95888 89 + 1 CAAUUUGUUGUUUGUUGUUGCUAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACCGAAAGAAA-----------------------AAAAAGCUGGAGAAGAGCAG .....((((.(((......(((..........((((..((((....))))..))))....(....)...-----------------------....)))....))).)))). ( -16.80) >DroEre_CAF1 97762 100 + 1 AAAUUUGUUGUCUGCUGUUGCUAAUACAAUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACAGAAG-----------AAAAAAAGCAUUUGA-AAAAAGCUGGAGAAGAGCAG ...(((((.(((((((.((((....(((((....))))).)))).))))))))))))........-----------......(((.(((..-..))))))............ ( -21.60) >DroYak_CAF1 108809 111 + 1 AAAUUUGUUGUUUGUUGCUGCUAAUACAAUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACAGAAAGAAAAAACAGAAAAAAAAACAUUUGA-AAAAAGCUGGAGAAGAGCAG ...((((((.((((((..(((....(((((....)))))(((....)))...)))..)))))).......))))))...............-.....(((.......))).. ( -19.20) >consensus AAAUUUGUUGUUUGUUGUUGCUAAUACAUUAAUUGUUGUUGCGGAAGCGGAUGCAAAAACAGAAAGAAA_________AAAAA_CAUUUGA_AAAAAGCUGGAGAAGAGCAG ...(((((.(((((((.((((....(((.....)))....)))).))))))))))))........................................(((.......))).. (-16.08 = -15.60 + -0.48)

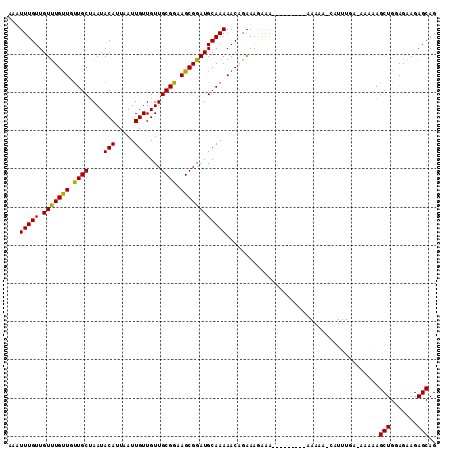
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:04:25 2006