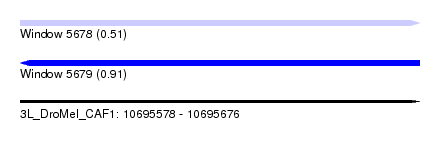

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,695,578 – 10,695,676 |
| Length | 98 |
| Max. P | 0.906150 |

| Location | 10,695,578 – 10,695,676 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 96.28 |
| Mean single sequence MFE | -26.66 |
| Consensus MFE | -23.64 |
| Energy contribution | -23.64 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.20 |
| Structure conservation index | 0.89 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.511588 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10695578 98 + 23771897 UUUUU--UUUUUAGGUGGUAAAUGGUGUUGGUGGUGCUGCUGCUGCUGGUGAUGCUUCCUGCGCUGCCGUUUAGAGAGGCUGCCAAAAAACAUUUAUACA .....--.......(((.((((((...((((..((.((.(((..((.(((..(((.....)))..))))).)))))..))..))))....))))))))). ( -27.30) >DroSec_CAF1 29978 93 + 1 -UUUU---UUUUAGGUGGUAAAUGGUGUUGGUGGUGCUG---CUGCUGGUGAUGCUUCCUGCGCUGCCGUUUAGAGAGGCUGCCAAAAAACAUUUAUACA -....---......(((.((((((...((((..((.((.---(((.((((..(((.....)))..))))..)))))..))..))))....))))))))). ( -26.70) >DroSim_CAF1 31628 94 + 1 -UUUU--UUUUUAGGUGGUAAAUGGUGUUGGUGGUGCUG---CUGCUGGUGAUGCUUCCUGCGCUGCCGUUUAGAGAGGCUGCCAAAAAACAUUUAUACA -....--.......(((.((((((...((((..((.((.---(((.((((..(((.....)))..))))..)))))..))..))))....))))))))). ( -26.70) >DroEre_CAF1 33554 96 + 1 -UUUUUUUUUUUAGGUGGUAAAUGGUGUUGGUGGUGGUG---CUGCUGGUGAUGCUUCCUGCGCUGCCGUUUAGAGAGGCUGCCAAAAAACAUUUAUACA -.............(((.((((((...(((((((..(((---(.(..((.....))..).))))..))((((....)))).)))))....))))))))). ( -26.30) >DroYak_CAF1 42779 95 + 1 -UUUUU-UUUUUAGGUGGUAAAUGGUGUUGGUGGUGGUG---CUGCUGGUGAUGCUUCCUGCGCUGCCGUUUAGAGAGGCUGCCAAAAAACAUUUAUACA -.....-.......(((.((((((...(((((((..(((---(.(..((.....))..).))))..))((((....)))).)))))....))))))))). ( -26.30) >consensus _UUUU__UUUUUAGGUGGUAAAUGGUGUUGGUGGUGCUG___CUGCUGGUGAUGCUUCCUGCGCUGCCGUUUAGAGAGGCUGCCAAAAAACAUUUAUACA ..............(((.((((((...((((..((..(......((.(((..(((.....)))..))))).....)..))..))))....))))))))). (-23.64 = -23.64 + 0.00)



| Location | 10,695,578 – 10,695,676 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 96.28 |
| Mean single sequence MFE | -16.32 |
| Consensus MFE | -15.80 |
| Energy contribution | -15.80 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.906150 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10695578 98 - 23771897 UGUAUAAAUGUUUUUUGGCAGCCUCUCUAAACGGCAGCGCAGGAAGCAUCACCAGCAGCAGCAGCACCACCAACACCAUUUACCACCUAAAAA--AAAAA (((....((((((((((((.(((.........))).)).)))))))))).....)))((....))............................--..... ( -17.00) >DroSec_CAF1 29978 93 - 1 UGUAUAAAUGUUUUUUGGCAGCCUCUCUAAACGGCAGCGCAGGAAGCAUCACCAGCAG---CAGCACCACCAACACCAUUUACCACCUAAAA---AAAA- (((....((((((((((((.(((.........))).)).)))))))))).....((..---..)).......))).................---....- ( -16.50) >DroSim_CAF1 31628 94 - 1 UGUAUAAAUGUUUUUUGGCAGCCUCUCUAAACGGCAGCGCAGGAAGCAUCACCAGCAG---CAGCACCACCAACACCAUUUACCACCUAAAAA--AAAA- (((....((((((((((((.(((.........))).)).)))))))))).....((..---..)).......)))..................--....- ( -16.50) >DroEre_CAF1 33554 96 - 1 UGUAUAAAUGUUUUUUGGCAGCCUCUCUAAACGGCAGCGCAGGAAGCAUCACCAGCAG---CACCACCACCAACACCAUUUACCACCUAAAAAAAAAAA- (((....((((((((((((.(((.........))).)).)))))))))).....))).---......................................- ( -15.80) >DroYak_CAF1 42779 95 - 1 UGUAUAAAUGUUUUUUGGCAGCCUCUCUAAACGGCAGCGCAGGAAGCAUCACCAGCAG---CACCACCACCAACACCAUUUACCACCUAAAAA-AAAAA- (((....((((((((((((.(((.........))).)).)))))))))).....))).---................................-.....- ( -15.80) >consensus UGUAUAAAUGUUUUUUGGCAGCCUCUCUAAACGGCAGCGCAGGAAGCAUCACCAGCAG___CAGCACCACCAACACCAUUUACCACCUAAAAA__AAAA_ (((....((((((((((((.(((.........))).)).)))))))))).....)))........................................... (-15.80 = -15.80 + -0.00)

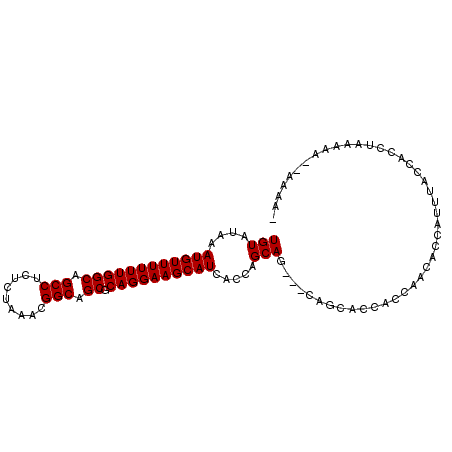

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:01 2006