
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,650,638 – 10,650,864 |
| Length | 226 |
| Max. P | 0.999317 |

| Location | 10,650,638 – 10,650,748 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 92.99 |
| Mean single sequence MFE | -27.08 |
| Consensus MFE | -25.04 |
| Energy contribution | -24.80 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.92 |
| SVM decision value | 3.51 |
| SVM RNA-class probability | 0.999317 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10650638 110 - 23771897 GACACUUACAAAUUUGUAUGUACAACACA------UAUACAUGCACACAAACGGACCAAGUACAUAAGUAUGCGUGUGCUUAAGGUCUGAAGAGUACGUAAUACUUUCAAUUUGAC ........((((((((((((((.......------..))))))))......((((((((((((((........))))))))..)))))).((((((.....)))))).)))))).. ( -27.10) >DroSec_CAF1 12964 110 - 1 GACACUUACAAAUUUGUAUGUACAACACA------UAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGCGUGUGCUUAGGGUCUGAAGAGUACGUAAUACUUUCAAUUUGAC ........((((((((((((((.......------..))))))))......((((((.(((((((........)))))))...)))))).((((((.....)))))).)))))).. ( -27.20) >DroSim_CAF1 12338 110 - 1 GACACUUACAAAUUUGUAUGUACAACACA------UAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGCGUGUGCUUAGGGUCUGAAGAGUACGUAAUACUUUCAAUUUGAC ........((((((((((((((.......------..))))))))......((((((.(((((((........)))))))...)))))).((((((.....)))))).)))))).. ( -27.20) >DroEre_CAF1 13400 116 - 1 GACACUUACAAAUUUGUAUGUACAACACAUGCAUAUGUACAUGCACACAAACGGACCAAGUACAUAAGUGUGAGUGUGUUUAGGGUCUGAAGAGUACGUAAUACUUUCAAUUUGAC ........((((((((((((((((...........))))))))))......((((((....(((((........)))))....)))))).((((((.....)))))).)))))).. ( -28.00) >DroYak_CAF1 13294 112 - 1 GACACUUACAAAUUUGUAUGUACAA----UGCAUACAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGAGUGUGUUUAGGGUCUGAAGAGUACGUAAUACUUUCAAUUUGAC ........((((((..(((((((..----(((((......)))))......((((((....(((((........)))))....))))))....)))))))........)))))).. ( -25.90) >consensus GACACUUACAAAUUUGUAUGUACAACACA______UAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGCGUGUGCUUAGGGUCUGAAGAGUACGUAAUACUUUCAAUUUGAC ........((((((((((((((...............))))))))......((((((((((((((........))))))))..)))))).((((((.....)))))).)))))).. (-25.04 = -24.80 + -0.24)



| Location | 10,650,678 – 10,650,771 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 92.80 |
| Mean single sequence MFE | -24.91 |
| Consensus MFE | -19.38 |
| Energy contribution | -19.58 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.945787 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10650678 93 - 23771897 GCUCUCUAGCUGAUAAGCAGGGCGACACUUACAAAUUUGUAUGUACAACACA------UAUACAUGCACACAAACGGACCAAGUACAUAAGUAUGCGUG (((((...(((....))))))))..............(((((((.....)))------))))((((((.((...................)).)))))) ( -18.11) >DroSec_CAF1 13004 93 - 1 GCUCUCUAGCUGAUAAGCAGGGCGACACUUACAAAUUUGUAUGUACAACACA------UAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGCGUG (((((...(((....))))))))..............(((((((.....)))------))))(((((((((...................))))))))) ( -24.11) >DroSim_CAF1 12378 93 - 1 GCUCUCUAGCUGAUAAGCAGGGCGACACUUACAAAUUUGUAUGUACAACACA------UAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGCGUG (((((...(((....))))))))..............(((((((.....)))------))))(((((((((...................))))))))) ( -24.11) >DroEre_CAF1 13440 99 - 1 GCUCUCUAGCUGAUAAGCAGGGCGACACUUACAAAUUUGUAUGUACAACACAUGCAUAUGUACAUGCACACAAACGGACCAAGUACAUAAGUGUGAGUG (((((...(((....))))))))..((((((((....(((((((.....)))))))(((((((.((..(......)...)).)))))))..)))))))) ( -30.50) >DroYak_CAF1 13334 95 - 1 GCUCUCUAGCUGAUAAGCAGGGCGACACUUACAAAUUUGUAUGUACAA----UGCAUACAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGAGUG (((((...(((....))))))))..((((((((......(((((((..----(((((......)))))(......)......)))))))..)))))))) ( -27.70) >consensus GCUCUCUAGCUGAUAAGCAGGGCGACACUUACAAAUUUGUAUGUACAACACA______UAUACAUGCACACAAACGGACCAAGUACAUAAGUGUGCGUG (((((...(((....)))))))).(((((((......((((((((...............)))))))).....((.......))...)))))))..... (-19.38 = -19.58 + 0.20)


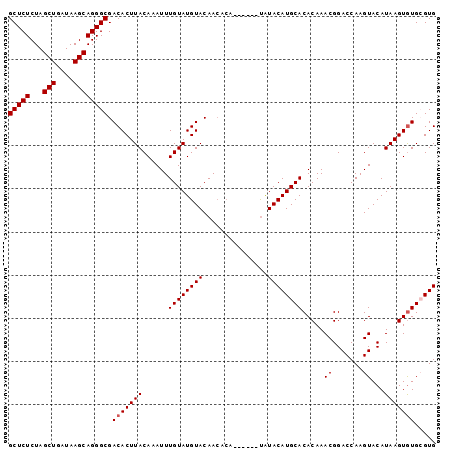
| Location | 10,650,771 – 10,650,864 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 82.34 |
| Mean single sequence MFE | -18.25 |
| Consensus MFE | -11.36 |
| Energy contribution | -11.92 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.682472 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
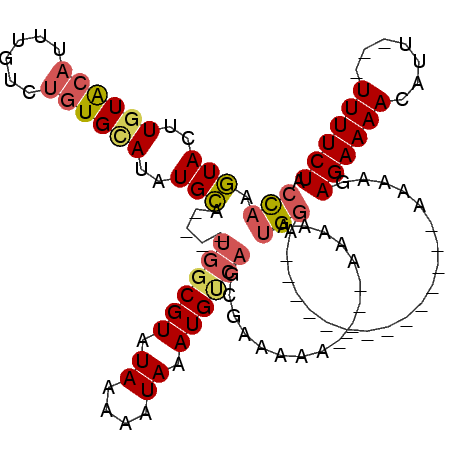
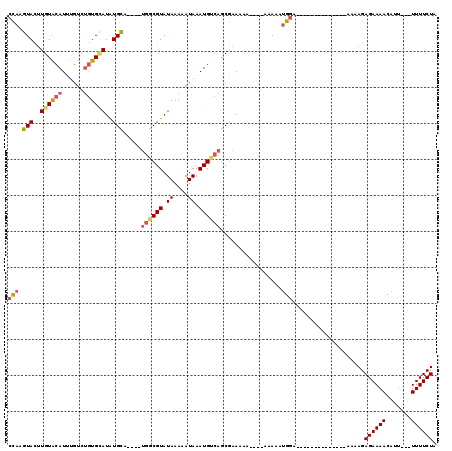
>3L_DroMel_CAF1 10650771 93 - 23771897 CCAAGUACUUGUACAUUUGUCUGUGAAUAUGCA----UGGCGUAUAAAAAUAAAUGUCAGCGAAAA-----AAAAAUGGA--------------AAAAGAGAAAACAUU---UUUUCUA (((.(((....)))...............(((.----((((((.((....)).)))))))))....-----.....))).--------------...(((((((....)---)))))). ( -14.10) >DroSec_CAF1 13097 94 - 1 CCAAGUACUUGUACAU-UGUCUGUGCAUAUGCA----UGGCGUAUAAAAAUAAAUGUCAGCGAAAAA---AAAAAAUGGA--------------AAAAGAGAAAACAUU---UUUUCUA (((.(((..((((((.-....))))))..))).----((((((.((....)).))))))........---......))).--------------...(((((((....)---)))))). ( -18.30) >DroSim_CAF1 12471 98 - 1 CCAAGUACUUGUACAUUUGUCUGUGCAUAUGCA----UGGCGUAUAAAAAUAAAUGUCAGCGAAAAAAAAAAAAAAUGGA--------------AAAAGAGAAAACAUU---UUUUCUA (((.(((..((((((......))))))..))).----((((((.((....)).)))))).................))).--------------...(((((((....)---)))))). ( -18.50) >DroEre_CAF1 13539 94 - 1 CCAAGUACUUGUACAUUUGUCUGUGCAUAUGCA----UGGCGUAUAAAAAUAAAUGCCAGCGAAAAA----AAAAAUGGA--------------AAAAGAGAAAACAUU---UUUUCUA (((.(((..((((((......))))))..))).----((((((.((....)).))))))........----.....))).--------------...(((((((....)---)))))). ( -21.20) >DroWil_CAF1 13575 116 - 1 AUACAUAUAUGUAGAUGUCCCGAUGUAUGUGUAAAAAGUACGUAUAAAAAUAAAUGCCAGCAGAAAAA---AAAAGCGGAAAAAUAUAUAAAGUAAAAGAGAAAACAUUCUGUUUUCUA .(((...((((((....(((...((((((((.......)))))))).............((.......---....)))))....))))))..)))....(((((((.....))))))). ( -19.10) >DroYak_CAF1 13429 98 - 1 CCAAGUACUUGUGCAUUUGUCUGUGCAUAUGCA---AUGGCGUAUAAAAAUAAAUGUCAGCGAAAAAA-AAAAAAAUGGA--------------AAAAGAGAAAACAUU---UUUUCUA (((......(((((((......)))))))(((.---.((((((.((....)).)))))))))......-.......))).--------------...(((((((....)---)))))). ( -18.30) >consensus CCAAGUACUUGUACAUUUGUCUGUGCAUAUGCA____UGGCGUAUAAAAAUAAAUGUCAGCGAAAAA____AAAAAUGGA______________AAAAGAGAAAACAUU___UUUUCUA (((.(((..((((((......))))))..))).....((((((.((....)).)))))).................)))....................((((((.......)))))). (-11.36 = -11.92 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:02:25 2006