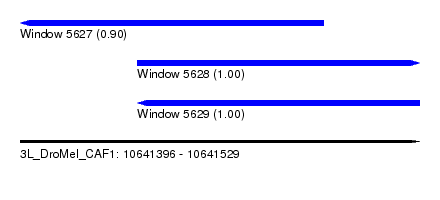

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,641,396 – 10,641,529 |
| Length | 133 |
| Max. P | 0.999351 |

| Location | 10,641,396 – 10,641,497 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 90.40 |
| Mean single sequence MFE | -35.50 |
| Consensus MFE | -28.25 |
| Energy contribution | -27.88 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.902289 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
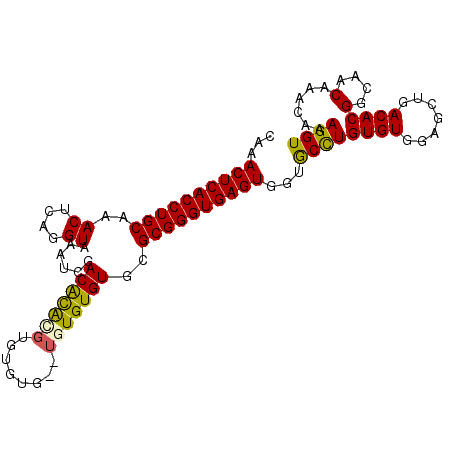
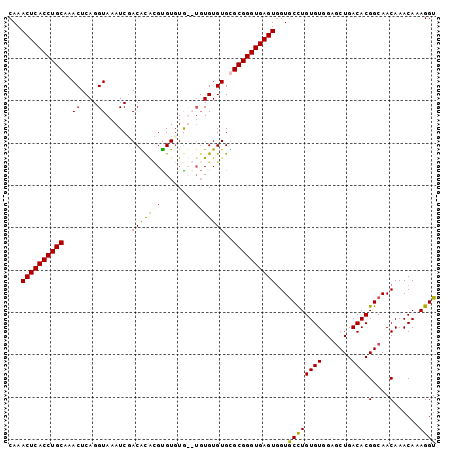
>3L_DroMel_CAF1 10641396 101 - 23771897 CAAACUCACCUGCAAACUCAGGUAAAUCGACACGCGUGUGUGUGUGUGUGUGCGCGGGUGAGUGGUACCUGUGUGGAGCUGACACGGCAACAAACAAAGGU ...((((((((((..((....)).....(.((((((..(....)..)))))))))))))))))...((((...((..((((...))))..)).....)))) ( -34.80) >DroSec_CAF1 3946 99 - 1 CAAACUCACCUGCAAACUCAGGUAAAUCGACGCGCGCGUGUGUG--UGUGUGUGCGGGUGAGUGGUGCCUGUGUGGAUCUGACACGGCAACAAACAAAGGU ...(((((((((((.((....))......((((((((....)))--))))).)))))))))))(.((((.((((.......)))))))).).......... ( -37.90) >DroEre_CAF1 3909 99 - 1 CAAACUCACCUGCAAACUCAGGUAAAUCGACACAUGUGUUUG--UGUGUGUGUGCGGGUGAGUUGUGCUUGUGUGGAGCUGACACGGCAACAAACAAAGGU ..((((((((((((.((....))......(((((..(....)--..))))).))))))))))))....((((.((..((((...))))..)).)))).... ( -33.40) >DroYak_CAF1 3923 99 - 1 CAAACUCACCUGCAAACUCAGGUAAAUCUACGUAAGUGUGUA--UGUGUGUGCGCGGGUGAGUGGUGCCUGUGUGGAGCUGACACGGCAACAAACAAAGGU ...((((((((((..((.((.(((....(((....)))..))--).)).))..))))))))))(.((((.((((.......)))))))).).......... ( -35.90) >consensus CAAACUCACCUGCAAACUCAGGUAAAUCGACACACGUGUGUG__UGUGUGUGCGCGGGUGAGUGGUGCCUGUGUGGAGCUGACACGGCAACAAACAAAGGU ...((((((((((..((....))......(((((((........)))))))..))))))))))...((((((((.......))))(....)......)))) (-28.25 = -27.88 + -0.37)

| Location | 10,641,435 – 10,641,529 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 67.82 |
| Mean single sequence MFE | -39.14 |
| Consensus MFE | -13.54 |
| Energy contribution | -15.22 |
| Covariance contribution | 1.68 |
| Combinations/Pair | 1.32 |
| Mean z-score | -4.63 |
| Structure conservation index | 0.35 |
| SVM decision value | 3.53 |
| SVM RNA-class probability | 0.999351 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
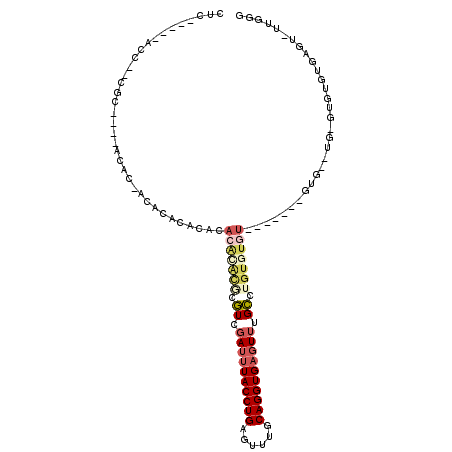
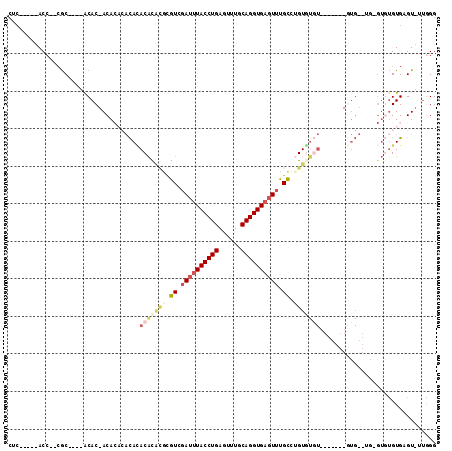
>3L_DroMel_CAF1 10641435 94 + 23771897 CUC-----ACC--CGC----GCAC-ACACACACACACACGCGUGUCGAUUUACCUGAGUUUGCAGGUGAGUUUGCCAGUGUCU-------GUGUUUGUGUGUGUGAGU-UUAGG ...-----((.--(((----((((-(((.(((((.((((..(.((.((((((((((......)))))))))).))).)))).)-------)))).)))))))))).))-..... ( -42.90) >DroSec_CAF1 3985 86 + 1 CUC-----ACC--CGC----ACAC-ACA--CACACACGCGCGCGUCGAUUUACCUGAGUUUGCAGGUGAGUUUGCCAGUGUGU-------GUG------AUUGUGAGU-UUGGG ...-----.((--((.----((.(-(((--..(((((((((..((.((((((((((......)))))))))).))..))))))-------)))------..)))).))-.)))) ( -39.50) >DroEre_CAF1 3948 91 + 1 CUC-----ACC--CGC----ACAC-ACACA--CAAACACAUGUGUCGAUUUACCUGAGUUUGCAGGUGAGUUUGCGUGUUUGU-------AUG--UGUGUGCGUGAGUUUUGGG (((-----((.--.((----((((-(((.(--(((((((....((.((((((((((......)))))))))).))))))))))-------.))--))))))))))))....... ( -44.30) >DroYak_CAF1 3962 80 + 1 CUC-----ACC--CGC----GCAC-ACACA--UACACACUUACGUAGAUUUACCUGAGUUUGCAGGUGAGUUUGCGUGU-------------------GUUUGUGAGU-UUGGG ...-----.((--((.----((.(-(((..--.((((((....(((((((((((((......)))))))))))))))))-------------------)).)))).))-.)))) ( -34.40) >DroAna_CAF1 3520 112 + 1 AUACAGAGACAGAGACAAAAACACAACA--CACAUACAUAAGCAUAAAUCUACCUGAGUUUGCAGGUGACUAUGUCUGUGUGUUCUGCUUGUGUGUGAGUGUGUGUGUUAGAGG ...................(((((((((--(.((((((((((((.......(((((......)))))(((...))).........)))))))))))).)))).))))))..... ( -34.60) >consensus CUC_____ACC__CGC____ACAC_ACACACACACACACACGCGUCGAUUUACCUGAGUUUGCAGGUGAGUUUGCCUGUGUGU_______GUG__UG_GUGUGUGAGU_UUGGG ...................................(((((((.((.((((((((((......)))))))))).)).)))))))............................... (-13.54 = -15.22 + 1.68)


| Location | 10,641,435 – 10,641,529 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 67.82 |
| Mean single sequence MFE | -32.15 |
| Consensus MFE | -8.79 |
| Energy contribution | -9.04 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.25 |
| Mean z-score | -3.88 |
| Structure conservation index | 0.27 |
| SVM decision value | 3.45 |
| SVM RNA-class probability | 0.999230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
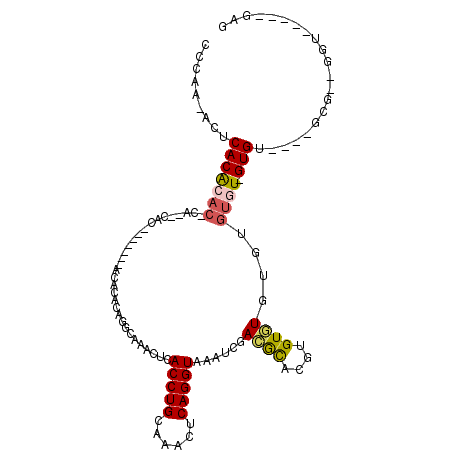
>3L_DroMel_CAF1 10641435 94 - 23771897 CCUAA-ACUCACACACACAAACAC-------AGACACUGGCAAACUCACCUGCAAACUCAGGUAAAUCGACACGCGUGUGUGUGUGUGU-GUGC----GCG--GGU-----GAG .....-((((.(.((((((.((((-------(.((((((........(((((......))))).........)).)))).))))).)))-))).----).)--)))-----... ( -32.93) >DroSec_CAF1 3985 86 - 1 CCCAA-ACUCACAAU------CAC-------ACACACUGGCAAACUCACCUGCAAACUCAGGUAAAUCGACGCGCGCGUGUGUG--UGU-GUGU----GCG--GGU-----GAG .....-.(((((...------(((-------((((((..........(((((......)))))......(((((....))))))--)))-))))----)..--.))-----))) ( -31.50) >DroEre_CAF1 3948 91 - 1 CCCAAAACUCACGCACACA--CAU-------ACAAACACGCAAACUCACCUGCAAACUCAGGUAAAUCGACACAUGUGUUUG--UGUGU-GUGU----GCG--GGU-----GAG .......((((((((((((--(((-------(((((((((.....(((((((......))))).....))....))))))))--)))))-))))----)).--.))-----))) ( -39.70) >DroYak_CAF1 3962 80 - 1 CCCAA-ACUCACAAAC-------------------ACACGCAAACUCACCUGCAAACUCAGGUAAAUCUACGUAAGUGUGUA--UGUGU-GUGC----GCG--GGU-----GAG .....-.(((((...(-------------------(((((((.((.((((((......)))))......((....))).)).--)))))-))).----...--.))-----))) ( -26.10) >DroAna_CAF1 3520 112 - 1 CCUCUAACACACACACUCACACACAAGCAGAACACACAGACAUAGUCACCUGCAAACUCAGGUAGAUUUAUGCUUAUGUAUGUG--UGUUGUGUUUUUGUCUCUGUCUCUGUAU ........(((.(.((....(((.(((((.((((((((.(((((((((((((......))))).))).......))))).))))--)))).))))).)))....)).).))).. ( -30.51) >consensus CCCAA_ACUCACACAC_CA__CAC_______ACACACAGGCAAACUCACCUGCAAACUCAGGUAAAUCGACGCACGUGUGUGUGUGUGU_GUGU____GCG__GGU_____GAG .........(((((((...............................(((((......)))))......((((....))))....)))).)))..................... ( -8.79 = -9.04 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:02:13 2006