
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,597,257 – 10,597,350 |
| Length | 93 |
| Max. P | 0.670400 |

| Location | 10,597,257 – 10,597,350 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 90.11 |
| Mean single sequence MFE | -35.04 |
| Consensus MFE | -26.12 |
| Energy contribution | -27.08 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.670400 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
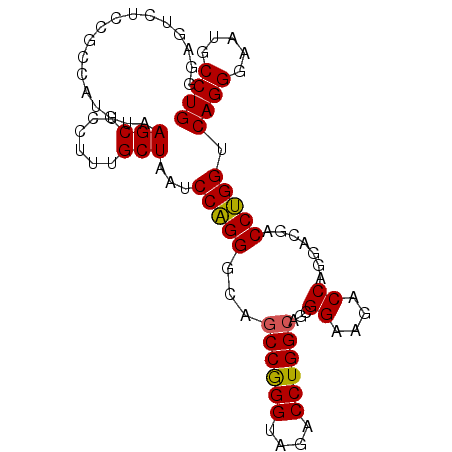
>3L_DroMel_CAF1 10597257 93 + 23771897 CAGGCAUUCCCUGACCAGGUCGUCCUGGUCUUCCGCUGCCAGGUCUACCCGGUUGCCCUGGAUUAGCAAAGGAGCUUACAUGUCGAAGACUCC ((((........(((((((....)))))))....((.(((.((....)).))).))))))..........(((((((........))).)))) ( -31.40) >DroSec_CAF1 479645 93 + 1 CAGGCAUUCCCUGACCAGGUCGUCCUGGUCUUCCGCUGCCAGGUCUACCCGGCUGCCCUGGAUUAGCAAAGGAGCUUACAUGGCGGAAACUCC .(((.....)))(((((((....)))))))((((((((((.((....)).))).(((((..........))).))......)))))))..... ( -35.70) >DroSim_CAF1 479703 93 + 1 CAGGCAUUCCCUGACCAGGUCGUCCUGGUCUUCCUCUGCCAGGUCUACCCGGCUGCCCUGGAUUAGCAAAGGAGCUUACAUGGCGGAGACUCC .(((.....)))(((((((....)))))))...((((((((((((.....))))(((((..........))).)).....))))))))..... ( -34.50) >DroEre_CAF1 486868 93 + 1 CAGGAAUUCCCUGGCCCGGUCGCCCUGGUAUUCCGGUGCCAGGUCUACCCGGCUGGCCAGGAUUAGCAAAGGAGCUUACAUGACGAAGACUUC .(((..((((((((((.(((((.((((((((....))))))))......))))))))))))...(((......)))........)))..))). ( -33.90) >DroYak_CAF1 489178 93 + 1 CAGGAAUUCCCUGGCCAGGUCGUCCUGGUAUUCCGGUGCCAGGUCUUCCUGGCUGUCCUGGAUUAGCGAAGUAGCUUACAUGGCGCAGACUUC (((((.......(((((((..(.((((((((....)))))))).)..))))))).))))).......(((((.(((.....)))....))))) ( -39.70) >consensus CAGGCAUUCCCUGACCAGGUCGUCCUGGUCUUCCGCUGCCAGGUCUACCCGGCUGCCCUGGAUUAGCAAAGGAGCUUACAUGGCGAAGACUCC ((((.....))))(((.(((.(.((((((........)))))).).))).)))((((.((....(((......)))..)).))))........ (-26.12 = -27.08 + 0.96)



| Location | 10,597,257 – 10,597,350 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 90.11 |
| Mean single sequence MFE | -36.72 |
| Consensus MFE | -25.72 |
| Energy contribution | -25.44 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.70 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10597257 93 - 23771897 GGAGUCUUCGACAUGUAAGCUCCUUUGCUAAUCCAGGGCAACCGGGUAGACCUGGCAGCGGAAGACCAGGACGACCUGGUCAGGGAAUGCCUG ((((((...))).....(((......)))..))).((((((((((((...(((((...(....).)))))...))))))).......))))). ( -34.31) >DroSec_CAF1 479645 93 - 1 GGAGUUUCCGCCAUGUAAGCUCCUUUGCUAAUCCAGGGCAGCCGGGUAGACCUGGCAGCGGAAGACCAGGACGACCUGGUCAGGGAAUGCCUG ((.(((((((((.((..(((......)))....)).))).((((((....)))))).......(((((((....))))))).)))))).)).. ( -41.30) >DroSim_CAF1 479703 93 - 1 GGAGUCUCCGCCAUGUAAGCUCCUUUGCUAAUCCAGGGCAGCCGGGUAGACCUGGCAGAGGAAGACCAGGACGACCUGGUCAGGGAAUGCCUG ((.((..((...........(((((((((.......))).((((((....)))))))))))).(((((((....))))))).))..)).)).. ( -38.90) >DroEre_CAF1 486868 93 - 1 GAAGUCUUCGUCAUGUAAGCUCCUUUGCUAAUCCUGGCCAGCCGGGUAGACCUGGCACCGGAAUACCAGGGCGACCGGGCCAGGGAAUUCCUG (((.((...........(((......)))...(((((((.((((((....))))))...((....)).((....)).))))))))).)))... ( -35.60) >DroYak_CAF1 489178 93 - 1 GAAGUCUGCGCCAUGUAAGCUACUUCGCUAAUCCAGGACAGCCAGGAAGACCUGGCACCGGAAUACCAGGACGACCUGGCCAGGGAAUUCCUG (((((..((.........)).))))).......(((((..((((((....)))))).((((....(((((....))))))).))....))))) ( -33.50) >consensus GGAGUCUCCGCCAUGUAAGCUCCUUUGCUAAUCCAGGGCAGCCGGGUAGACCUGGCAGCGGAAGACCAGGACGACCUGGUCAGGGAAUGCCUG .................(((......)))...(((((...((((((....))))))...((....)).......))))).((((.....)))) (-25.72 = -25.44 + -0.28)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:57 2006