
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,515,256 – 10,515,391 |
| Length | 135 |
| Max. P | 0.999960 |

| Location | 10,515,256 – 10,515,372 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 65.35 |
| Mean single sequence MFE | -22.50 |
| Consensus MFE | -13.52 |
| Energy contribution | -14.56 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.60 |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.943663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
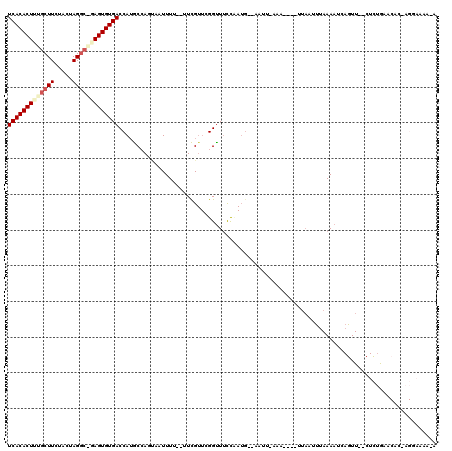
>3L_DroMel_CAF1 10515256 116 - 23771897 UCACACUUUGCUUCUACUAGGCGGAGUGUGACCAUGCAAGUAAUUUU--UUCGUUCGGUUUCCAAUGUUAAUUAAAAUACUUUAAUUUAAAAUCAGUU--CUCUAAACACCAGGAAAAGA (((((((((((((.....)))))))))))))((..(((((......)--)).))..))(((((......(((((((....)))))))........(((--.....)))....)))))... ( -23.30) >DroSec_CAF1 394755 93 - 1 UCACACUUUGCUUCUACUAG---AAGUGUGACCAUGCCAGUAAUUUU--UUCG---GGUUUUCU-----------------CUAAUUUAAAAUCAGUU--CUCUGAACACAAGGAAAAAA (((((((((..........)---))))))))............((((--((((---((....))-----------------)..........((((..--..))))......))))))). ( -18.00) >DroSim_CAF1 403140 102 - 1 UCACACUUUGCUUCUACUAGGCAGAGUGUGACCAUGCAAUGAAUUUU--UUCGUUCGGUUUCCAAUG--------------UUAAUUUAAAAUCAGUU--CUCUGAACACAAGGAAAAGA (((((((((((((.....)))))))))))))...((.((((((....--)))))))).((((((((.--------------...))).....((((..--..))))......)))))... ( -26.80) >DroEre_CAF1 401765 93 - 1 UCACACUUUGCUUUAGGUAGGC-UAGUGUGACCAUGCCACCCAUUUU--UUCGUUUGGCUUCUCAUGCGAAUUGAAAA-------UCGAAA--AAGUUUACUUGG--------------- (((((((..((((.....))))-.)))))))(((........(((((--((((((..(.(((......))))..))..-------.)))))--)))).....)))--------------- ( -20.72) >DroYak_CAF1 403487 103 - 1 UCACACUUUGCUUCAGGUAGGCGUAGUGUGACCAAACCAUUCAUUUUUCUUUGUUUGGUUACCCAUGUGAAUUGAAAUCCUUUAAUUUAAAAUAAGUU--CUUUG--------------- (((((((.(((((.....))))).)))))))...(((((..((........))..))))).......(((((((((....))))))))).........--.....--------------- ( -23.70) >consensus UCACACUUUGCUUCUACUAGGC_GAGUGUGACCAUGCCAGUAAUUUU__UUCGUUCGGUUUCCAAUG__AAUU_AAA____UUAAUUUAAAAUCAGUU__CUCUGAACAC_AGGAAAA_A (((((((((((((.....)))))))))))))......................................................................................... (-13.52 = -14.56 + 1.04)


| Location | 10,515,294 – 10,515,391 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 66.47 |
| Mean single sequence MFE | -25.85 |
| Consensus MFE | -17.19 |
| Energy contribution | -17.25 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.98 |
| Structure conservation index | 0.66 |
| SVM decision value | 4.90 |
| SVM RNA-class probability | 0.999960 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
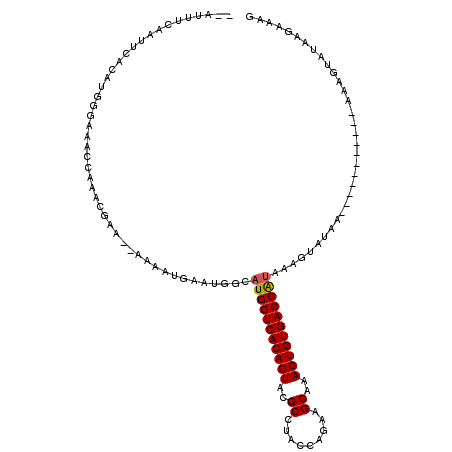
>3L_DroMel_CAF1 10515294 97 + 23771897 GUAUUUUAAUUAACAUUGGAAACCGAACGAA--AAAAUUACUUGCAUGGUCACACUCCGCCUAGUAGAAGCAAAGUGUGACCAUA-------------------AUAGUAUCAGAAAG ...............((((...)))).....--.....((((...(((((((((((..((.........))..))))))))))).-------------------..))))........ ( -22.00) >DroSim_CAF1 403177 92 + 1 -------------CAUUGGAAACCGAACGAA--AAAAUUCAUUGCAUGGUCACACUCUGCCUAGUAGAAGCAAAGUGUGACCGUAAACUCAAA-----------AUAGUAUCAGAAAU -------------....(....).(((((((--....))).....(((((((((((.(((.........))).))))))))))).........-----------...)).))...... ( -21.90) >DroEre_CAF1 401783 112 + 1 --UUUUCAAUUCGCAUGAGAAGCCAAACGAA--AAAAUGGGUGGCAUGGUCACACUA-GCCUACCUAAAGCAAAGUGUGACCAUAAAGUAUAAUUUUUGGCAAA-AAUUAUACGAUAG --((((((.......))))))((((..((..--....))..))))(((((((((((.-((.........))..)))))))))))...((((((((((.....))-))))))))..... ( -33.90) >DroYak_CAF1 403510 118 + 1 GGAUUUCAAUUCACAUGGGUAACCAAACAAAGAAAAAUGAAUGGUUUGGUCACACUACGCCUACCUGAAGCAAAGUGUGACCAUCAAGUAUAAUUUUUGGUAAAAAAGUAUAAGAAUG ........((((...(((....)))..((((((...(((..(((..((((((((((..((.........))..)))))))))))))..)))..))))))..............)))). ( -25.60) >consensus __AUUUCAAUUCACAUGGGAAACCAAACGAA__AAAAUGAAUGGCAUGGUCACACUACGCCUACCAGAAGCAAAGUGUGACCAUAAAGUAUAA___________AAAGUAUAAGAAAG .............................................(((((((((((..((.........))..))))))))))).................................. (-17.19 = -17.25 + 0.06)


| Location | 10,515,294 – 10,515,391 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 66.47 |
| Mean single sequence MFE | -31.23 |
| Consensus MFE | -22.92 |
| Energy contribution | -24.30 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.06 |
| Mean z-score | -4.42 |
| Structure conservation index | 0.73 |
| SVM decision value | 4.83 |
| SVM RNA-class probability | 0.999954 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
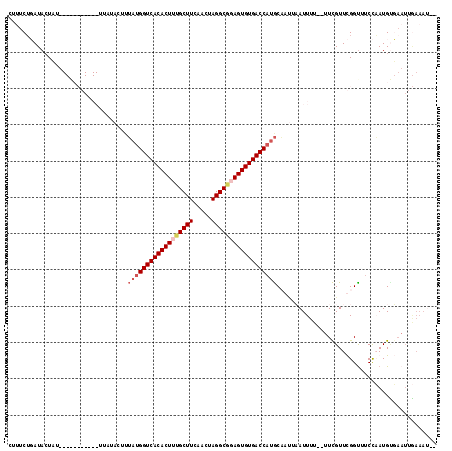
>3L_DroMel_CAF1 10515294 97 - 23771897 CUUUCUGAUACUAU-------------------UAUGGUCACACUUUGCUUCUACUAGGCGGAGUGUGACCAUGCAAGUAAUUUU--UUCGUUCGGUUUCCAAUGUUAAUUAAAAUAC ........((((..-------------------((((((((((((((((((.....))))))))))))))))))..))))...((--..((((.((...))))))..))......... ( -30.10) >DroSim_CAF1 403177 92 - 1 AUUUCUGAUACUAU-----------UUUGAGUUUACGGUCACACUUUGCUUCUACUAGGCAGAGUGUGACCAUGCAAUGAAUUUU--UUCGUUCGGUUUCCAAUG------------- ......((.(((..-----------...........(((((((((((((((.....)))))))))))))))..(.((((((....--)))))))))).)).....------------- ( -29.80) >DroEre_CAF1 401783 112 - 1 CUAUCGUAUAAUU-UUUGCCAAAAAUUAUACUUUAUGGUCACACUUUGCUUUAGGUAGGC-UAGUGUGACCAUGCCACCCAUUUU--UUCGUUUGGCUUCUCAUGCGAAUUGAAAA-- .....((((((((-((.....))))))))))..((((((((((((..((((.....))))-.)))))))))))).........((--((((.((.((.......)).)).))))))-- ( -35.00) >DroYak_CAF1 403510 118 - 1 CAUUCUUAUACUUUUUUACCAAAAAUUAUACUUGAUGGUCACACUUUGCUUCAGGUAGGCGUAGUGUGACCAAACCAUUCAUUUUUCUUUGUUUGGUUACCCAUGUGAAUUGAAAUCC ..............(((((................((((((((((.(((((.....))))).))))))))))(((((..((........))..)))))......)))))......... ( -30.00) >consensus CUUUCUGAUACUAU___________UUAUACUUUAUGGUCACACUUUGCUUCAACUAGGCGGAGUGUGACCAUGCAAUUAAUUUU__UUCGUUCGGUUUCCAAUGUGAAUUGAAAU__ .................................((((((((((((((((((.....))))))))))))))))))............................................ (-22.92 = -24.30 + 1.38)
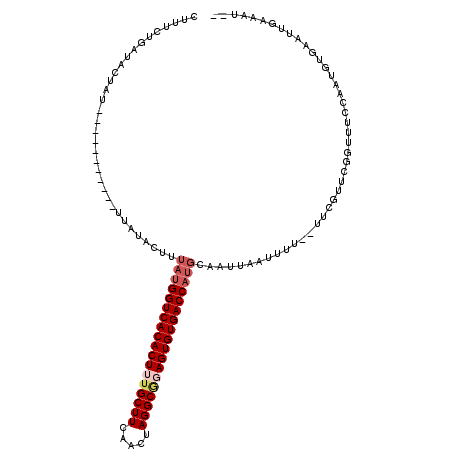


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:18 2006