
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,427,268 – 10,427,361 |
| Length | 93 |
| Max. P | 0.997197 |

| Location | 10,427,268 – 10,427,361 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.91 |
| Mean single sequence MFE | -30.47 |
| Consensus MFE | -26.70 |
| Energy contribution | -27.20 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.88 |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.986052 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
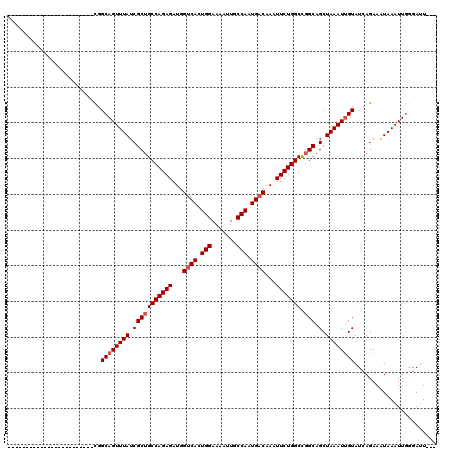
>3L_DroMel_CAF1 10427268 93 + 23771897 ------------------------CGGCAGUUUAUCGCUGCCAGAGAUGGUCACUGGAAAAUUGCCAAUGACAAAUUCUGGCCGGCAGCUAAAUUGUAUCAGAAAUAAAUUGGGAUU--- ------------------------..((((((((.(((((((((((...((((.(((.......))).))))...)))))).)))).).))))))))....................--- ( -29.00) >DroSec_CAF1 298274 93 + 1 ------------------------CGGCAGUUUAUCGCUGCCAGAGAUGGUCACUGGAAAAUUGCCAAUGACAAAUUCUGGCCGGCAGCUAAAUUGUAUCAGAAAUAAAUUGGGAUU--- ------------------------..((((((((.(((((((((((...((((.(((.......))).))))...)))))).)))).).))))))))....................--- ( -29.00) >DroSim_CAF1 313950 93 + 1 ------------------------CGGCAGUUUAUCGCUGCCAGAGAUGGUCACUGGAAAAUUGCCAAUGACAAAUUCUGGCCGGCAGCUAAAUUGUAUCAGAAAUAAAUUUGGAUU--- ------------------------..((((((((.(((((((((((...((((.(((.......))).))))...)))))).)))).).)))))))).(((((......)))))...--- ( -30.00) >DroEre_CAF1 309644 93 + 1 ------------------------CGGCAGUUUAUCGCUGCCAGAGAUGGUCACUGGAAAAUUGCCAAUGACAAAUUCUGGCCGGCAGCUAAAUUGUAUCAGAAGUAAAUUGGGAUU--- ------------------------..((((((((.(((((((((((...((((.(((.......))).))))...)))))).)))).).))))))))....................--- ( -29.00) >DroYak_CAF1 311637 93 + 1 ------------------------CGGCAGUUUAUCGCUGCCAGAGAUGGUCACUGGAAAAUUGCCAAUGACAAAUUCUGGCCGGCAGCUAAAUUGUAUCAGAAAUAAAUUGGGAUU--- ------------------------..((((((((.(((((((((((...((((.(((.......))).))))...)))))).)))).).))))))))....................--- ( -29.00) >DroAna_CAF1 303962 120 + 1 AGUUGCUGGUUGCCAGUUGGCGCUUGGCAGUUUAUCGCUGCCAGAGAUGGGCACUGGAAAAUUGCCAAUGACAAAUUCUGGCUUGCAGCUAAAUGGUAUAAGAAAUAAAUUGAGAAUAAG ((((((.....((((((((.((.((((((((((.(((.((((.......)))).))).)))))))))))).)))...)))))..)))))).............................. ( -36.80) >consensus ________________________CGGCAGUUUAUCGCUGCCAGAGAUGGUCACUGGAAAAUUGCCAAUGACAAAUUCUGGCCGGCAGCUAAAUUGUAUCAGAAAUAAAUUGGGAUU___ ..........................((((((((.(((((((((((...((((.(((.......))).))))...))))))).))).).))))))))....................... (-26.70 = -27.20 + 0.50)
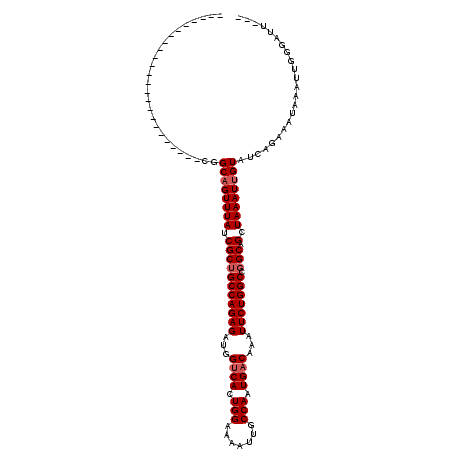


| Location | 10,427,268 – 10,427,361 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.91 |
| Mean single sequence MFE | -26.53 |
| Consensus MFE | -24.16 |
| Energy contribution | -24.35 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.98 |
| Structure conservation index | 0.91 |
| SVM decision value | 2.82 |
| SVM RNA-class probability | 0.997197 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
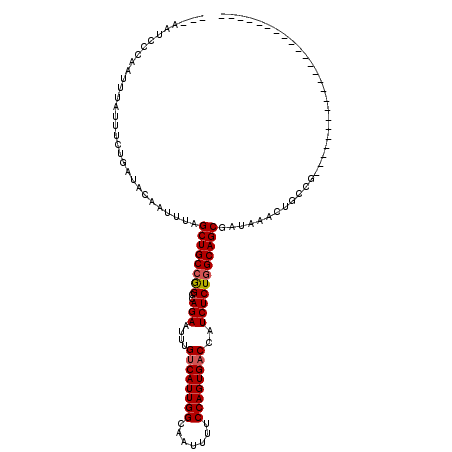
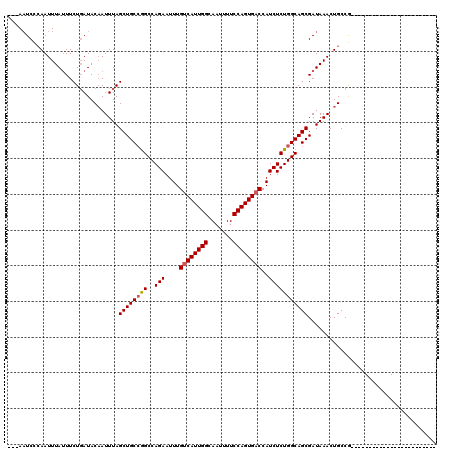
>3L_DroMel_CAF1 10427268 93 - 23771897 ---AAUCCCAAUUUAUUUCUGAUACAAUUUAGCUGCCGGCCAGAAUUUGUCAUUGGCAAUUUUCCAGUGACCAUCUCUGGCAGCGAUAAACUGCCG------------------------ ---.....((.((((((..(((......)))((((((((..(((....((((((((.......))))))))..))))))))))))))))).))...------------------------ ( -26.20) >DroSec_CAF1 298274 93 - 1 ---AAUCCCAAUUUAUUUCUGAUACAAUUUAGCUGCCGGCCAGAAUUUGUCAUUGGCAAUUUUCCAGUGACCAUCUCUGGCAGCGAUAAACUGCCG------------------------ ---.....((.((((((..(((......)))((((((((..(((....((((((((.......))))))))..))))))))))))))))).))...------------------------ ( -26.20) >DroSim_CAF1 313950 93 - 1 ---AAUCCAAAUUUAUUUCUGAUACAAUUUAGCUGCCGGCCAGAAUUUGUCAUUGGCAAUUUUCCAGUGACCAUCUCUGGCAGCGAUAAACUGCCG------------------------ ---............................((((((((..(((....((((((((.......))))))))..)))))))))))............------------------------ ( -25.90) >DroEre_CAF1 309644 93 - 1 ---AAUCCCAAUUUACUUCUGAUACAAUUUAGCUGCCGGCCAGAAUUUGUCAUUGGCAAUUUUCCAGUGACCAUCUCUGGCAGCGAUAAACUGCCG------------------------ ---............................((((((((..(((....((((((((.......))))))))..)))))))))))............------------------------ ( -25.90) >DroYak_CAF1 311637 93 - 1 ---AAUCCCAAUUUAUUUCUGAUACAAUUUAGCUGCCGGCCAGAAUUUGUCAUUGGCAAUUUUCCAGUGACCAUCUCUGGCAGCGAUAAACUGCCG------------------------ ---.....((.((((((..(((......)))((((((((..(((....((((((((.......))))))))..))))))))))))))))).))...------------------------ ( -26.20) >DroAna_CAF1 303962 120 - 1 CUUAUUCUCAAUUUAUUUCUUAUACCAUUUAGCUGCAAGCCAGAAUUUGUCAUUGGCAAUUUUCCAGUGCCCAUCUCUGGCAGCGAUAAACUGCCAAGCGCCAACUGGCAACCAGCAACU ...............................((((...(((((...(((.(.((((((.(((..(..((((.......))))..)..))).))))))..).))))))))...)))).... ( -28.80) >consensus ___AAUCCCAAUUUAUUUCUGAUACAAUUUAGCUGCCGGCCAGAAUUUGUCAUUGGCAAUUUUCCAGUGACCAUCUCUGGCAGCGAUAAACUGCCG________________________ ...............................((((((((..(((....((((((((.......))))))))..))))))))))).................................... (-24.16 = -24.35 + 0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:04 2006