
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,379,825 – 10,379,927 |
| Length | 102 |
| Max. P | 0.919932 |

| Location | 10,379,825 – 10,379,927 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 78.13 |
| Mean single sequence MFE | -26.20 |
| Consensus MFE | -18.79 |
| Energy contribution | -19.65 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.919932 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10379825 102 + 23771897 UAUAUACAUGCUGCUGAUAUGAAGAGCAUGUGU--UGUAGUGAAGUCACUUUCUACCCUCUCGCUCAAAUAGAGGGAUCACAUGACUUGAUUGCUUACACAGCC ...(((((((((.((.......)))))))))))--.(((((.((((((.......((((((.........))))))......)))))).))))).......... ( -31.02) >DroPse_CAF1 278653 87 + 1 UGUAAG-GGA---U------G-----GUGGUGU--GGCAGUAAAGUCACUUUCUACCCUCUCGCUUAAAUAGAGGGAUCACAUGACUUGAUUGCUUGCACAGAC ((((((-.((---(------(-----((((.((--(((......)))))...)))))(((((.........))))).............))).))))))..... ( -25.80) >DroEre_CAF1 268815 99 + 1 UACAUACAUA---CUGAUAUGGGGAGCAUGUGU--UGUAGUGAAGUCACUUUCUACCCUCUCGCUCAAAUAGAGGGAUCACAUGACUUGAUUGCUUACACAGCC ..(((((...---..).))))....((.(((((--.(((((.((((((.......((((((.........))))))......)))))).)))))..))))))). ( -28.62) >DroWil_CAF1 405270 90 + 1 U--AUG-UUU---GUUAU--UUGGGGCU--AGC----UGCUAAAGUCACUUUCUACCCUCUCGCUUAAAUAGAGGGAUCACAUGACUUGAUUGCUUACACAGCC .--...-...---.....--....((((--(((----.....((((((.......((((((.........))))))......))))))....))).....)))) ( -20.02) >DroYak_CAF1 266180 99 + 1 UAAAUACAUA---CCGAUAUGAAGAGCAUGUGU--UGUAGUGAAGUCACUUUCUACCCUCUCGCUCAAAUAGAGGGAUCACAUGACUUGAUUGCUUACACAGCC ......((((---....))))....((.(((((--.(((((.((((((.......((((((.........))))))......)))))).)))))..))))))). ( -28.32) >DroAna_CAF1 259482 97 + 1 UGUAAA-GGA---UUCAU--G-GGGGGAUGAGAUACGGAGUUGAGUCACUUUCUACCCUCUCGCUCAAAUAGAGGGAUCACAUGACUUGAUUGCUUACACAGCC (((((.-...---(((((--.-.....)))))....(.(((..(((((.......((((((.........))))))......)))))..))).))))))..... ( -23.42) >consensus UAUAUA_AUA___CUGAU__G_GGAGCAUGUGU__UGUAGUGAAGUCACUUUCUACCCUCUCGCUCAAAUAGAGGGAUCACAUGACUUGAUUGCUUACACAGCC .............................((((...(((((.((((((.......((((((.........))))))......)))))).)))))..)))).... (-18.79 = -19.65 + 0.86)



| Location | 10,379,825 – 10,379,927 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 78.13 |
| Mean single sequence MFE | -23.93 |
| Consensus MFE | -18.02 |
| Energy contribution | -18.18 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813181 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10379825 102 - 23771897 GGCUGUGUAAGCAAUCAAGUCAUGUGAUCCCUCUAUUUGAGCGAGAGGGUAGAAAGUGACUUCACUACA--ACACAUGCUCUUCAUAUCAGCAGCAUGUAUAUA .(((.....)))....(((((((.....((((((.........))))))......))))))).......--..(((((((((.......)).)))))))..... ( -27.80) >DroPse_CAF1 278653 87 - 1 GUCUGUGCAAGCAAUCAAGUCAUGUGAUCCCUCUAUUUAAGCGAGAGGGUAGAAAGUGACUUUACUGCC--ACACCAC-----C------A---UCC-CUUACA ...((((...(((.(.(((((((.....((((((.........))))))......))))))).).))))--)))....-----.------.---...-...... ( -21.90) >DroEre_CAF1 268815 99 - 1 GGCUGUGUAAGCAAUCAAGUCAUGUGAUCCCUCUAUUUGAGCGAGAGGGUAGAAAGUGACUUCACUACA--ACACAUGCUCCCCAUAUCAG---UAUGUAUGUA .(((.....)))....(((((((.....((((((.........))))))......)))))))...((((--..(((((((.........))---))))).)))) ( -25.60) >DroWil_CAF1 405270 90 - 1 GGCUGUGUAAGCAAUCAAGUCAUGUGAUCCCUCUAUUUAAGCGAGAGGGUAGAAAGUGACUUUAGCA----GCU--AGCCCCAA--AUAAC---AAA-CAU--A (((((.((..((....(((((((.....((((((.........))))))......)))))))..)).----)))--))))....--.....---...-...--. ( -24.40) >DroYak_CAF1 266180 99 - 1 GGCUGUGUAAGCAAUCAAGUCAUGUGAUCCCUCUAUUUGAGCGAGAGGGUAGAAAGUGACUUCACUACA--ACACAUGCUCUUCAUAUCGG---UAUGUAUUUA .(((.....)))....(((((((.....((((((.........))))))......))))))).......--..(((((((.........))---)))))..... ( -24.60) >DroAna_CAF1 259482 97 - 1 GGCUGUGUAAGCAAUCAAGUCAUGUGAUCCCUCUAUUUGAGCGAGAGGGUAGAAAGUGACUCAACUCCGUAUCUCAUCCCCC-C--AUGAA---UCC-UUUACA .(((.....))).....((((((.....((((((.........))))))......)))))).......(((..((((.....-.--)))).---...-..))). ( -19.30) >consensus GGCUGUGUAAGCAAUCAAGUCAUGUGAUCCCUCUAUUUGAGCGAGAGGGUAGAAAGUGACUUCACUACA__ACACAUGCUCC_C__AUCAG___UAU_UAUAUA (..(((....)))..)(((((((.....((((((.........))))))......))))))).......................................... (-18.02 = -18.18 + 0.17)


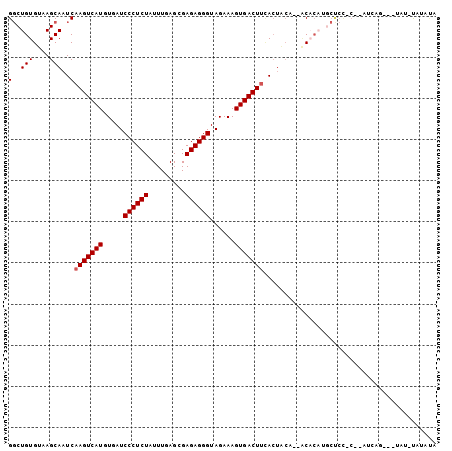
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:00:42 2006