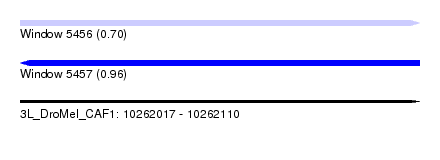
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,262,017 – 10,262,110 |
| Length | 93 |
| Max. P | 0.958461 |

| Location | 10,262,017 – 10,262,110 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 84.31 |
| Mean single sequence MFE | -16.45 |
| Consensus MFE | -11.99 |
| Energy contribution | -12.03 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.697858 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
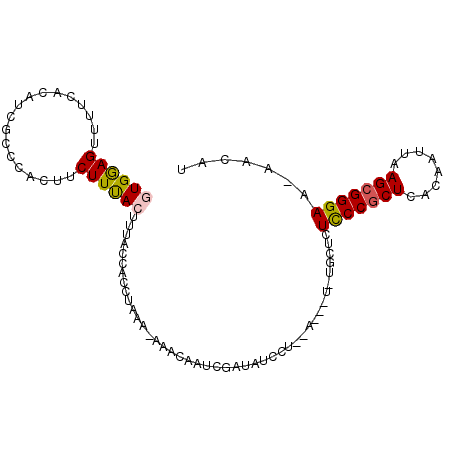
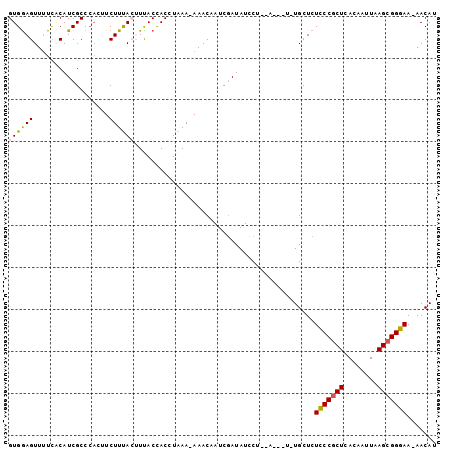
>3L_DroMel_CAF1 10262017 93 + 23771897 GUGAAGUUUUCGCAUCGCCCACUUCUUUACUUUACCACCUAAAAAAACAAUCGAUAUCCU--------UGCUCUUCCGCUCACAAUUAAGCGGGAA-AACAA .....(((((((((..............................................--------)))....(((((........))))))))-))).. ( -13.45) >DroSec_CAF1 137300 92 + 1 GUGGAGUUUUCACAUCGCCCACUUCUUUACCUUACCACCUAAA-AAACAAUCGAUAUCCU--------UGCUCUCCCGCUCACAAUUAAGCGGGAA-AACAU ((((.((.........)))))).....................-................--------.....(((((((........))))))).-..... ( -16.40) >DroSim_CAF1 142697 97 + 1 GUGGAGUUUUCACAUCGCCCACUUCUUUACCUUACCACCUAAA-AAACAAUCGAUAUCCUCGA---UAUGCUCUCCCGCUCACAAUUAAGCGGGAA-AACAU ((((.((.........)))))).....................-.....(((((.....))))---)......(((((((........))))))).-..... ( -19.90) >DroEre_CAF1 147177 99 + 1 GUGGAGUU-UUCCAUCGCCCAGCUCUUCAGUUCACCACCUAAA-AAACAAUCAAUAUCCUUCAGCUUUUGCUCUCCCGCUCACAAUUAAGCGGGAA-AACAA (.((((((-...........)))))).).(((...........-..................(((....))).(((((((........))))))).-))).. ( -17.80) >DroYak_CAF1 144615 97 + 1 GUGGAGUUUUUCCAUCGCC----UCUUCAGUUCACUACCUAAA-AAACAAUCGAUAUACUUCAGCUUUUGCUCUCCCACUCACAAUUAAGCGGGAAAAACAU .....((((((((..(((.----....................-........((......))(((....))).................))))))))))).. ( -14.70) >consensus GUGGAGUUUUCACAUCGCCCACUUCUUUACUUUACCACCUAAA_AAACAAUCGAUAUCCU__A___U_UGCUCUCCCGCUCACAAUUAAGCGGGAA_AACAU ((((((..................))))))...........................................(((((((........)))))))....... (-11.99 = -12.03 + 0.04)

| Location | 10,262,017 – 10,262,110 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 84.31 |
| Mean single sequence MFE | -30.72 |
| Consensus MFE | -18.37 |
| Energy contribution | -18.01 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.13 |
| Mean z-score | -3.26 |
| Structure conservation index | 0.60 |
| SVM decision value | 1.49 |
| SVM RNA-class probability | 0.958461 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
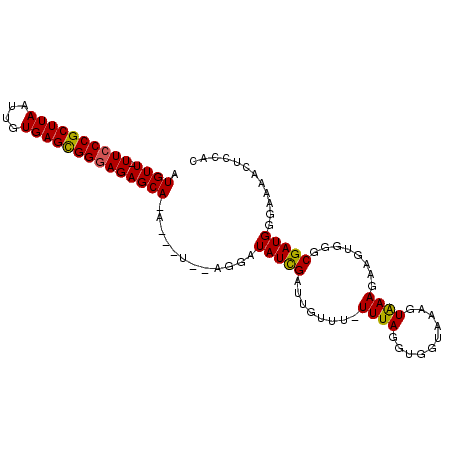
>3L_DroMel_CAF1 10262017 93 - 23771897 UUGUU-UUCCCGCUUAAUUGUGAGCGGAAGAGCA--------AGGAUAUCGAUUGUUUUUUUAGGUGGUAAAGUAAAGAAGUGGGCGAUGCGAAAACUUCAC (((((-((.(((((((....))))))).))))))--------).(.(((((.(..((((((((..(....)..))))))))..).))))))........... ( -25.60) >DroSec_CAF1 137300 92 - 1 AUGUU-UUCCCGCUUAAUUGUGAGCGGGAGAGCA--------AGGAUAUCGAUUGUUU-UUUAGGUGGUAAGGUAAAGAAGUGGGCGAUGUGAAAACUCCAC .((((-((((((((((....))))))))))))))--------.(((.((((.(..(((-(((.(.(.....).).))))))..).))))((....))))).. ( -31.40) >DroSim_CAF1 142697 97 - 1 AUGUU-UUCCCGCUUAAUUGUGAGCGGGAGAGCAUA---UCGAGGAUAUCGAUUGUUU-UUUAGGUGGUAAGGUAAAGAAGUGGGCGAUGUGAAAACUCCAC (((((-((((((((((....))))))))))))))).---..(((.((((((.(..(((-(((.(.(.....).).))))))..).)))))).....)))... ( -32.20) >DroEre_CAF1 147177 99 - 1 UUGUU-UUCCCGCUUAAUUGUGAGCGGGAGAGCAAAAGCUGAAGGAUAUUGAUUGUUU-UUUAGGUGGUGAACUGAAGAGCUGGGCGAUGGAA-AACUCCAC (((((-((((((((((....)))))))))))))))........((((((((...((((-(((((........)))))))))....)))))...-...))).. ( -35.40) >DroYak_CAF1 144615 97 - 1 AUGUUUUUCCCGCUUAAUUGUGAGUGGGAGAGCAAAAGCUGAAGUAUAUCGAUUGUUU-UUUAGGUAGUGAACUGAAGA----GGCGAUGGAAAAACUCCAC .((((((.((((((((....)))))))))))))).....((.(((.(.((.(((((((-(((.(((.....))).))))----)))))).)).).))).)). ( -29.00) >consensus AUGUU_UUCCCGCUUAAUUGUGAGCGGGAGAGCA_A___U__AGGAUAUCGAUUGUUU_UUUAGGUGGUAAAGUAAAGAAGUGGGCGAUGGGAAAACUCCAC .((((.((((((((((....))))))))))))))............(((((........((((..........))))........)))))............ (-18.37 = -18.01 + -0.36)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:59:30 2006