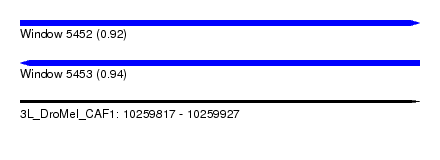
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,259,817 – 10,259,927 |
| Length | 110 |
| Max. P | 0.935454 |

| Location | 10,259,817 – 10,259,927 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 95.24 |
| Mean single sequence MFE | -25.95 |
| Consensus MFE | -23.44 |
| Energy contribution | -23.22 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.915732 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
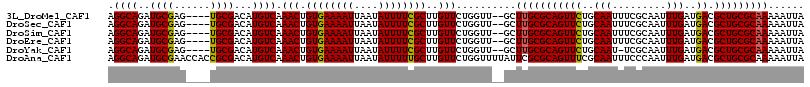
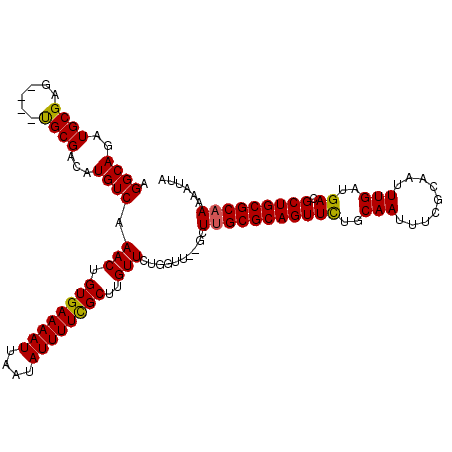
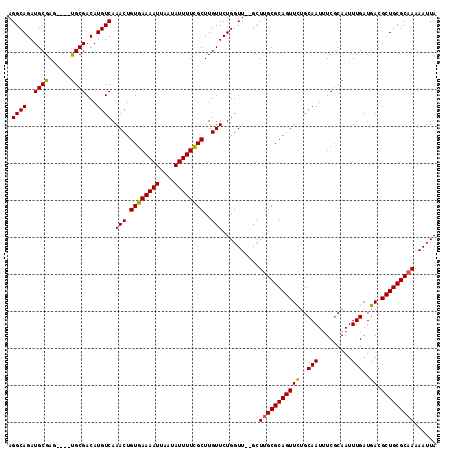
>3L_DroMel_CAF1 10259817 110 + 23771897 UAAUUUUUGCGCAGCGUCAUCAAAUUGCGAAAUUGCAGAACUGCGCAAGC--AACCAGAACAAGCGAAAAUAUUAAUUUUCACAGUUUGACAUGUCGCA----CUCGCAUCUGCCU .......((((..(((.((((((((((..............(((....))--)............((((((....)))))).))))))))..)).))).----..))))....... ( -26.00) >DroSec_CAF1 135163 110 + 1 UAAUUUUUGCGCAGCGUCAUCAAAUUGCGAAAUUGCAGAACUGCGCAAGC--AACCAGAACAAGCGAAAAUAUUAAUUUUCACAGUUUGACAUGUCGCA----CUCGCAUCUGCCU .......((((..(((.((((((((((..............(((....))--)............((((((....)))))).))))))))..)).))).----..))))....... ( -26.00) >DroSim_CAF1 140525 110 + 1 UAAUUUUUGCGCAGCGUCAUCAAAUUGCGAAAUUGCAGAACUGCGCAAGC--AACCAGAACAAGCGAAAAUAUUAAUUUUCACAGUUUGACAUGUCGCA----CUCGCAUCUGCCU .......((((..(((.((((((((((..............(((....))--)............((((((....)))))).))))))))..)).))).----..))))....... ( -26.00) >DroEre_CAF1 144823 110 + 1 UAAUUUUUGCGCAGCGUCAUCAAAUUGCGAAAUUGCAGAACUGCGCAAGC--AACCAGAACAAGCGAAAAUAUUAAUUUUCACAGUUUGACAUGUCGCA----CUCGCAUCUGCCU .......((((..(((.((((((((((..............(((....))--)............((((((....)))))).))))))))..)).))).----..))))....... ( -26.00) >DroYak_CAF1 142161 109 + 1 UAAUUUUUGCGCAGCGUCAUCAAAUUGCGA-AUUGCAGAACUGCGCAAGC--AACCAGAACAAGCGAAAAUAUUAAUUUUCACAGUUUGACAUGUCGCA----CUCGCAUCUGCCU .......((((..(((.((((((((((...-..........(((....))--)............((((((....)))))).))))))))..)).))).----..))))....... ( -26.00) >DroAna_CAF1 148641 116 + 1 UAAUUUUUGCGCAGCGUCAUCAAAUUGGGAAAUUGCGAAACUGCGCGAAUAAAACCAGAACAAGCAAAAAUAUUAAUUUUCACAGUUUGACAUGUCGCGGUGGUUCGCAUCUGCCU .............(((.((((((((((((((((((((....)))((.................)).........))))))).))))))))..)).)))((..(.......)..)). ( -25.73) >consensus UAAUUUUUGCGCAGCGUCAUCAAAUUGCGAAAUUGCAGAACUGCGCAAGC__AACCAGAACAAGCGAAAAUAUUAAUUUUCACAGUUUGACAUGUCGCA____CUCGCAUCUGCCU .....(((((((((.(....)...(((((....)))))..)))))))))......((((.(((((((((((....))))))...))))).......((........)).))))... (-23.44 = -23.22 + -0.22)
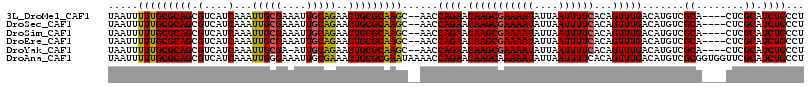
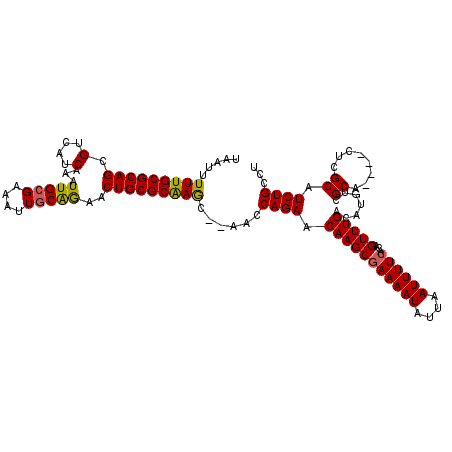
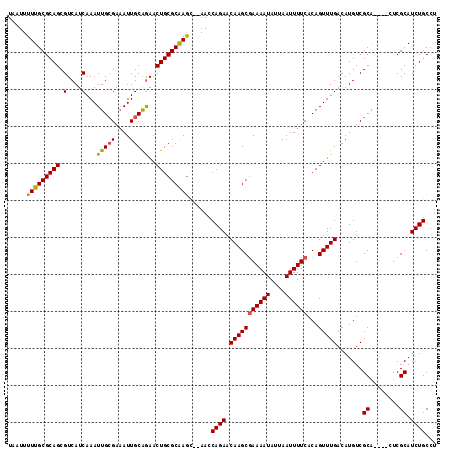
| Location | 10,259,817 – 10,259,927 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 95.24 |
| Mean single sequence MFE | -30.22 |
| Consensus MFE | -27.63 |
| Energy contribution | -27.38 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.935454 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10259817 110 - 23771897 AGGCAGAUGCGAG----UGCGACAUGUCAAACUGUGAAAAUUAAUAUUUUCGCUUGUUCUGGUU--GCUUGCGCAGUUCUGCAAUUUCGCAAUUUGAUGACGCUGCGCAAAAAUUA ..(((((((((..----.(((((......(((.((((((((....))))))))..)))...)))--))...))))..)))))..(((((((............)))).)))..... ( -30.30) >DroSec_CAF1 135163 110 - 1 AGGCAGAUGCGAG----UGCGACAUGUCAAACUGUGAAAAUUAAUAUUUUCGCUUGUUCUGGUU--GCUUGCGCAGUUCUGCAAUUUCGCAAUUUGAUGACGCUGCGCAAAAAUUA ..(((((((((..----.(((((......(((.((((((((....))))))))..)))...)))--))...))))..)))))..(((((((............)))).)))..... ( -30.30) >DroSim_CAF1 140525 110 - 1 AGGCAGAUGCGAG----UGCGACAUGUCAAACUGUGAAAAUUAAUAUUUUCGCUUGUUCUGGUU--GCUUGCGCAGUUCUGCAAUUUCGCAAUUUGAUGACGCUGCGCAAAAAUUA ..(((((((((..----.(((((......(((.((((((((....))))))))..)))...)))--))...))))..)))))..(((((((............)))).)))..... ( -30.30) >DroEre_CAF1 144823 110 - 1 AGGCAGAUGCGAG----UGCGACAUGUCAAACUGUGAAAAUUAAUAUUUUCGCUUGUUCUGGUU--GCUUGCGCAGUUCUGCAAUUUCGCAAUUUGAUGACGCUGCGCAAAAAUUA ..(((((((((..----.(((((......(((.((((((((....))))))))..)))...)))--))...))))..)))))..(((((((............)))).)))..... ( -30.30) >DroYak_CAF1 142161 109 - 1 AGGCAGAUGCGAG----UGCGACAUGUCAAACUGUGAAAAUUAAUAUUUUCGCUUGUUCUGGUU--GCUUGCGCAGUUCUGCAAU-UCGCAAUUUGAUGACGCUGCGCAAAAAUUA ..((((...((((----(((((..(((..((((((((((((....))))))((..((.......--))..))))))))..)))..-)))).)))))......)))).......... ( -30.30) >DroAna_CAF1 148641 116 - 1 AGGCAGAUGCGAACCACCGCGACAUGUCAAACUGUGAAAAUUAAUAUUUUUGCUUGUUCUGGUUUUAUUCGCGCAGUUUCGCAAUUUCCCAAUUUGAUGACGCUGCGCAAAAAUUA .((((..((((......))))...)))).(((.(..(((((....)))))..)..)))............(((((((.((((((.........))).))).)))))))........ ( -29.80) >consensus AGGCAGAUGCGAG____UGCGACAUGUCAAACUGUGAAAAUUAAUAUUUUCGCUUGUUCUGGUU__GCUUGCGCAGUUCUGCAAUUUCGCAAUUUGAUGACGCUGCGCAAAAAUUA .((((..((((......))))...)))).(((.((((((((....))))))))..)))..........(((((((((((..(((.........)))..)).)))))))))...... (-27.63 = -27.38 + -0.25)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:59:27 2006